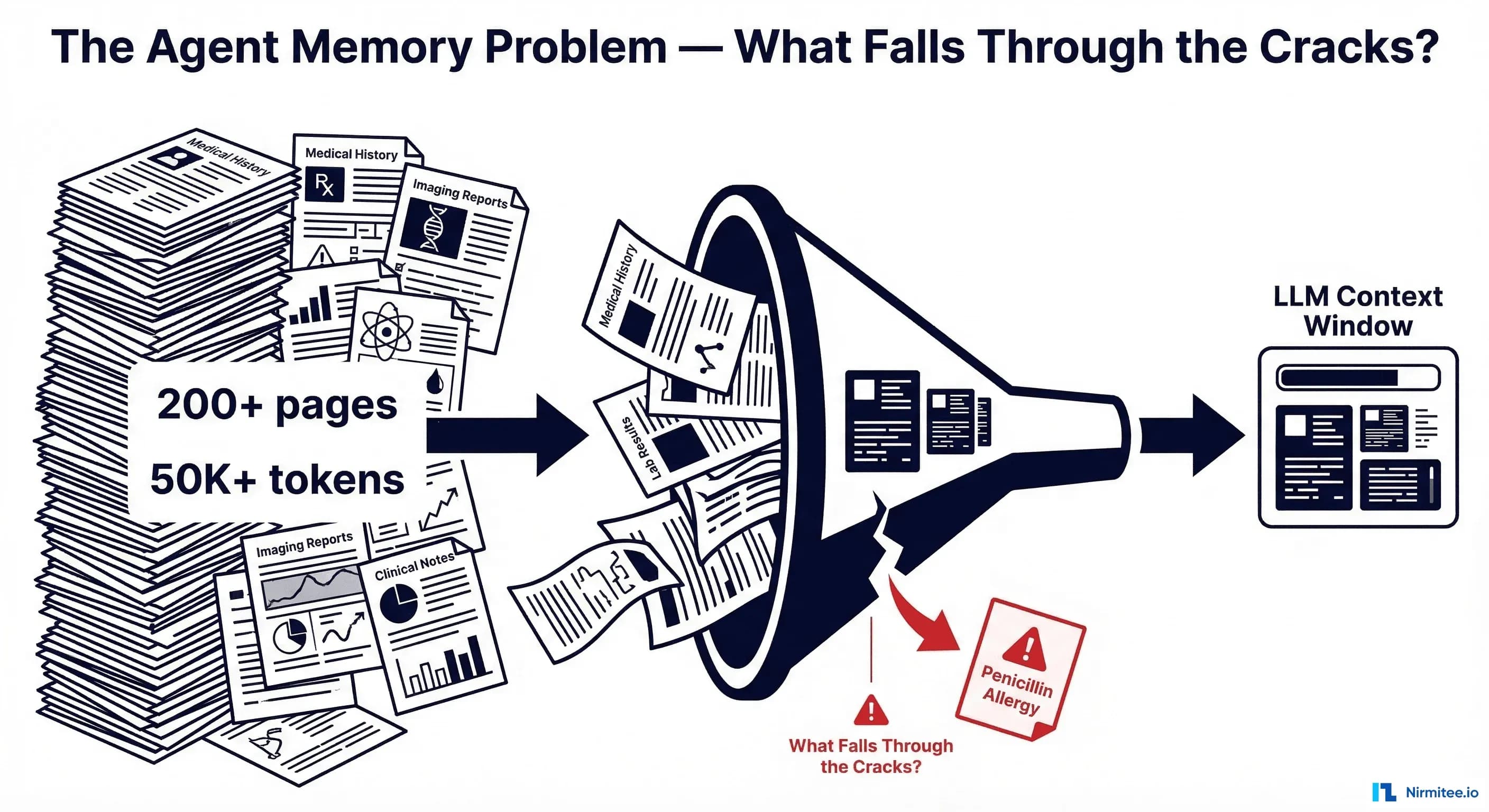
A 63-year-old patient walks into an emergency department with chest pain. The attending physician asks the clinical AI agent for a medication recommendation. The agent processes the query, searches the patient's records, and suggests a beta-blocker. Reasonable. Except that buried on page 47 of the patient's medical history — in a 2019 encounter note from a different health system — there's a documented severe bradycardia episode triggered by a beta-blocker. The agent never saw it.
This is the agent memory problem in healthcare. It is not a theoretical concern. It is a patient safety risk that every engineering team building clinical AI agents must solve before deploying to production.
The Scale of the Problem: 200 Pages, 50,000 Tokens, One Missed Allergy
A typical adult patient with chronic conditions accumulates between 50 and 200 pages of medical records per year of care. A patient who has been in the healthcare system for 20 years may have records exceeding 1,000 pages — clinical notes, lab results, imaging reports, medication lists, allergy documentation, procedure notes, referral letters, and discharge summaries. When converted to tokens, a comprehensive patient history can easily exceed 50,000 tokens. Complex patients with multiple comorbidities can reach 150,000+ tokens.
Meanwhile, the critical information your agent needs might be a single sentence: "Patient experienced anaphylaxis to penicillin on 03/15/2019." That sentence is approximately 12 tokens. Finding those 12 tokens in a 50,000-token haystack is the core engineering challenge.
The stakes are high. According to research published in the Journal of Emergency Medicine, nearly 41% of patients had at least one discrepancy between their self-reported allergies and what was documented in their electronic medical record. Over 10% of inpatient medication errors are attributable to accidental delivery of a medication to which a patient has a known allergy. When an AI agent misses this information, it compounds an already dangerous documentation gap.
The Naive Approach: Stuff Everything Into Context
The first instinct of most engineering teams is straightforward: just put everything into the context window. Modern LLMs support 128K to 200K tokens. Some support up to 1M. Why not just feed the entire patient history?
Four reasons this fails in production:
1. Cost scales linearly with input tokens. At current 2026 pricing, processing 50K input tokens through Claude Sonnet 4 costs approximately $0.15 per call. Through GPT-5.2, approximately $0.09. That sounds manageable until you realize a clinical agent handles hundreds of queries per patient encounter, and a health system processes thousands of encounters per day. At 200 queries per encounter across 500 daily encounters, you are looking at $15,000/day in API costs for a single facility — just for input tokens.
2. Latency becomes clinically unacceptable. Processing 50K+ tokens adds 8-15 seconds of latency before the model even begins generating a response. In an emergency department, where decisions happen in minutes, a 15-second delay per query is a non-starter. Multiply that across a multi-turn clinical reasoning session, and the agent becomes slower than looking it up manually.
3. The "Lost in the Middle" phenomenon. Research by Liu et al. (2024), published in Transactions of the Association for Computational Linguistics, demonstrated that LLMs exhibit a U-shaped attention curve: they attend well to information at the beginning and end of the context window, but accuracy degrades significantly for information in the middle. Their study showed a 30%+ accuracy drop on multi-document question answering when the answer was positioned in the middle versus the start of the context. For clinical safety data — an allergy buried in the middle of a 50K-token context — this means a substantial probability of the agent simply not finding it.
4. Token limits are still real. Even with 200K-token context windows, many patients' complete records exceed this limit. When you add the system prompt, few-shot examples, tool definitions, and conversation history, the available space for patient data shrinks considerably. You are often working with 80-120K tokens of effective capacity for clinical data.
RAG for Clinical Data: Building the Retrieval Pipeline
Retrieval-Augmented Generation (RAG) is the standard architectural answer to the context window limitation. Instead of stuffing everything into context, you index the patient's records in a vector database, retrieve only the relevant chunks for each query, and inject those into the context alongside the query. Recent surveys confirm RAG as the dominant approach for healthcare LLM applications, significantly reducing hallucination and improving factual accuracy.
But clinical RAG is not the same as building a chatbot over documentation. The data is structured differently, the stakes are higher, and the failure modes are more dangerous.
What to Index: FHIR Resources as Your Source of Truth
If your source data follows the HL7 FHIR standard (and in 2026, under ONC and CMS interoperability mandates, it increasingly does), you have a natural unit of indexing: the FHIR resource. The key resource types for clinical agent retrieval are:
- AllergyIntolerance — documented allergies and adverse reactions
- Condition — diagnoses, both active and resolved
- MedicationRequest — prescribed medications with status
- Observation — lab results, vital signs, social history
- Encounter — visit records with associated notes
- DiagnosticReport — imaging, pathology, and other diagnostic results
- DocumentReference — unstructured clinical notes, discharge summaries
FHIR is a strong starting point because it already breaks data into consumable chunks. However, as researchers at the NLP Summit noted, raw FHIR JSON is not directly suitable for embedding — it needs to be transformed into natural language representations that capture the clinical meaning.
Chunking Strategies: Per-Resource vs. Per-Encounter vs. Per-Concept
How you chunk clinical data determines the quality of your retrieval. There are three primary strategies:
Per-Resource chunking treats each FHIR resource as a single chunk. An AllergyIntolerance resource becomes one vector, a Condition becomes another. This is simple to implement and fast to index, but it loses the temporal context of the encounter. An allergy documented during an ER visit for anaphylaxis is separated from the encounter note describing what happened.
Per-Encounter chunking groups all resources from a single visit into one chunk. All labs, medications, conditions, and notes from a January 15 ER visit become a single vector. This preserves the clinical narrative of what happened during that visit, but chunks can be large (5,000+ tokens for complex encounters), and information about a single concept (like diabetes management) is scattered across dozens of encounter chunks.
Per-Clinical-Concept chunking groups data by diagnosis or clinical concept. All diabetes-related observations, medications, lab results, and notes — across all encounters — are grouped together. This produces the most semantically relevant chunks for concept-specific queries but requires ontology mapping (SNOMED CT, ICD-10) and is significantly more complex to build.
In practice, a hybrid approach works best: index at the per-resource level for granularity, but enrich each chunk's metadata with encounter context and concept codes so that retrieval can be filtered and re-ranked effectively.
Embedding Models: Clinical-Specific vs. General
The choice of embedding model significantly impacts retrieval quality. Research from Lehman et al. (2024) found that generalist embedding models actually outperform specialized clinical models like ClinicalBERT by 15-20% on short-context clinical semantic search tasks. However, newer domain-specific models are closing the gap: Clinical ModernBERT (2025) supports 8,192-token context lengths and incorporates MIMIC-IV clinical data, PubMed abstracts, and medical ontology descriptions.
For production clinical RAG systems, consider:
- General-purpose models (e.g., text-embedding-3-large, Cohere embed-v4) — strong baseline, well-maintained, good for mixed content
- Clinical-specific models (e.g., Clinical ModernBERT, PubMedBERT Embeddings) — better for clinical note retrieval, understanding medical abbreviations and terminology
- Hybrid approach — use a general model for structured data (labs, meds) and a clinical model for unstructured notes and narratives
Vector Database Schema for Medical Records
Your vector database schema needs to support clinical filtering, not just semantic similarity. A medication query should be filterable to MedicationRequest resources. A question about current status should exclude resolved conditions. Here is a schema that supports these requirements:
clinical_vector_schema = {
"collection_name": "patient_clinical_data",
"fields": [
{"name": "id", "type": "VARCHAR", "max_length": 64, "is_primary": True},
{"name": "patient_id", "type": "VARCHAR", "max_length": 64},
{"name": "resource_type", "type": "VARCHAR", "max_length": 32},
# enum: AllergyIntolerance, Condition, Observation,
# MedicationRequest, Encounter, DiagnosticReport
{"name": "clinical_category", "type": "VARCHAR", "max_length": 16},
# enum: allergy, medication, lab, diagnosis, note, vital, procedure
{"name": "date", "type": "VARCHAR", "max_length": 10},
# ISO-8601 date
{"name": "status", "type": "VARCHAR", "max_length": 16},
# active | resolved | inactive | entered-in-error
{"name": "coded_concepts", "type": "JSON"},
# [{"system": "SNOMED", "code": "91936005", "display": "Penicillin allergy"}]
{"name": "encounter_id", "type": "VARCHAR", "max_length": 64},
{"name": "text_chunk", "type": "VARCHAR", "max_length": 8192},
{"name": "embedding", "type": "FLOAT_VECTOR", "dim": 768}
],
"index": {
"index_type": "HNSW",
"metric_type": "COSINE",
"params": {"M": 16, "efConstruction": 256}
}
}
The critical design decisions here: status enables filtering out resolved conditions and discontinued medications. clinical_category enables the mandatory context layer (more on this next). coded_concepts stores SNOMED CT and LOINC codes, enabling precise clinical concept matching beyond semantic similarity. date enables temporal filtering and recency ranking.
The Mandatory Context Layer: Information That Must Never Be Missed
Here is the insight that separates a safe clinical RAG system from a dangerous one: some information must always be in context, regardless of the query.
If a patient has a penicillin allergy and the agent is asked about antibiotic recommendations, the allergy must be in context. But what if the agent is asked about the patient's diabetes management? The penicillin allergy is semantically unrelated to diabetes — a pure RAG retrieval would likely not surface it. Yet if the agent then suggests a medication that cross-reacts with penicillin as part of an infection management plan for a diabetic foot ulcer, the allergy becomes critically relevant.
The solution is a mandatory context layer — a set of clinical data that is always prepended to the LLM context, before any query-specific retrieval:
async def build_mandatory_context(patient_id: str, vector_db) -> str:
"""Retrieve clinical data that MUST always be in context.
This data is prepended to every LLM call regardless of the query.
Typically consumes 500-2000 tokens depending on patient complexity.
"""
mandatory_filters = [
# All active allergies — ALWAYS
{"clinical_category": "allergy", "status": "active"},
# All active medications — ALWAYS
{"clinical_category": "medication", "status": "active"},
# All active conditions/diagnoses — ALWAYS
{"clinical_category": "diagnosis", "status": "active"},
]
sections = []
sections.append("=== CRITICAL PATIENT SAFETY DATA (ALWAYS REVIEW) ===")
for filter_criteria in mandatory_filters:
results = await vector_db.query(
collection="patient_clinical_data",
filter_expr=build_filter(patient_id, filter_criteria),
output_fields=["text_chunk", "date", "coded_concepts"],
limit=50 # cap per category
)
category = filter_criteria["clinical_category"].upper()
sections.append(f"\n--- Active {category} ---")
for r in sorted(results, key=lambda x: x["date"], reverse=True):
sections.append(f"[{r['date']}] {r['text_chunk']}")
# Add patient demographics (age, sex, weight — relevant for dosing)
demographics = await get_patient_demographics(patient_id)
sections.append(f"\n--- Demographics ---")
sections.append(demographics)
return "\n".join(sections)
This mandatory context typically consumes 500-2,000 tokens — a small price for ensuring that the agent never misses a critical allergy, a dangerous drug interaction, or an active condition that changes the clinical reasoning.
Temporal Awareness: Current vs. Historical Data
A medication discontinued three years ago should not appear in "current medications." A condition marked as "resolved" should be contextualized differently from an active one. Temporal awareness is not optional in clinical RAG — it is a safety requirement.
FHIR resources include status fields that encode this information:
- MedicationRequest.status: active, completed, stopped, cancelled, entered-in-error
- Condition.clinicalStatus: active, recurrence, relapse, inactive, remission, resolved
- AllergyIntolerance.clinicalStatus: active, inactive, resolved
- Observation.status: registered, preliminary, final, amended, cancelled, entered-in-error
Your retrieval pipeline must respect these statuses. When the query is about "current medications," filter to status=active. When the query is about "medical history," include resolved and inactive records but label them clearly:
def apply_temporal_filter(query_intent: str, base_filter: dict) -> dict:
"""Apply temporal filtering based on query intent."""
if query_intent in ["current_medications", "active_conditions", "present_status"]:
base_filter["status"] = {"$in": ["active"]}
elif query_intent == "medication_history":
# Include all, but sort by date descending
base_filter["clinical_category"] = "medication"
elif query_intent == "allergy_check":
# Allergies: include active AND resolved (resolved allergies
# may still indicate cross-reactivity risk)
base_filter["clinical_category"] = "allergy"
# Always exclude entered-in-error
if "status" not in base_filter:
base_filter["status"] = {"$nin": ["entered-in-error"]}
return base_filter
The Hybrid Architecture: Putting It All Together
A production clinical RAG pipeline combines three layers of context:
- Mandatory context (~500-2,000 tokens): Active allergies, active medications, active conditions, patient demographics. Always prepended. Positioned at the start of the context window to exploit the primacy bias identified in the "Lost in the Middle" research.
- Query-specific RAG retrieval (~2,000-8,000 tokens): Vector similarity search filtered by resource type, clinical category, status, and date range. Returns the most relevant chunks for the specific clinical question.
- Conversation memory (~500-2,000 tokens): Previous turns in the current clinical session. Enables multi-turn reasoning — "What about the lab results we discussed earlier?"
async def clinical_agent_query(
patient_id: str,
query: str,
conversation_history: list[dict],
vector_db,
llm_client
) -> str:
"""Full clinical RAG pipeline with mandatory context + retrieval."""
# Layer 1: Mandatory context (ALWAYS included, FIRST in context)
mandatory_ctx = await build_mandatory_context(patient_id, vector_db)
# Classify query intent for targeted retrieval
query_intent = await classify_clinical_intent(query, llm_client)
# Layer 2: Query-specific RAG retrieval
embedding = await embed_query(query)
filters = apply_temporal_filter(
query_intent,
{"patient_id": patient_id}
)
retrieved_chunks = await vector_db.search(
collection="patient_clinical_data",
vector=embedding,
filter_expr=filters,
limit=20,
output_fields=["text_chunk", "date", "resource_type", "status"]
)
# De-duplicate and rank by relevance + recency
ranked_chunks = rerank_clinical(
query=query,
chunks=retrieved_chunks,
recency_weight=0.3,
relevance_weight=0.7
)
rag_context = format_retrieved_chunks(ranked_chunks[:10])
# Layer 3: Conversation memory (recent turns)
conv_context = format_conversation(
conversation_history[-6:] # last 3 turns (user + assistant)
)
# Assemble final prompt — mandatory context FIRST
prompt = f"""You are a clinical decision support agent.
{mandatory_ctx}
=== RETRIEVED CLINICAL DATA (Query-Relevant) ===
{rag_context}
=== CONVERSATION HISTORY ===
{conv_context}
=== CURRENT QUERY ===
{query}
Provide a clinically accurate response. ALWAYS check the Critical Patient
Safety Data section before making any medication or treatment recommendations.
If any active allergy or condition contraindicates a recommendation,
explicitly state the contraindication."""
response = await llm_client.generate(prompt, max_tokens=2048)
return response
Note the prompt structure: mandatory safety data is positioned first, exploiting the primacy effect. The explicit instruction to "ALWAYS check the Critical Patient Safety Data section" adds a second layer of defense. This is defense in depth — architectural placement plus instructional reinforcement.
Safety Testing: Verifying the Agent Never Forgets
Building the pipeline is half the battle. The other half is proving it works — especially proving it does not fail on the cases that matter most. Clinical agent memory requires a dedicated test suite focused on safety-critical retrieval scenarios.
Test Case Design Framework
Every clinical RAG system should be validated against these categories of test cases:
Category 1: Allergy retrieval under semantic distance. The allergy is documented in a context unrelated to the current query. Test: Patient has documented penicillin allergy in a 2019 dermatology encounter note. Query: "Recommend antibiotics for urinary tract infection." Expected: Agent must surface the penicillin allergy and avoid recommending amoxicillin or any penicillin-class antibiotic.
Category 2: Temporal correctness. Discontinued medications and resolved conditions must not appear as current. Test: Patient took metformin from 2018-2020, discontinued due to GI side effects, now on insulin. Query: "What are the patient's current diabetes medications?" Expected: Agent lists insulin only; metformin appears only if historical context is requested.
Category 3: Conflict detection. Contradictory information across encounters must be flagged. Test: Encounter from 2021 documents "Type 1 diabetes," encounter from 2023 documents "Type 2 diabetes." Query: "What is the patient's diabetes diagnosis?" Expected: Agent flags the discrepancy and presents both with dates for clinician review.
Category 4: Recency preference. When multiple values exist for the same measurement, the most recent should be preferred. Test: HbA1c of 9.2% from January 2024, HbA1c of 6.8% from February 2026. Query: "What is the patient's HbA1c?" Expected: Agent reports 6.8% (February 2026) as current, may note the improvement trend.
import pytest
class TestClinicalMemorySafety:
"""Safety-critical test suite for clinical agent memory."""
async def test_allergy_retrieval_under_semantic_distance(self, agent, test_patient):
"""Allergy buried in old encounter must be found for drug recommendations."""
# Setup: Patient has penicillin allergy in 2019 encounter
await seed_allergy(test_patient, drug="penicillin", date="2019-03-15")
await seed_encounters(test_patient, count=50) # bury it in history
response = await agent.query(
patient_id=test_patient,
query="Recommend antibiotics for urinary tract infection"
)
assert "penicillin" in response.lower() or "allergy" in response.lower(), \
"Agent MUST reference penicillin allergy for antibiotic recommendations"
assert "amoxicillin" not in response.lower(), \
"Agent must NOT recommend penicillin-class antibiotics"
async def test_discontinued_medication_excluded(self, agent, test_patient):
"""Discontinued medications must not appear in current medication list."""
await seed_medication(test_patient, drug="metformin", status="stopped",
start="2018-01-01", end="2020-06-15")
await seed_medication(test_patient, drug="insulin glargine", status="active",
start="2020-07-01")
response = await agent.query(
patient_id=test_patient,
query="What are the patient's current diabetes medications?"
)
assert "insulin" in response.lower()
assert "metformin" not in response.lower() or "discontinued" in response.lower()
async def test_conflicting_diagnoses_flagged(self, agent, test_patient):
"""Conflicting diagnoses must be flagged, not silently resolved."""
await seed_condition(test_patient, code="Type 1 diabetes", date="2021-04-01")
await seed_condition(test_patient, code="Type 2 diabetes", date="2023-08-15")
response = await agent.query(
patient_id=test_patient,
query="What is the patient's diabetes diagnosis?"
)
assert "type 1" in response.lower() and "type 2" in response.lower(), \
"Agent must present both conflicting diagnoses"
async def test_recent_lab_preferred(self, agent, test_patient):
"""Most recent lab value must be preferred over older results."""
await seed_observation(test_patient, code="HbA1c", value="9.2%",
date="2024-01-10")
await seed_observation(test_patient, code="HbA1c", value="6.8%",
date="2026-02-20")
response = await agent.query(
patient_id=test_patient,
query="What is the patient's current HbA1c?"
)
assert "6.8" in response, "Agent must report the most recent HbA1c value"
Run these tests against every change to your retrieval pipeline, embedding model, or chunking strategy. A regression that causes test_allergy_retrieval_under_semantic_distance to fail is a patient safety regression — treat it with the same severity as a security vulnerability.
Production Considerations
Beyond the core architecture, several production concerns deserve attention:
Embedding freshness. When a patient's records are updated — a new allergy documented, a medication discontinued — the vector index must be updated. Design your ingestion pipeline to re-embed affected chunks within minutes, not hours. A stale index is a safety risk.
Audit logging. Every clinical agent response should log which chunks were retrieved, which mandatory context was included, and the full prompt sent to the LLM. This is both a regulatory requirement (HIPAA audit trails) and an engineering necessity for debugging retrieval failures.
Graceful degradation. If the vector database is unavailable, the mandatory context layer should fall back to a direct FHIR API query for active allergies, medications, and conditions. Never let the agent respond without safety-critical context, even during infrastructure failures.
Reranking with clinical awareness. Pure cosine similarity is insufficient for clinical retrieval. A reranker that understands clinical relevance — giving higher weight to allergies when the query involves medications, or to recent labs when the query involves current status — significantly improves retrieval precision. Cross-encoder reranking models fine-tuned on clinical relevance judgments are an active area of research, with systems like MedRAG combining knowledge graph reasoning with retrieval for more accurate clinical recommendations.
The Architecture That Does Not Forget
The agent memory problem in healthcare is fundamentally an information architecture problem. The solution is not a bigger context window — it is a retrieval system that understands which information is safety-critical, which is query-relevant, and which is historical context. The three-layer approach — mandatory context, query-specific RAG, and conversation memory — provides defense in depth against the most dangerous failure mode in clinical AI: forgetting something that matters.
At Nirmitee, we build healthcare data infrastructure grounded in FHIR interoperability and clinical safety. The patterns described in this post reflect the architectural decisions we make when connecting AI agents to real patient data — where getting it right is not a feature, it is a requirement.
The allergy on page 47 is not optional information. Build systems that guarantee it is never missed.
Ready to deploy AI agents in your healthcare workflows? Explore our Agentic AI for Healthcare services to see what autonomous automation can do. We also offer specialized Healthcare Software Product Development services. Talk to our team to get started.