Your FHIR server holds millions of clinical resources: patient demographics, lab observations, conditions, encounters, medications. Extracting them one REST call at a time is like draining a swimming pool with a coffee mug. At 100 resources per page, pulling 2 million Patient resources requires 20,000 API calls. Add Observations, Conditions, and MedicationRequests, and you are looking at days of sequential fetching, hammering your production server the entire time.
The FHIR Bulk Data Access specification (STU 3, v3.0.0) solves this with the $export operation: a standardized, asynchronous API for extracting large datasets as NDJSON files. Combined with Mirth Connect as an orchestration engine and a cloud data warehouse as the destination, you can build a production-grade population health data pipeline that moves millions of resources on a nightly schedule.
This guide walks through every layer of that pipeline: the $export operation itself, the async polling pattern, Mirth channel architecture for orchestration, NDJSON processing at scale, and loading into Snowflake, BigQuery, PostgreSQL, or Databricks. We include working code and configuration you can adapt for your environment.
What Is FHIR Bulk Data Export?
The Bulk Data Access Implementation Guide defines a FHIR-native mechanism for exporting large volumes of data from a FHIR server to an authorized client. Instead of paginating through individual REST endpoints, the client kicks off an asynchronous export job and downloads the results as NDJSON (Newline Delimited JSON) files, where each line is a self-contained FHIR resource.
The specification was developed by the HL7 FHIR Infrastructure Work Group and Boston Children's Hospital SMART Health IT team. It is now a required capability under CMS-0057-F, the CMS Interoperability and Prior Authorization Final Rule, which mandates that Medicare Advantage organizations, Medicaid managed care plans, CHIP entities, and Qualified Health Plan issuers support FHIR-based bulk data exchange starting January 1, 2026.
Key characteristics of FHIR Bulk Data Export:
- Asynchronous by design — the client does not wait for the export to complete. It receives a status polling URL and checks back periodically.
- NDJSON output — each output file contains one FHIR resource per line, organized by resource type (Patient.ndjson, Observation.ndjson, Condition.ndjson).
- Incremental exports — the
_sinceparameter enables delta exports, returning only resources modified after a given timestamp. - Resource type filtering — the
_typeparameter restricts the export to specific resource types, reducing export volume by 60-80%. - SMART Backend Services auth — bulk exports use OAuth 2.0 client credentials with a signed JWT assertion, suitable for system-to-system communication without user interaction.
Three Export Levels
The specification defines three scopes for the $export operation, each serving different use cases in population health and data engineering workflows.
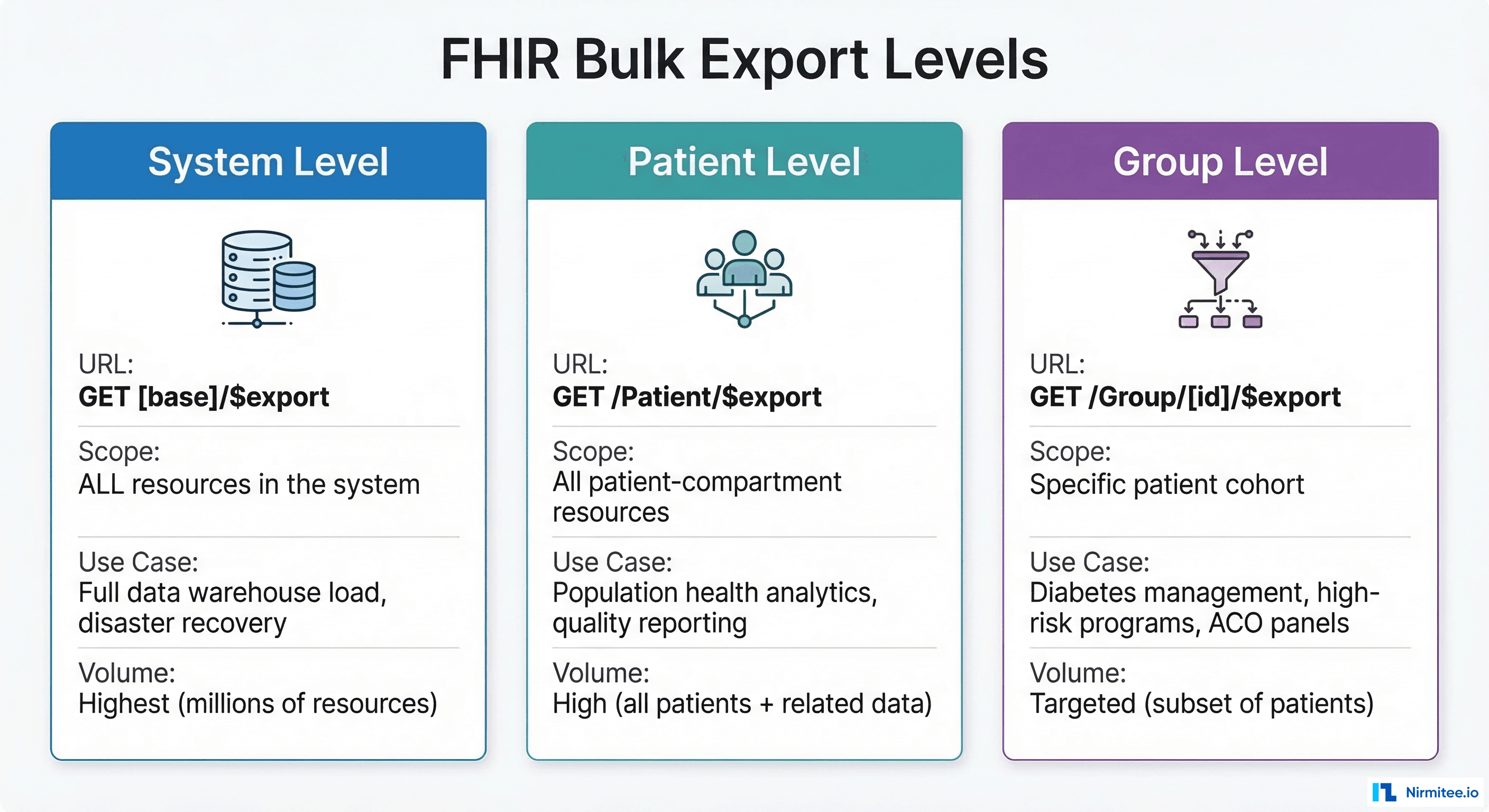
System-Level Export
POST [fhir-base]/$export
Accept: application/fhir+json
Prefer: respond-async
Exports all data in the FHIR server. This is the broadest scope: every Patient, Observation, Encounter, Condition, Procedure, MedicationRequest, and supporting resource. Use it for initial data warehouse loads, disaster recovery snapshots, or regulatory data submissions. A typical health system with 500,000 patients generates 50-200 GB of NDJSON at this level.
Patient-Level Export
POST [fhir-base]/Patient/$export
Accept: application/fhir+json
Prefer: respond-async
Content-Type: application/fhir+json
{"parameter": [{"name": "_type", "valueString": "Patient,Observation,Condition,Encounter"}]}
Exports all resources in the Patient compartment. This is the most common level for population health analytics because it captures the complete clinical picture: demographics, labs, diagnoses, encounters, medications, and procedures tied to patients. It excludes non-patient resources like Organization, Practitioner, and Location unless explicitly requested.
Group-Level Export
POST [fhir-base]/Group/diabetes-cohort/$export
Accept: application/fhir+json
Prefer: respond-async
Exports data for a specific patient cohort defined by a FHIR Group resource. This is the precision tool: export data for your ACO panel (4,200 attributed lives), your diabetes management program (1,800 patients with HbA1c > 7%), or your high-risk pregnancy cohort (340 patients). Group-level exports produce dramatically smaller datasets and complete faster, making them ideal for targeted quality measure reporting and clinical AI/ML pipelines.
The Async Pattern: How $export Actually Works
Unlike standard FHIR REST operations that return results synchronously, $export follows a four-phase asynchronous pattern defined in the FHIR Asynchronous Request Pattern.
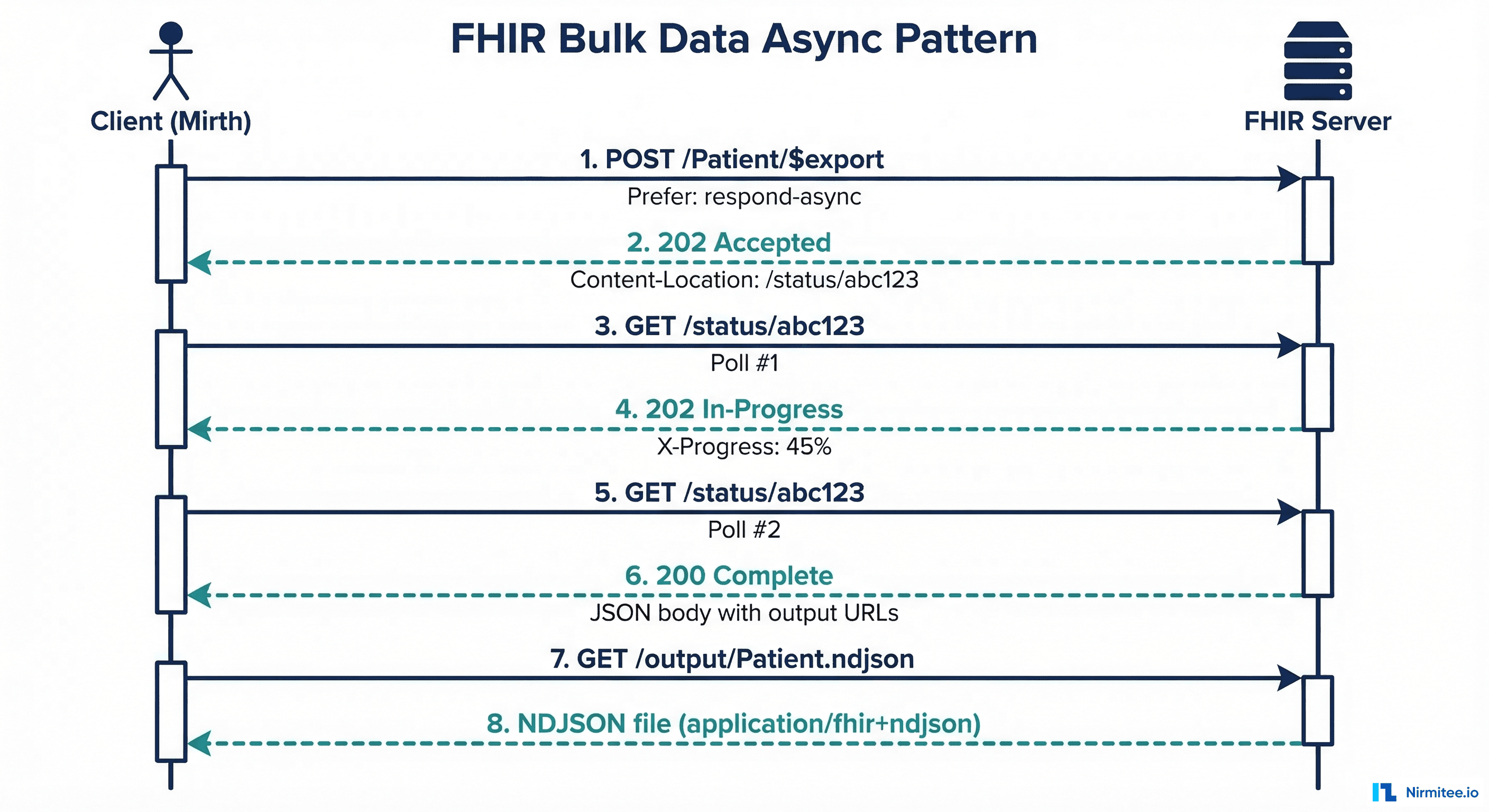

Phase 1: Kick-off
The client sends a POST (or GET) request to the $export endpoint with the Prefer: respond-async header. The server validates the request, creates an export job, and immediately returns 202 Accepted with a Content-Location header pointing to the status polling URL.
HTTP/1.1 202 Accepted
Content-Location: https://fhir.example.org/bulk-status/job-abc-123
The client stores this URL. The export job runs asynchronously on the server, potentially for minutes or hours depending on data volume.
Phase 2: Status Polling
The client periodically polls the status URL with a GET request. While the job is in progress, the server returns 202 Accepted with an optional X-Progress header indicating completion percentage and a Retry-After header suggesting the polling interval.
HTTP/1.1 202 Accepted
X-Progress: Processing: 45% complete
Retry-After: 30
Phase 3: Completion
When the export finishes, the status endpoint returns 200 OK with a JSON body listing the output files, organized by resource type:
{
"transactionTime": "2026-03-16T02:15:00Z",
"request": "https://fhir.example.org/Patient/$export",
"requiresAccessToken": true,
"output": [
{"type": "Patient", "url": "https://fhir.example.org/output/Patient-1.ndjson", "count": 125000},
{"type": "Patient", "url": "https://fhir.example.org/output/Patient-2.ndjson", "count": 125000},
{"type": "Observation", "url": "https://fhir.example.org/output/Observation-1.ndjson", "count": 500000},
{"type": "Condition", "url": "https://fhir.example.org/output/Condition-1.ndjson", "count": 340000}
],
"error": []
}
Note that large resource types may be split across multiple files (Patient-1.ndjson, Patient-2.ndjson). The transactionTime value is critical: store it and use it as the _since parameter for your next incremental export.
Phase 4: Download
The client downloads each NDJSON file from the output URLs. These files can be large (hundreds of MB to several GB), so streaming downloads with proper error handling and retry logic are essential.
Using Mirth Connect as the Orchestrator
Mirth Connect is an ideal orchestrator for bulk export pipelines because it handles HTTP communication, scheduling, JavaScript-based logic, and channel chaining natively. Here is the three-channel architecture that manages the full async lifecycle.
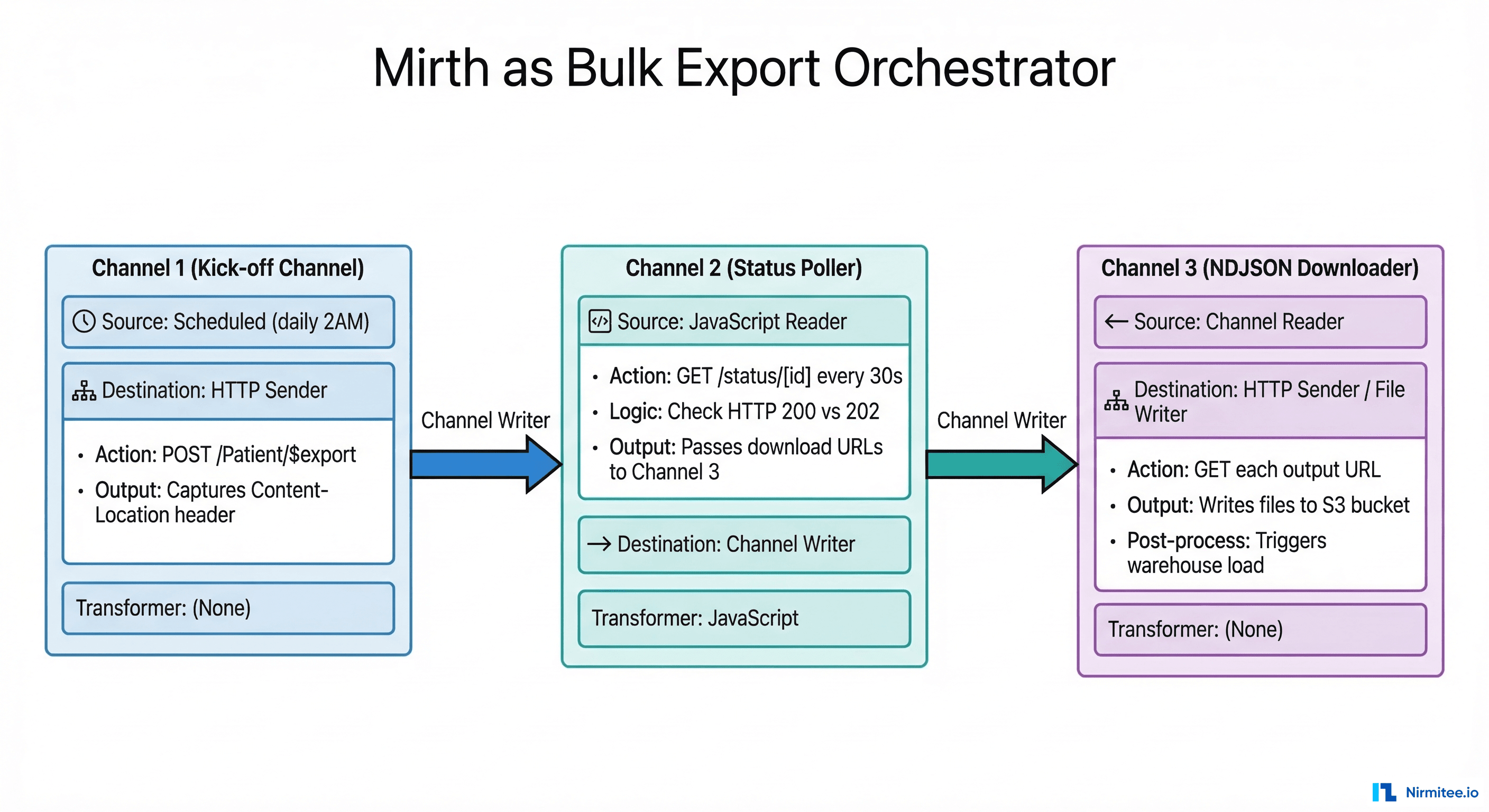
Channel 1: Export Kick-off
This channel runs on a schedule (e.g., daily at 2:00 AM) and initiates the $export request. Configure an HTTP Sender destination with:
- URL:
https://fhir.example.org/Patient/$export - Method: POST
- Headers:
Accept: application/fhir+json,Prefer: respond-async,Authorization: Bearer [token] - Parameters:
_type=Patient,Observation,Condition,Encounter,MedicationRequest,_since=[last_export_timestamp]
In the response transformer, extract the Content-Location header and persist it to a channel map or database table:
// Response transformer — extract Content-Location
var statusCode = $('responseStatusCode');
if (statusCode == 202) {
var contentLocation = $('responseHeaders').getHeader('Content-Location');
var dbConn = DatabaseConnectionFactory.createDatabaseConnection(
'org.postgresql.Driver',
'jdbc:postgresql://localhost:5432/mirth_bulk',
'mirth_user', 'mirth_pass'
);
dbConn.executeUpdate(
"INSERT INTO bulk_export_jobs (job_id, status_url, started_at, status) " +
"VALUES (gen_random_uuid(), '" + contentLocation + "', NOW(), 'polling')"
);
dbConn.close();
logger.info('Bulk export kicked off. Status URL: ' + contentLocation);
} else {
logger.error('Unexpected status: ' + statusCode);
}
Channel 2: Status Poller
This channel uses a JavaScript Reader source that runs on a timer (every 30-60 seconds). It queries the database for active polling jobs and checks each status URL:
// JavaScript Reader source — poll active export jobs
var dbConn = DatabaseConnectionFactory.createDatabaseConnection(
'org.postgresql.Driver',
'jdbc:postgresql://localhost:5432/mirth_bulk',
'mirth_user', 'mirth_pass'
);
var result = dbConn.executeCachedQuery(
"SELECT id, status_url FROM bulk_export_jobs WHERE status = 'polling'"
);
var messages = new java.util.ArrayList();
while (result.next()) {
var jobId = result.getString('id');
var statusUrl = result.getString('status_url');
// HTTP GET to status URL
var httpClient = new org.apache.http.impl.client.HttpClients.createDefault();
var httpGet = new org.apache.http.client.methods.HttpGet(statusUrl);
httpGet.setHeader('Authorization', 'Bearer ' + getAccessToken());
var response = httpClient.execute(httpGet);
var statusCode = response.getStatusLine().getStatusCode();
if (statusCode == 200) {
// Export complete — parse output URLs
var body = org.apache.commons.io.IOUtils.toString(
response.getEntity().getContent(), 'UTF-8'
);
dbConn.executeUpdate(
"UPDATE bulk_export_jobs SET status='complete', " +
"output_manifest='" + body.replace(/'/g, "''") + "', " +
"completed_at=NOW() WHERE id='" + jobId + "'"
);
// Create message for downstream channel
messages.add(body);
logger.info('Export job ' + jobId + ' complete.');
} else if (statusCode == 202) {
var progress = response.getFirstHeader('X-Progress');
logger.info('Job ' + jobId + ': ' + (progress ? progress.getValue() : 'in progress'));
} else {
logger.error('Job ' + jobId + ' unexpected status: ' + statusCode);
}
httpClient.close();
}
dbConn.close();
return messages;
Channel 3: NDJSON Downloader
When Channel 2 detects a completed export, it sends the manifest JSON to Channel 3 via a Channel Writer. Channel 3 parses the output array and downloads each NDJSON file to cloud storage:
// Source transformer — parse manifest and download each file
var manifest = JSON.parse(msg);
var outputFiles = manifest.output;
for (var i = 0; i < outputFiles.length; i++) {
var fileEntry = outputFiles[i];
var resourceType = fileEntry.type;
var downloadUrl = fileEntry.url;
var count = fileEntry.count || 'unknown';
logger.info('Downloading ' + resourceType + ' (' + count + ' resources) from: ' + downloadUrl);
// Stream download to local staging directory
var httpClient = org.apache.http.impl.client.HttpClients.createDefault();
var httpGet = new org.apache.http.client.methods.HttpGet(downloadUrl);
httpGet.setHeader('Authorization', 'Bearer ' + getAccessToken());
httpGet.setHeader('Accept', 'application/fhir+ndjson');
var response = httpClient.execute(httpGet);
var timestamp = new java.text.SimpleDateFormat('yyyyMMdd_HHmmss').format(new java.util.Date());
var filename = resourceType + '_' + timestamp + '_' + i + '.ndjson';
var outputPath = '/data/bulk-export/staging/' + filename;
var inputStream = response.getEntity().getContent();
var outputStream = new java.io.FileOutputStream(outputPath);
org.apache.commons.io.IOUtils.copy(inputStream, outputStream);
outputStream.close();
httpClient.close();
logger.info('Saved: ' + outputPath + ' (' + new java.io.File(outputPath).length() + ' bytes)');
// Upload to S3 (via AWS CLI or SDK)
var s3Key = 'fhir-bulk/' + timestamp + '/' + filename;
java.lang.Runtime.getRuntime().exec(
'aws s3 cp ' + outputPath + ' s3://your-data-lake/' + s3Key
);
}
NDJSON Processing: Parsing at Scale
NDJSON (Newline Delimited JSON) is the output format for all FHIR bulk exports. Each line is a complete, valid JSON object representing a single FHIR resource. A 500,000-resource Observation file might be 2-4 GB. The cardinal rule: never load the entire file into memory.
Stream Processing Pattern
Process NDJSON files line by line using streaming I/O. In Java (within Mirth), Python, or Node.js, the pattern is the same: read one line, parse it, process it, discard it.
# Python streaming NDJSON parser with resource type splitting
import json
import os
from collections import defaultdict
def process_ndjson_file(input_path, output_dir):
"""Split a mixed NDJSON file into per-resource-type files."""
writers = {}
counts = defaultdict(int)
with open(input_path, 'r') as f:
for line_num, line in enumerate(f, 1):
line = line.strip()
if not line:
continue
try:
resource = json.loads(line)
resource_type = resource.get('resourceType', 'Unknown')
# Get or create writer for this resource type
if resource_type not in writers:
out_path = os.path.join(output_dir, f'{resource_type}.ndjson')
writers[resource_type] = open(out_path, 'a')
writers[resource_type].write(line + '\n')
counts[resource_type] += 1
except json.JSONDecodeError as e:
print(f'Line {line_num}: JSON parse error — {e}')
for w in writers.values():
w.close()
return dict(counts)
# Usage
result = process_ndjson_file('/data/staging/export.ndjson', '/data/processed/')
# Output: {'Patient': 125000, 'Observation': 487000, 'Condition': 92000}
Memory usage stays constant regardless of file size: roughly 50 MB for the Python process, compared to 8+ GB if you loaded the entire file with json.load(). For files exceeding 10 GB, consider using ijson for incremental JSON parsing or orjson for 3-5x faster deserialization.
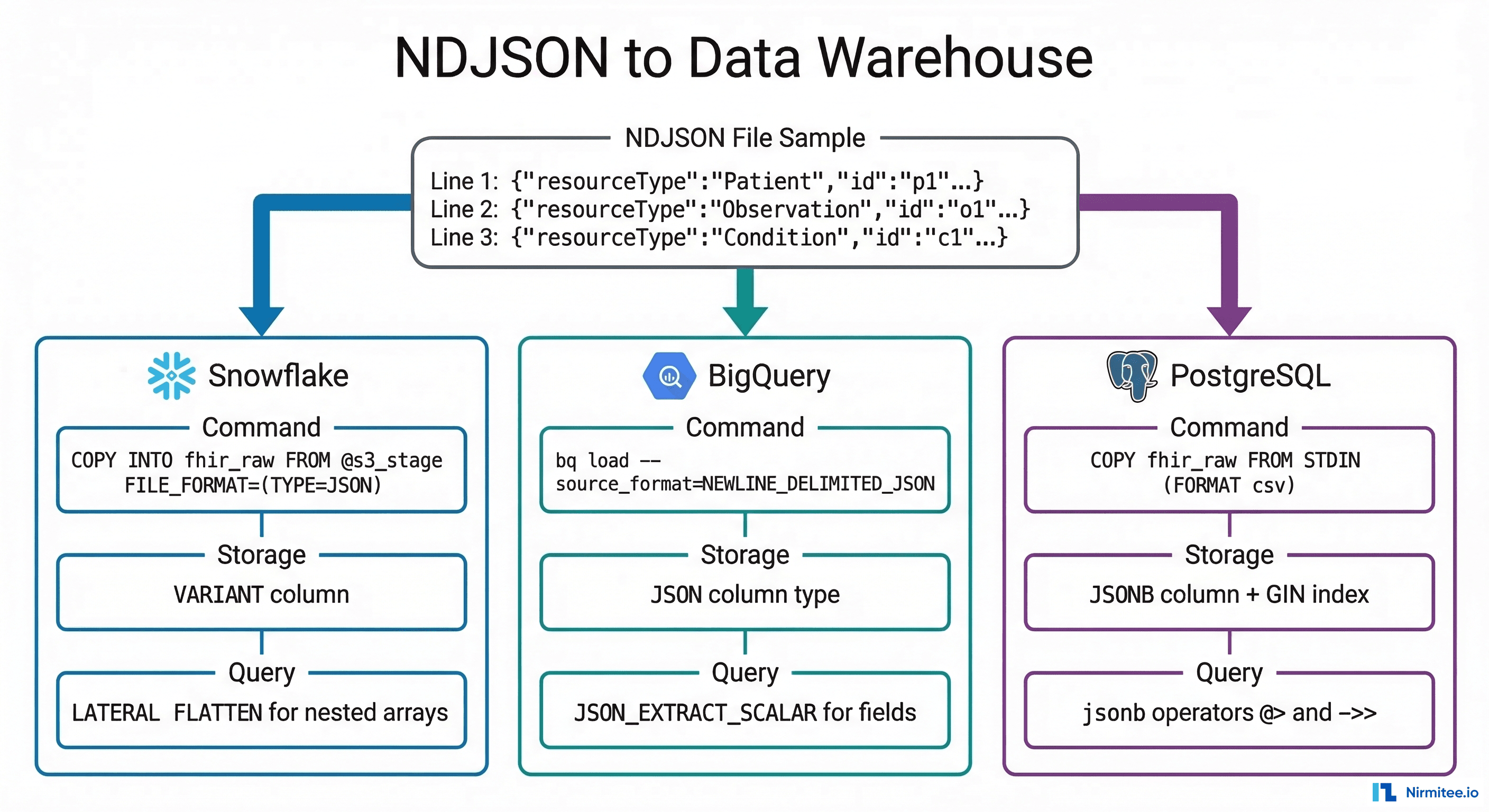
Loading NDJSON into Data Warehouses
Once NDJSON files are staged in cloud storage (S3, GCS, or Azure Blob), the next step is loading them into your analytics warehouse. Each platform has a native, high-performance path for JSON ingestion.
Snowflake: VARIANT Column + COPY INTO
Snowflake's VARIANT data type stores semi-structured JSON natively, and COPY INTO ingests NDJSON files directly from an S3 stage:
-- Create raw staging table
CREATE TABLE fhir_raw.patient_raw (
load_id VARCHAR DEFAULT UUID_STRING(),
loaded_at TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP(),
resource_type VARCHAR,
resource VARIANT
);
-- Create external stage pointing to S3
CREATE OR REPLACE STAGE fhir_s3_stage
URL = 's3://your-data-lake/fhir-bulk/'
CREDENTIALS = (AWS_KEY_ID='...' AWS_SECRET_KEY='...')
FILE_FORMAT = (TYPE=JSON STRIP_OUTER_ARRAY=FALSE);
-- Load NDJSON into raw table
COPY INTO fhir_raw.patient_raw (resource_type, resource)
FROM (
SELECT 'Patient', $1
FROM @fhir_s3_stage/Patient.ndjson
);
-- Query: Extract structured patient data
SELECT
resource:id::VARCHAR AS patient_id,
resource:name[0].family::VARCHAR AS last_name,
resource:name[0].given[0]::VARCHAR AS first_name,
resource:birthDate::DATE AS birth_date,
resource:gender::VARCHAR AS gender,
f.value:coding[0].code::VARCHAR AS condition_code
FROM fhir_raw.patient_raw p,
LATERAL FLATTEN(input => p.resource:extension) f
WHERE resource:resourceType = 'Patient';
BigQuery: Load Job with JSON Column
BigQuery supports native NDJSON ingestion from Google Cloud Storage with automatic schema detection or a JSON column type:
# Load NDJSON into BigQuery with JSON column
bq load \
--source_format=NEWLINE_DELIMITED_JSON \
--autodetect \
your_project:fhir_warehouse.patient_raw \
gs://your-bucket/fhir-bulk/Patient.ndjson
# Or using the Healthcare API FHIR store export
gcloud healthcare fhir-stores export bq \
--dataset=your-healthcare-dataset \
--fhir-store=your-fhir-store \
--bq-dataset=bq://your_project.fhir_warehouse \
--resource-type=Patient,Observation,Condition
-- BigQuery: Query FHIR resources
SELECT
JSON_EXTRACT_SCALAR(resource, '$.id') AS patient_id,
JSON_EXTRACT_SCALAR(resource, '$.name[0].family') AS last_name,
JSON_EXTRACT_SCALAR(resource, '$.birthDate') AS birth_date,
JSON_EXTRACT_SCALAR(resource, '$.gender') AS gender
FROM `your_project.fhir_warehouse.patient_raw`;
PostgreSQL: JSONB + GIN Index
For teams already running PostgreSQL in their healthcare stack, JSONB columns with GIN indexes provide fast semi-structured querying without a separate warehouse:
-- Create raw FHIR resource table
CREATE TABLE fhir_raw (
id UUID DEFAULT gen_random_uuid() PRIMARY KEY,
resource_type VARCHAR(64) NOT NULL,
resource_id VARCHAR(128),
resource JSONB NOT NULL,
loaded_at TIMESTAMPTZ DEFAULT NOW()
);
-- GIN index for fast JSONB queries
CREATE INDEX idx_fhir_raw_resource ON fhir_raw USING GIN (resource);
CREATE INDEX idx_fhir_raw_type ON fhir_raw (resource_type);
-- Load NDJSON using psql
\COPY fhir_raw (resource_type, resource_id, resource)
FROM PROGRAM 'cat Patient.ndjson | jq -r ''[.resourceType, .id, (. | tojson)] | @csv''' WITH CSV;
-- Query: Find diabetic patients
SELECT resource->>'id' AS patient_id,
resource->'name'->0->>'family' AS last_name
FROM fhir_raw
WHERE resource_type = 'Condition'
AND resource @> '{"code": {"coding": [{"system": "http://snomed.info/sct", "code": "73211009"}]}}';
Databricks: Delta Lake
For teams on the Lakehouse architecture, load NDJSON into Delta tables for unified batch and streaming analytics:
# PySpark: Load NDJSON into Delta Lake
df = spark.read.json("s3://your-data-lake/fhir-bulk/Patient.ndjson")
df.write.format("delta").mode("append").saveAsTable("fhir.patient_raw")
# Query with SQL
SELECT id, name[0].family AS last_name, birthDate, gender
FROM fhir.patient_raw
WHERE resourceType = 'Patient'
CMS Bulk FHIR Requirements: Why This Matters Now
The CMS Interoperability and Prior Authorization Final Rule (CMS-0057-F) makes bulk FHIR data exchange a regulatory requirement, not just a technical convenience. As of January 1, 2026, impacted payers must support:
- Provider Access API — payers must make claims, encounter data, USCDI elements, and prior authorization information available to in-network providers via FHIR APIs. Bulk Data Access is the expected mechanism for large-scale data retrieval.
- Payer-to-Payer API (January 2027) — when a member switches plans, the new payer can request up to 5 years of clinical and claims history via bulk FHIR exchange.
- Prior Authorization API — payers must support FHIR-based prior auth with bulk reporting of metrics starting March 31, 2026.
For provider organizations building CMS interoperability compliance, this means your integration engine needs to consume bulk FHIR exports from multiple payers and load them into your analytics platform. The Mirth-based pipeline described in this guide is exactly that infrastructure.
Production Architecture: End-to-End Pipeline
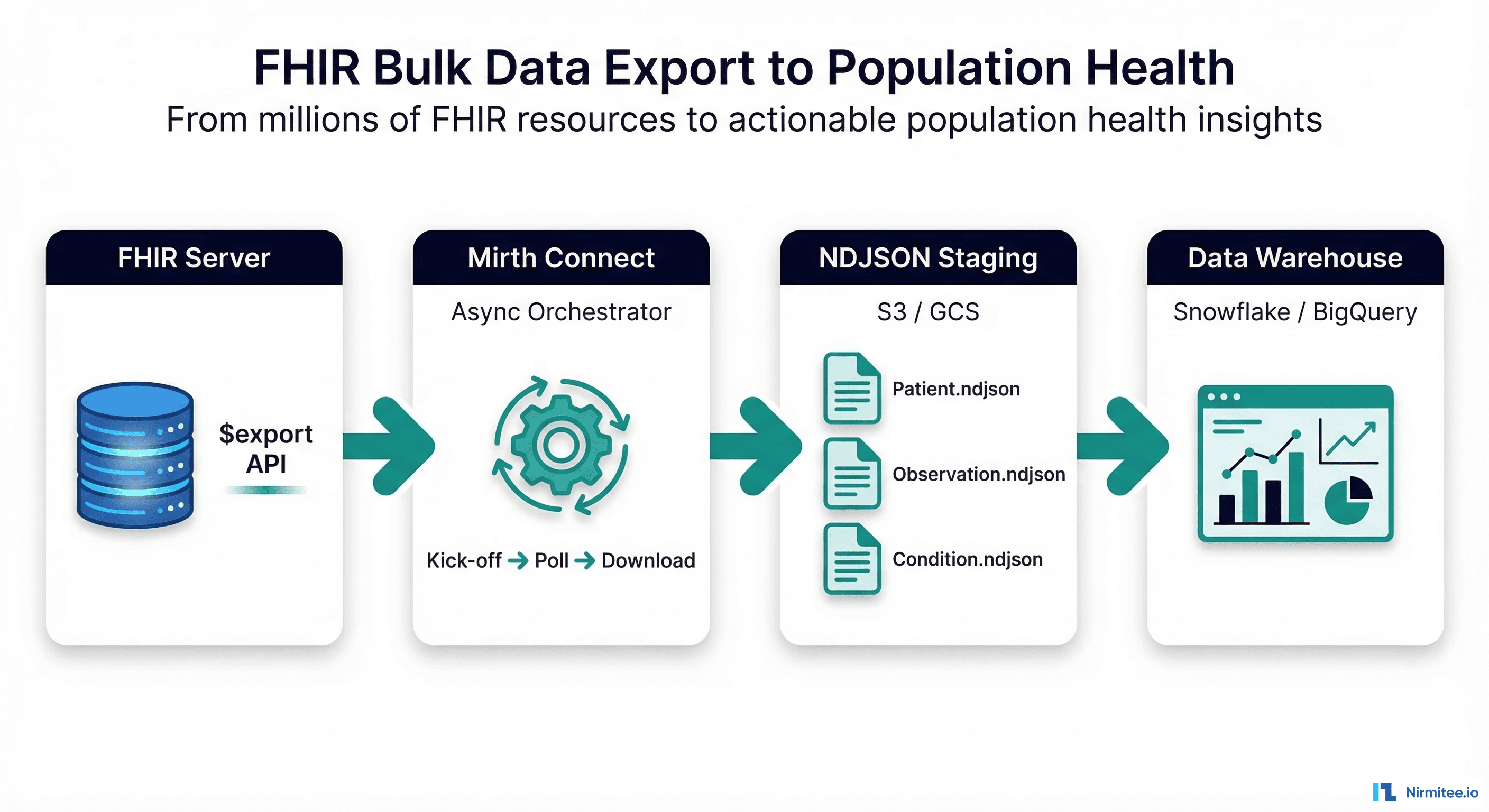
The complete production architecture connects five layers:
- FHIR Server (HAPI FHIR, Microsoft FHIR Server, Google Healthcare API, Smile CDR) — hosts the clinical data and exposes the
$exportendpoint. - Mirth Connect — orchestrates the async export lifecycle: kick-off, polling, downloading. Three channels handle the workflow as described above. Deployed in Docker on AWS/Azure alongside the FHIR server.
- Cloud Object Storage (S3, GCS, Azure Blob) — NDJSON files are staged here before warehouse loading. Lifecycle policies auto-delete files after 30 days. Versioning enabled for audit trail.
- Data Warehouse (Snowflake, BigQuery, Databricks, PostgreSQL) — raw NDJSON loaded into semi-structured columns, then transformed into star-schema tables via dbt or SQL views for analytics.
- Analytics Layer — population health dashboards (Tableau, Looker, Power BI), quality measure calculators (HEDIS, CQMs), and ML model training pipelines consume the structured data.
Performance: Handling Millions of Resources
A 500,000-patient health system generates roughly 15-25 million FHIR resources across core clinical types. Exporting, downloading, and loading this volume requires deliberate performance engineering.
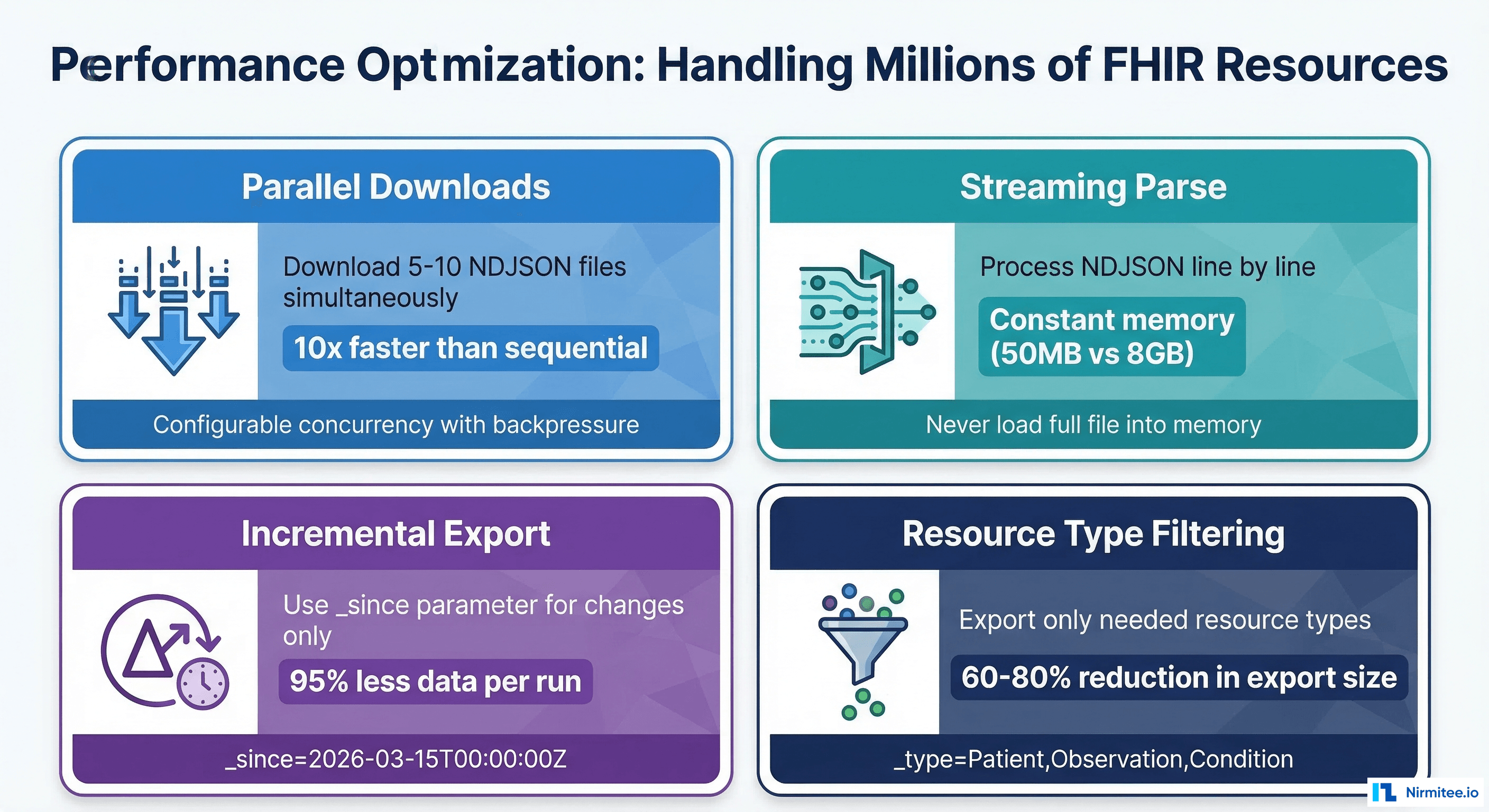

Parallel Downloads
The completion manifest typically contains 10-50 output files. Download them in parallel with configurable concurrency (5-10 simultaneous downloads). In Mirth, use a destination set with multiple HTTP Sender destinations or spawn threads in a JavaScript transformer. This reduces download time from 45 minutes (sequential) to 5-8 minutes (10x parallel).
Incremental Exports with _since
After the initial full export, subsequent runs should use the _since parameter set to the transactionTime from the previous export. This typically reduces export volume by 90-95% because only modified resources are included.
POST /Patient/$export?_since=2026-03-15T02:15:00Z&_type=Patient,Observation,Condition
Accept: application/fhir+json
Prefer: respond-async
Resource Type Filtering
If your population health dashboard only needs Patient, Observation, Condition, Encounter, and MedicationRequest, do not export all 100+ FHIR resource types. The _type parameter reduces export size by 60-80% and proportionally reduces download and load times.
Streaming and Backpressure
For NDJSON files exceeding 1 GB, stream downloads directly to disk or object storage. Never buffer the full response body in memory. In Mirth's HTTP Sender, configure the response handler to stream to a file destination. Monitor Mirth heap usage and set appropriate JVM memory limits (4-8 GB for bulk export channels).
From Pipeline to Population Health Insights
Once FHIR resources land in your data warehouse, the population health use cases unlock:
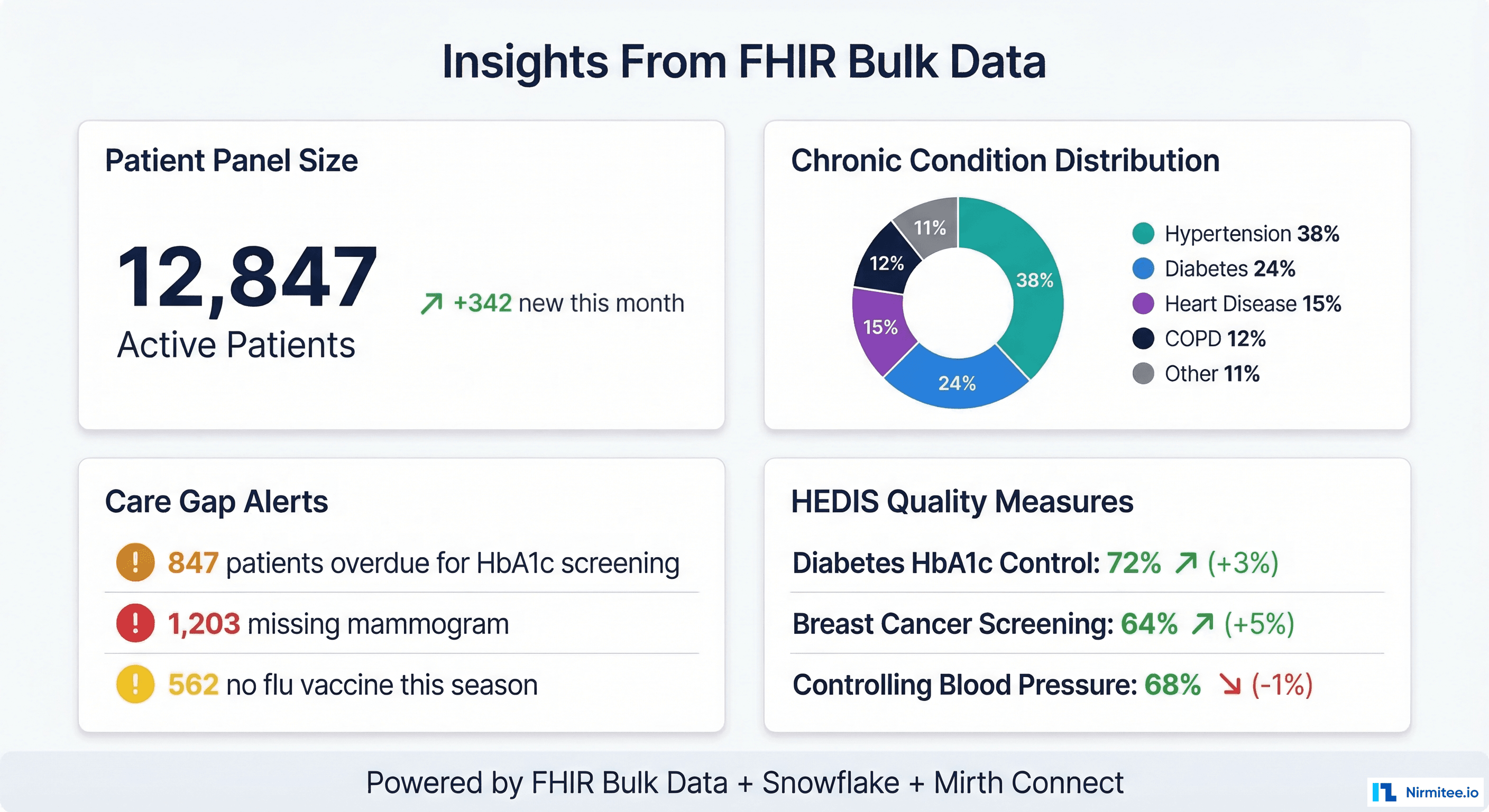
- Care gap identification — query Observation and Condition resources to find patients overdue for HbA1c screening, mammograms, colonoscopies, or annual wellness visits. Cross-reference with Encounter data to identify patients who have not been seen in 12+ months.
- HEDIS quality measure calculation — compute Comprehensive Diabetes Care (CDC), Controlling Blood Pressure (CBP), Breast Cancer Screening (BCS), and other HEDIS measures directly from FHIR resources using SQL.
- Risk stratification — combine Condition, Observation (labs), MedicationRequest, and Encounter data to calculate risk scores and identify rising-risk patients before they become high-cost.
- Chronic disease cohort management — build and maintain patient registries for diabetes, hypertension, heart failure, and COPD programs using Group-level exports and condition-based queries.
- Network adequacy reporting — analyze encounter patterns, referral flows, and geographic coverage using patient and provider data from bulk exports.
Getting Started: Implementation Checklist
If you are building a FHIR Bulk Data pipeline for the first time, here is the practical sequence:
- Verify $export support — check your FHIR server's CapabilityStatement for
$exportoperation support. HAPI FHIR (v6.4+), Smile CDR, Microsoft FHIR Server, and Google Healthcare API all support it. - Configure SMART Backend Services auth — register a client with the FHIR server, generate an RS256 key pair, and implement the SMART Backend Services client credentials flow in Mirth.
- Build the three Mirth channels — kick-off, poller, downloader. Start with a simple Group-level export of 100 patients before scaling to full Patient-level exports.
- Stage to cloud storage — configure S3 or GCS bucket with appropriate IAM roles, encryption at rest, and lifecycle policies.
- Load into your warehouse — start with raw VARIANT/JSONB tables. Build materialized views or dbt models to flatten FHIR resources into analytics-friendly star schemas.
- Schedule and monitor — set up nightly scheduling with alerting on export failures, download errors, and load anomalies.
At Nirmitee, we build FHIR integration pipelines and Mirth Connect architectures for health systems and digital health companies. If you are implementing bulk data export for population health analytics, CMS compliance, or clinical AI pipelines, our team can help you get from zero to production. Get in touch.
Shipping healthcare software that scales requires deep domain expertise. See how our Healthcare Software Product Development practice can accelerate your roadmap. We also offer specialized Healthcare Interoperability Solutions services. Talk to our team to get started.