Your FHIR Data Parses Fine — But Is It Clinically Meaningful?
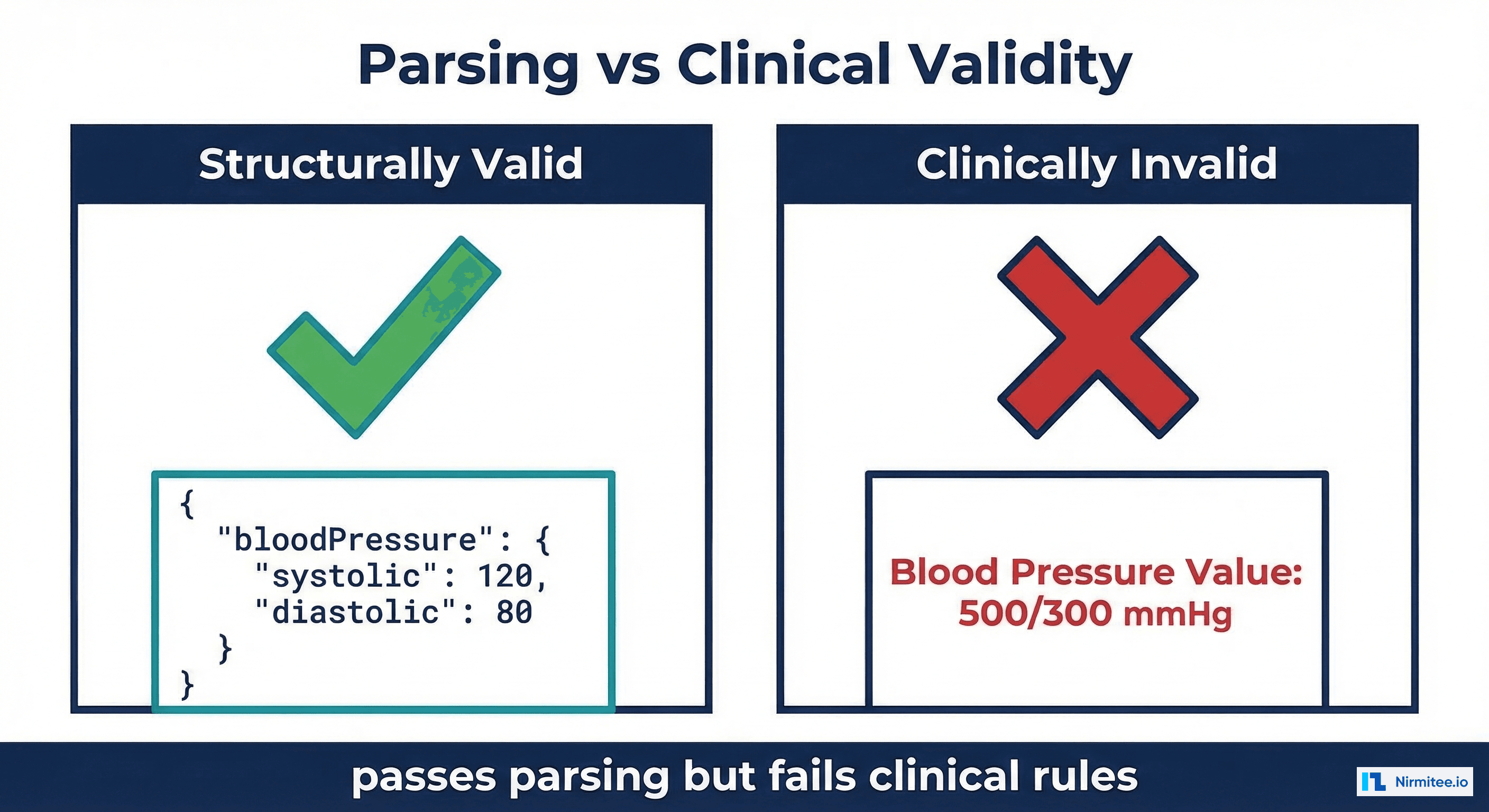
A blood pressure reading of 500/300 mmHg passes every structural validation check. It's valid JSON. The resource type is correct. The required fields are present. The LOINC codes are real. The units are from UCUM. According to the FHIR specification, it's a perfectly valid Observation.
It's also clinically impossible. No human has ever had a blood pressure of 500/300. If this data enters your EHR and triggers a clinical decision support rule, the consequences could range from a false alarm to a dangerous intervention. Yet most FHIR implementations validate only the first two layers (structural and profile conformance) and skip clinical validation entirely.
According to a study published in JAMIA, data quality issues in EHRs affect 10-25% of clinical records. These aren't syntax errors — they're semantically invalid values that passed every automated check but fail the "does this make medical sense?" test. This guide covers all three validation layers and shows you how to build a pipeline that catches what parsers miss.

Layer 1: Structural Validation
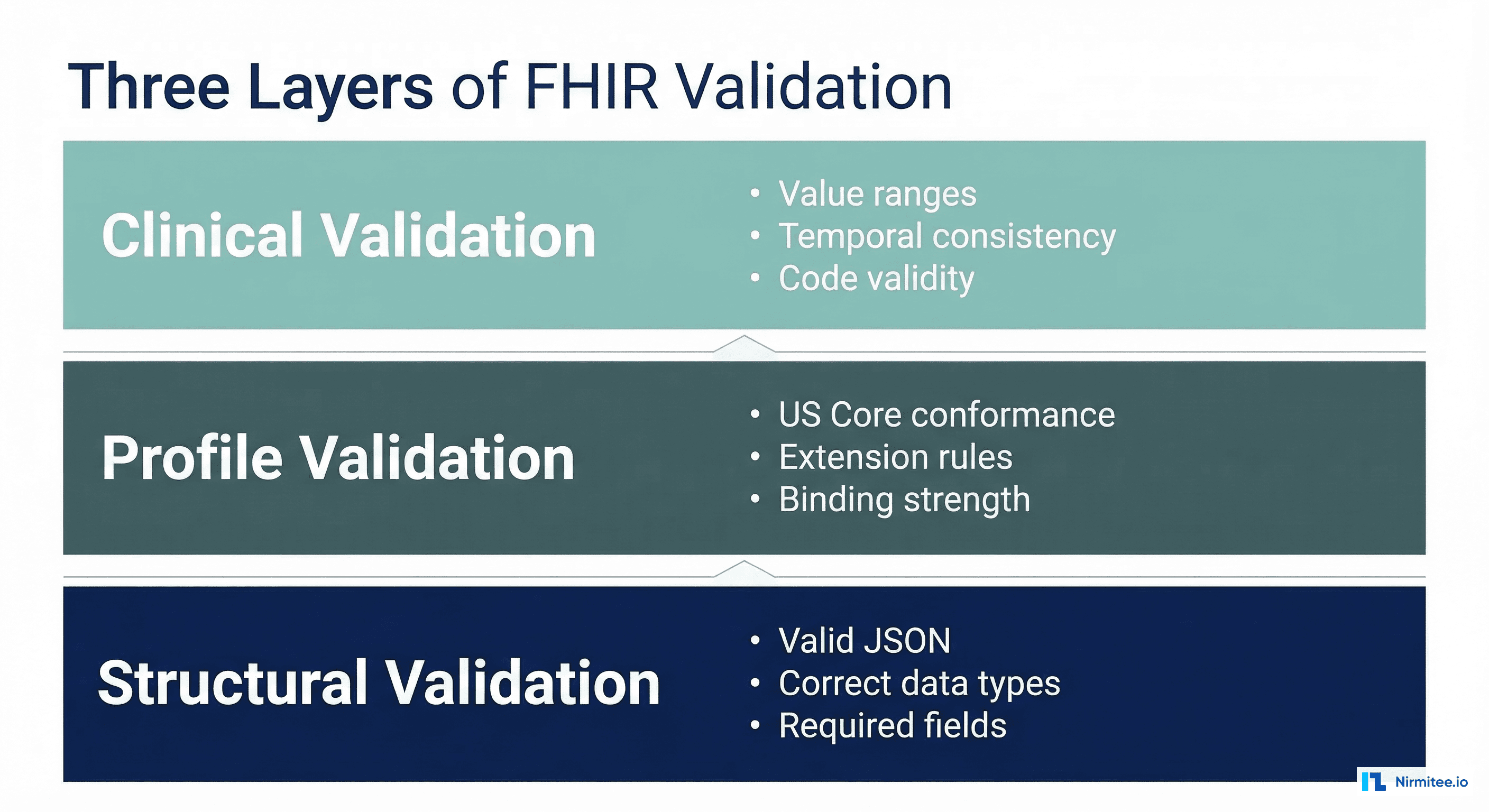
Structural validation confirms that the data conforms to the base FHIR specification. This is the minimum bar — if data fails structural validation, it's not FHIR.
What Structural Validation Checks
| Check | What It Validates | Example Failure |
|---|---|---|
| JSON/XML well-formedness | Valid JSON syntax, proper encoding | Missing closing brace, invalid UTF-8 |
| Resource type | resourceType is a known FHIR type | "resourceType": "Pateint" (typo) |
| Required elements | Elements with min cardinality > 0 exist | Observation without status or code |
| Data types | Values match expected FHIR types | String where integer expected, invalid dateTime format |
| Cardinality | Arrays don't exceed max; single values aren't arrays | "status": ["active", "inactive"] for a 0..1 element |
| Value sets (required binding) | Coded values from required value sets | Observation.status = "pending" (not in required ValueSet) |
| Reference targets | References point to valid resource types | Observation.subject referencing a Medication |
Implementation: HAPI FHIR Validator
The HAPI FHIR Validator is the most widely used structural validator in the Java ecosystem:
// Java: HAPI FHIR structural validation
import ca.uhn.fhir.context.FhirContext;
import ca.uhn.fhir.validation.FhirValidator;
import ca.uhn.fhir.validation.ValidationResult;
import ca.uhn.fhir.validation.SingleValidationMessage;
FhirContext ctx = FhirContext.forR4();
FhirValidator validator = ctx.newValidator();
// Parse and validate
String observationJson = "{ ... }";
ValidationResult result = validator.validateWithResult(observationJson);
if (result.isSuccessful()) {
System.out.println("Structural validation passed");
} else {
for (SingleValidationMessage msg : result.getMessages()) {
System.out.println(msg.getSeverity() + ": " + msg.getMessage()
+ " at " + msg.getLocationString());
}
}Layer 2: Profile Validation
Profile validation checks that data conforms to a specific Implementation Guide (IG) — most commonly US Core. A resource can be structurally valid FHIR but violate profile constraints required for regulatory compliance.

What Profile Validation Checks
| Check | What It Validates | Example Failure |
|---|---|---|
| Must-support elements | Required profile elements are present | US Core Patient without name, gender, or identifier |
| Slicing constraints | Array elements match required slices | US Core Vital Signs without the required LOINC code slice |
| Extension requirements | Required extensions are present | US Core Patient missing race/ethnicity extensions |
| Binding strength | Coded values match extensible/preferred bindings | Using a local code instead of standard SNOMED for Condition.code |
| Fixed values | Elements with fixed values match exactly | US Core Blood Pressure with wrong category code |
| Invariants (FHIRPath) | Cross-element constraints via FHIRPath expressions | Blood pressure without both systolic and diastolic components |
US Core Profile Validation Example
// Java: US Core profile validation with HAPI
import ca.uhn.fhir.context.support.DefaultProfileValidationSupport;
import org.hl7.fhir.common.hapi.validation.support.*;
import org.hl7.fhir.common.hapi.validation.validator.FhirInstanceValidator;
// Build validation support chain
ValidationSupportChain chain = new ValidationSupportChain(
new DefaultProfileValidationSupport(ctx),
new InMemoryTerminologyServerValidationSupport(ctx),
new SnapshotGeneratingValidationSupport(ctx),
// Load US Core IG package
new NpmPackageValidationSupport(ctx, "hl7.fhir.us.core", "6.1.0")
);
FhirInstanceValidator instanceValidator = new FhirInstanceValidator(chain);
validator.registerValidatorModule(instanceValidator);
// Validate against US Core Patient profile
ValidationResult result = validator.validateWithResult(
patientResource,
new ValidationOptions().addProfile(
"http://hl7.org/fhir/us/core/StructureDefinition/us-core-patient"
)
);Layer 3: Clinical Validation
This is where most implementations fall short. Clinical validation checks whether data is medically plausible — not just technically correct.
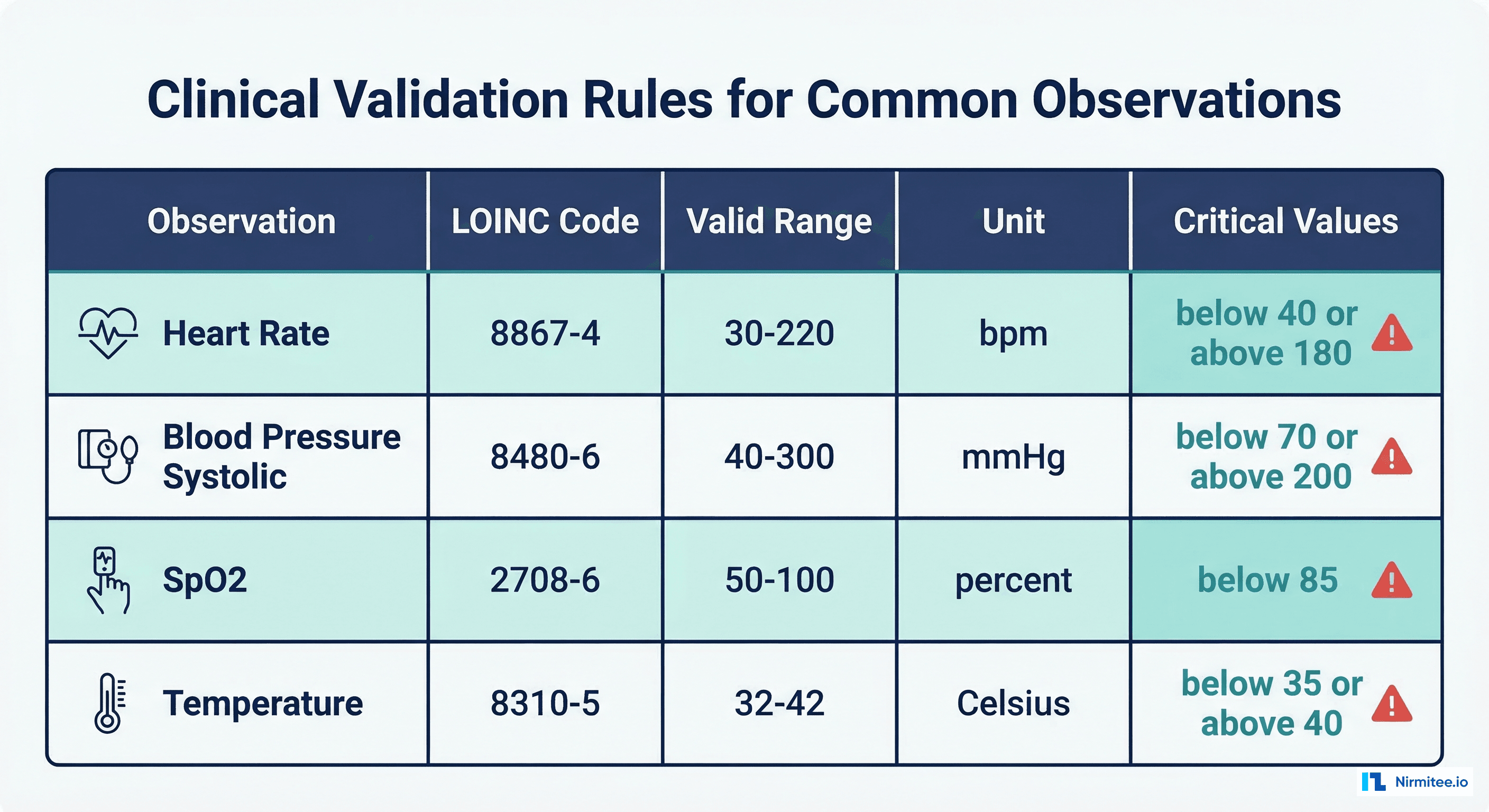
Clinical Validation Categories
- Value range checks — Is the numeric value within a physiologically possible range?
- Temporal consistency — Are dates logically ordered? (admission before discharge, birth before death)
- Code validity — Is the diagnosis code appropriate for the patient's demographics? (pediatric diagnosis on 80-year-old)
- Cross-resource consistency — Does the medication match the condition? Does the procedure align with the diagnosis?
- Unit consistency — Are units correct for the observation type? (temperature in mmHg is wrong)
Python Clinical Validation Framework
"""Clinical validation rules for FHIR Observations."""
from dataclasses import dataclass
from typing import Optional
from datetime import datetime
@dataclass
class ValidationRule:
name: str
severity: str # "error", "warning", "info"
message: str
@dataclass
class ValidationResult:
passed: bool
rule: str
severity: str
message: str
path: str = ""
# Clinical value ranges for common vital signs
VITAL_SIGN_RANGES = {
"8867-4": { # Heart Rate
"name": "Heart Rate",
"unit": "/min",
"absolute_min": 20,
"absolute_max": 300,
"normal_min": 60,
"normal_max": 100,
"critical_low": 40,
"critical_high": 180,
},
"8480-6": { # Systolic BP
"name": "Systolic Blood Pressure",
"unit": "mm[Hg]",
"absolute_min": 30,
"absolute_max": 350,
"normal_min": 90,
"normal_max": 140,
"critical_low": 70,
"critical_high": 200,
},
"8462-4": { # Diastolic BP
"name": "Diastolic Blood Pressure",
"unit": "mm[Hg]",
"absolute_min": 20,
"absolute_max": 200,
"normal_min": 60,
"normal_max": 90,
"critical_low": 40,
"critical_high": 120,
},
"2708-6": { # SpO2
"name": "Oxygen Saturation",
"unit": "%",
"absolute_min": 0,
"absolute_max": 100,
"normal_min": 95,
"normal_max": 100,
"critical_low": 85,
"critical_high": None,
},
"8310-5": { # Body Temperature
"name": "Body Temperature",
"unit": "Cel",
"absolute_min": 25,
"absolute_max": 45,
"normal_min": 36.1,
"normal_max": 37.2,
"critical_low": 32,
"critical_high": 42,
},
"29463-7": { # Body Weight
"name": "Body Weight",
"unit": "kg",
"absolute_min": 0.3,
"absolute_max": 700,
"normal_min": 2.5,
"normal_max": 300,
"critical_low": None,
"critical_high": None,
},
"8302-2": { # Body Height
"name": "Body Height",
"unit": "cm",
"absolute_min": 20,
"absolute_max": 280,
"normal_min": 45,
"normal_max": 220,
"critical_low": None,
"critical_high": None,
},
}
class ClinicalValidator:
def __init__(self):
self.results = []
def validate_observation(self, observation: dict) -> list:
"""Validate an Observation resource for clinical plausibility."""
self.results = []
# Check 1: Value range for vital signs
self._check_value_range(observation)
# Check 2: Unit consistency
self._check_unit_consistency(observation)
# Check 3: Temporal validity
self._check_temporal(observation)
# Check 4: Status consistency
self._check_status(observation)
# Check 5: Component consistency (e.g., BP)
self._check_components(observation)
return self.results
def _check_value_range(self, obs):
"""Check if numeric values are within physiological range."""
loinc_code = self._get_loinc_code(obs)
if not loinc_code or loinc_code not in VITAL_SIGN_RANGES:
return
rules = VITAL_SIGN_RANGES[loinc_code]
value = self._get_value(obs)
if value is None:
return
if value < rules["absolute_min"] or value > rules["absolute_max"]:
self.results.append(ValidationResult(
passed=False,
rule="clinical-value-range",
severity="error",
message=f"{rules['name']} value {value} is outside "
f"physiological range ({rules['absolute_min']}-"
f"{rules['absolute_max']})",
path="Observation.valueQuantity.value"
))
elif rules.get("critical_low") and value <= rules["critical_low"]:
self.results.append(ValidationResult(
passed=True,
rule="clinical-critical-value",
severity="warning",
message=f"{rules['name']} value {value} is at critical "
f"low level (threshold: {rules['critical_low']})",
path="Observation.valueQuantity.value"
))
elif rules.get("critical_high") and value >= rules["critical_high"]:
self.results.append(ValidationResult(
passed=True,
rule="clinical-critical-value",
severity="warning",
message=f"{rules['name']} value {value} is at critical "
f"high level (threshold: {rules['critical_high']})",
path="Observation.valueQuantity.value"
))
def _check_unit_consistency(self, obs):
"""Verify units match the expected unit for the observation type."""
loinc_code = self._get_loinc_code(obs)
if not loinc_code or loinc_code not in VITAL_SIGN_RANGES:
return
expected_unit = VITAL_SIGN_RANGES[loinc_code]["unit"]
actual_unit = obs.get("valueQuantity", {}).get("code", "")
if actual_unit and actual_unit != expected_unit:
self.results.append(ValidationResult(
passed=False,
rule="clinical-unit-consistency",
severity="error",
message=f"Expected unit '{expected_unit}' for "
f"{VITAL_SIGN_RANGES[loinc_code]['name']}, "
f"got '{actual_unit}'",
path="Observation.valueQuantity.code"
))
def _check_temporal(self, obs):
"""Check temporal validity of the observation."""
effective = obs.get("effectiveDateTime", "")
issued = obs.get("issued", "")
if effective:
try:
eff_dt = datetime.fromisoformat(effective.replace("Z", "+00:00"))
if eff_dt > datetime.now(eff_dt.tzinfo):
self.results.append(ValidationResult(
passed=False,
rule="clinical-temporal",
severity="error",
message="Observation effectiveDateTime is in the future",
path="Observation.effectiveDateTime"
))
except ValueError:
pass
def _check_status(self, obs):
"""Check status consistency."""
status = obs.get("status", "")
value = self._get_value(obs)
if status == "final" and value is None:
data_absent = obs.get("dataAbsentReason")
if not data_absent:
self.results.append(ValidationResult(
passed=False,
rule="clinical-status-value",
severity="warning",
message="Observation status is 'final' but no value or "
"dataAbsentReason is present",
path="Observation.status"
))
def _check_components(self, obs):
"""Check component consistency (e.g., BP systolic > diastolic)."""
components = obs.get("component", [])
systolic = diastolic = None
for comp in components:
code = self._get_loinc_from_codeable(comp.get("code", {}))
val = comp.get("valueQuantity", {}).get("value")
if code == "8480-6":
systolic = val
elif code == "8462-4":
diastolic = val
if systolic is not None and diastolic is not None:
if diastolic >= systolic:
self.results.append(ValidationResult(
passed=False,
rule="clinical-bp-consistency",
severity="error",
message=f"Diastolic BP ({diastolic}) >= Systolic BP "
f"({systolic}). Diastolic must be lower.",
path="Observation.component"
))
def _get_loinc_code(self, obs):
return self._get_loinc_from_codeable(obs.get("code", {}))
def _get_loinc_from_codeable(self, codeable):
for coding in codeable.get("coding", []):
if coding.get("system") == "http://loinc.org":
return coding.get("code")
return None
def _get_value(self, obs):
return obs.get("valueQuantity", {}).get("value")
# Usage
validator = ClinicalValidator()
results = validator.validate_observation({
"resourceType": "Observation",
"status": "final",
"code": {"coding": [{"system": "http://loinc.org", "code": "8480-6"}]},
"valueQuantity": {"value": 500, "unit": "mmHg", "code": "mm[Hg]"}
})
for r in results:
print(f"[{r.severity}] {r.rule}: {r.message}")Building a Complete Validation Pipeline
A production validation pipeline chains all three layers together, producing a comprehensive OperationOutcome resource:
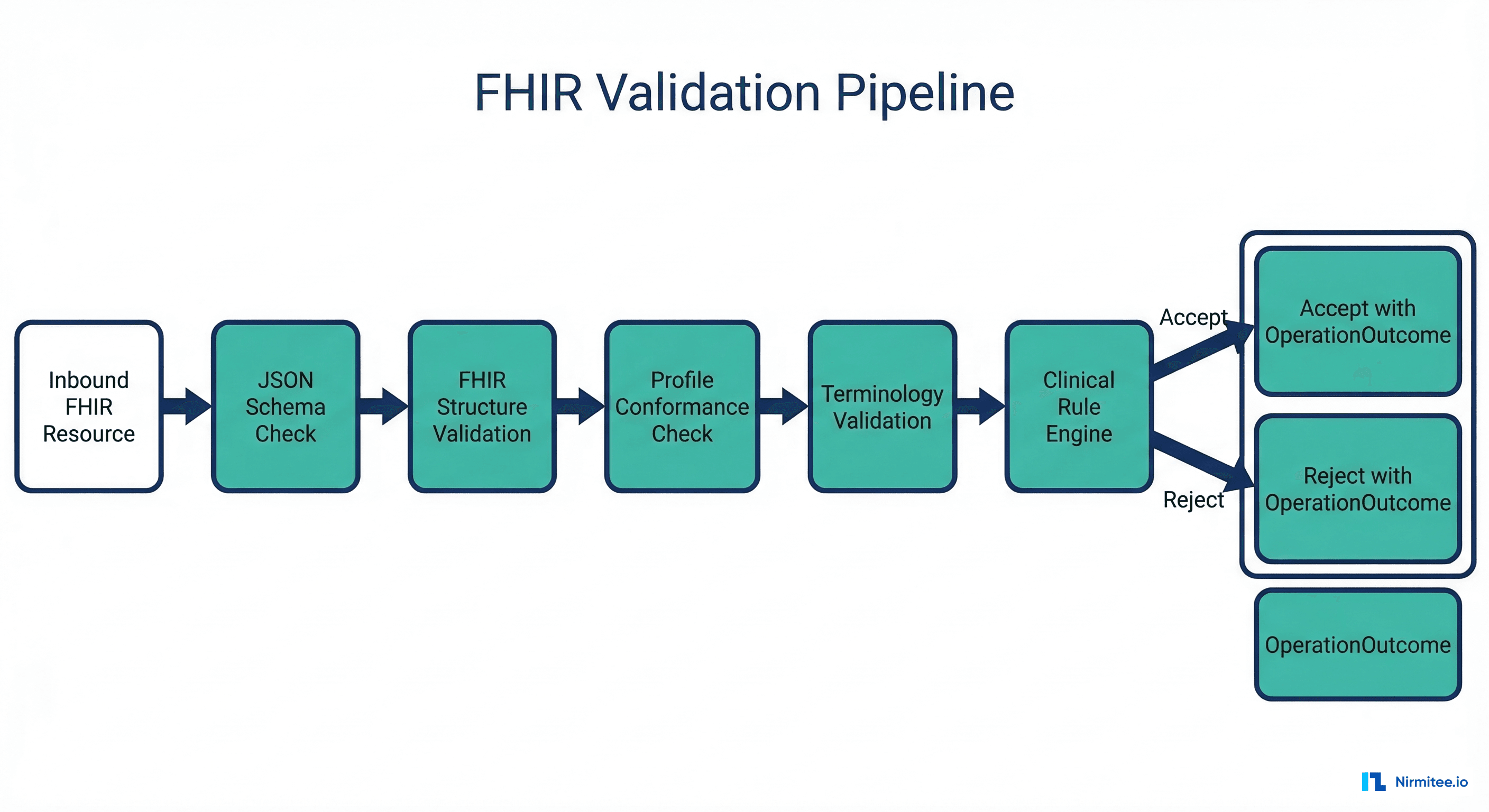
"""Complete FHIR validation pipeline combining all three layers."""
import json
from enum import Enum
class Severity(Enum):
ERROR = "error"
WARNING = "warning"
INFO = "information"
class ValidationPipeline:
def __init__(self):
self.clinical_validator = ClinicalValidator()
self.issues = []
def validate(self, resource_json: str, profile_url: str = None) -> dict:
"""Run full validation pipeline and return OperationOutcome."""
self.issues = []
# Layer 1: Structural
resource = self._validate_structure(resource_json)
if resource is None:
return self._to_operation_outcome()
# Layer 2: Profile (if specified)
if profile_url:
self._validate_profile(resource, profile_url)
# Layer 3: Clinical
resource_type = resource.get("resourceType", "")
if resource_type == "Observation":
clinical_results = self.clinical_validator.validate_observation(resource)
for result in clinical_results:
self.issues.append({
"severity": result.severity,
"code": "business-rule",
"details": {"text": result.message},
"diagnostics": f"Rule: {result.rule}",
"expression": [result.path] if result.path else []
})
return self._to_operation_outcome()
def _validate_structure(self, resource_json):
"""Layer 1: Basic structural validation."""
try:
resource = json.loads(resource_json)
except json.JSONDecodeError as e:
self.issues.append({
"severity": "error",
"code": "structure",
"details": {"text": f"Invalid JSON: {str(e)}"}
})
return None
if "resourceType" not in resource:
self.issues.append({
"severity": "error",
"code": "structure",
"details": {"text": "Missing required element: resourceType"}
})
return None
# Check required elements based on resource type
required = self._get_required_elements(resource["resourceType"])
for elem in required:
if elem not in resource:
self.issues.append({
"severity": "error",
"code": "required",
"details": {"text": f"Missing required element: {elem}"},
"expression": [f"{resource['resourceType']}.{elem}"]
})
return resource
def _validate_profile(self, resource, profile_url):
"""Layer 2: Profile validation (simplified)."""
# In production, delegate to HAPI FHIR validator or HL7 validator
self.issues.append({
"severity": "information",
"code": "informational",
"details": {"text": f"Profile validation against {profile_url} "
"requires external validator (HAPI/HL7)"}
})
def _get_required_elements(self, resource_type):
"""Return required elements for common resource types."""
required_map = {
"Observation": ["status", "code"],
"Patient": [],
"Condition": ["subject"],
"MedicationRequest": ["status", "intent", "medication", "subject"],
"AllergyIntolerance": ["patient"],
}
return required_map.get(resource_type, [])
def _to_operation_outcome(self):
"""Convert issues to a FHIR OperationOutcome."""
return {
"resourceType": "OperationOutcome",
"issue": self.issues if self.issues else [{
"severity": "information",
"code": "informational",
"details": {"text": "All validation checks passed"}
}]
}Terminology Validation
Terminology validation verifies that coded values (SNOMED CT, LOINC, ICD-10, RxNorm) are valid and appropriate. This sits between structural and clinical validation.

Binding Strength Matters
FHIR defines four levels of binding strength for coded elements:
| Binding Strength | Validation Behavior | Example |
|---|---|---|
| required | Code MUST come from the specified ValueSet. Reject if not. | Observation.status must be from ObservationStatus ValueSet |
| extensible | Code SHOULD come from ValueSet. Allow other codes if no suitable match exists. | Condition.code should be from US Core Condition Codes |
| preferred | Code is RECOMMENDED from ValueSet but alternatives are acceptable. | AllergyIntolerance.code preferred from SNOMED but allows others |
| example | ValueSet is illustrative only. Any valid code is acceptable. | Observation.code example binding to LOINC |
"""Terminology validation for FHIR coded elements."""
import requests
class TerminologyValidator:
def __init__(self, tx_server_url="https://tx.fhir.org/r4"):
self.tx_url = tx_server_url.rstrip("/")
def validate_code(self, system, code, value_set_url=None):
"""Validate a code against a terminology server."""
params = {
"system": system,
"code": code,
}
if value_set_url:
params["url"] = value_set_url
endpoint = f"{self.tx_url}/ValueSet/$validate-code"
resp = requests.get(endpoint, params=params, headers={
"Accept": "application/fhir+json"
})
if resp.status_code == 200:
result = resp.json()
for param in result.get("parameter", []):
if param.get("name") == "result":
return param.get("valueBoolean", False)
return False
def validate_coding(self, coding, binding_strength, value_set_url):
"""Validate a Coding element respecting binding strength."""
system = coding.get("system", "")
code = coding.get("code", "")
if not system or not code:
if binding_strength == "required":
return False, "Missing system or code in required binding"
return True, "No code to validate"
is_valid = self.validate_code(system, code, value_set_url)
if not is_valid and binding_strength == "required":
return False, f"Code {system}|{code} not in required ValueSet"
elif not is_valid and binding_strength == "extensible":
return True, f"Code {system}|{code} not in preferred ValueSet (extensible binding)"
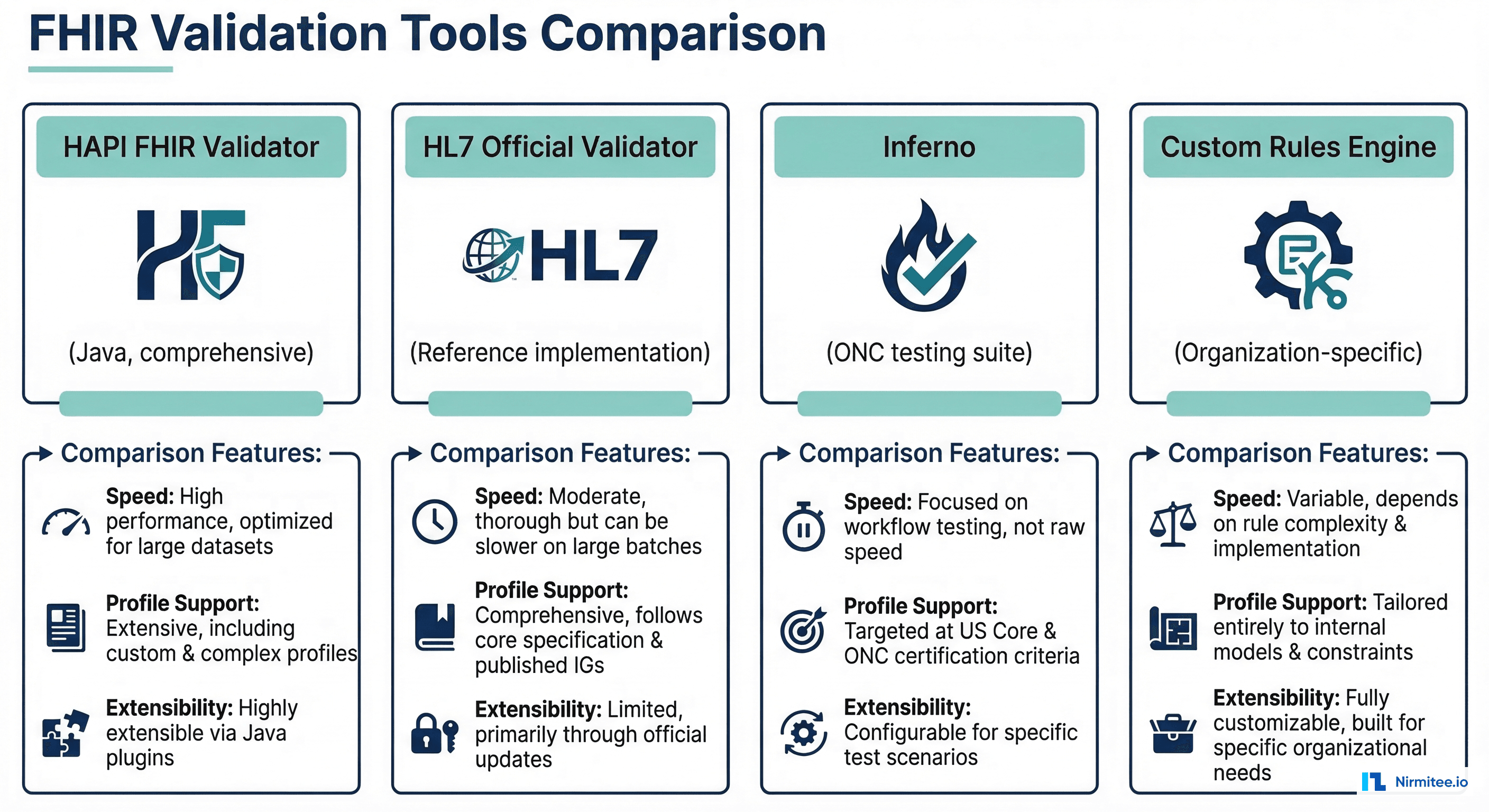
return True, "Valid"Validation Tools Comparison
Several tools exist for FHIR validation, each with different strengths:

| Tool | Language | Strengths | Limitations | Best For |
|---|---|---|---|---|
| HAPI FHIR Validator | Java | Fast, embeddable, IG package support | Java-only, memory-heavy | Server-side validation in Java apps |
| HL7 Official Validator | Java (CLI) | Reference implementation, most complete | Slow startup, complex setup | Conformance testing, CI/CD pipelines |
| Inferno | Ruby | ONC certification testing, scenario-based | Not real-time validation, test-focused | Regulatory conformance testing |
| firely-terminal | .NET | Interactive, profile authoring | .NET ecosystem only | Profile development and testing |
| Custom Rules Engine | Any | Clinical rules, business logic, fast | Must build and maintain | Clinical validation layer |
For teams using the HAPI FHIR server, the validator can be integrated directly into the server's request interceptor chain, validating every resource on write. Our guide on FHIR CapabilityStatements covers how to check whether a server supports the $validate operation.

Integrating Validation into Your Pipeline
Here's how to wire validation into a FHIR server's inbound data flow:
"""FHIR server middleware for validation."""
class FHIRValidationMiddleware:
def __init__(self, pipeline, strict_mode=False):
self.pipeline = pipeline
self.strict_mode = strict_mode
def validate_on_create(self, resource_json, profile_url=None):
"""Validate resource before accepting a CREATE operation."""
outcome = self.pipeline.validate(resource_json, profile_url)
errors = [i for i in outcome["issue"] if i["severity"] == "error"]
warnings = [i for i in outcome["issue"] if i["severity"] == "warning"]
if errors:
return {
"status": 422,
"outcome": outcome,
"accepted": False,
"message": f"Validation failed: {len(errors)} error(s)"
}
if warnings and self.strict_mode:
return {
"status": 422,
"outcome": outcome,
"accepted": False,
"message": f"Strict mode: {len(warnings)} warning(s)"
}
return {
"status": 201,
"outcome": outcome,
"accepted": True,
"message": "Resource accepted"
+ (f" with {len(warnings)} warning(s)" if warnings else "")
}Teams integrating with payer systems for the Patient Access API should validate inbound ExplanationOfBenefit resources against the CARIN IG profiles before persisting them.

Frequently Asked Questions
What is the difference between structural and clinical FHIR validation?
Structural validation checks that data conforms to the FHIR specification: valid JSON, correct data types, required fields present, proper resource type. Clinical validation checks that data is medically plausible: a blood pressure of 120/80 is structurally identical to 500/300 (both valid FHIR), but only the first is clinically real. Most FHIR implementations only do structural validation, missing clinically impossible data.
What tools validate FHIR resources against profiles like US Core?
The HAPI FHIR Validator (Java), HL7 Official FHIR Validator (Java CLI), and Firely Terminal (.NET) all support profile validation. They load Implementation Guide packages (NPM format) and validate resources against StructureDefinitions, including must-support elements, slicing constraints, FHIRPath invariants, and terminology bindings. The HL7 validator is considered the reference implementation for conformance testing.
How do I validate terminology bindings in FHIR?
Use the $validate-code operation on a terminology server (tx.fhir.org for public validation, or a local HAPI FHIR server with terminology loaded). Pass the code system, code value, and target ValueSet URL. The server returns whether the code is a member of the ValueSet. For required bindings, non-member codes must be rejected. For extensible bindings, non-member codes are allowed if no suitable match exists in the ValueSet.
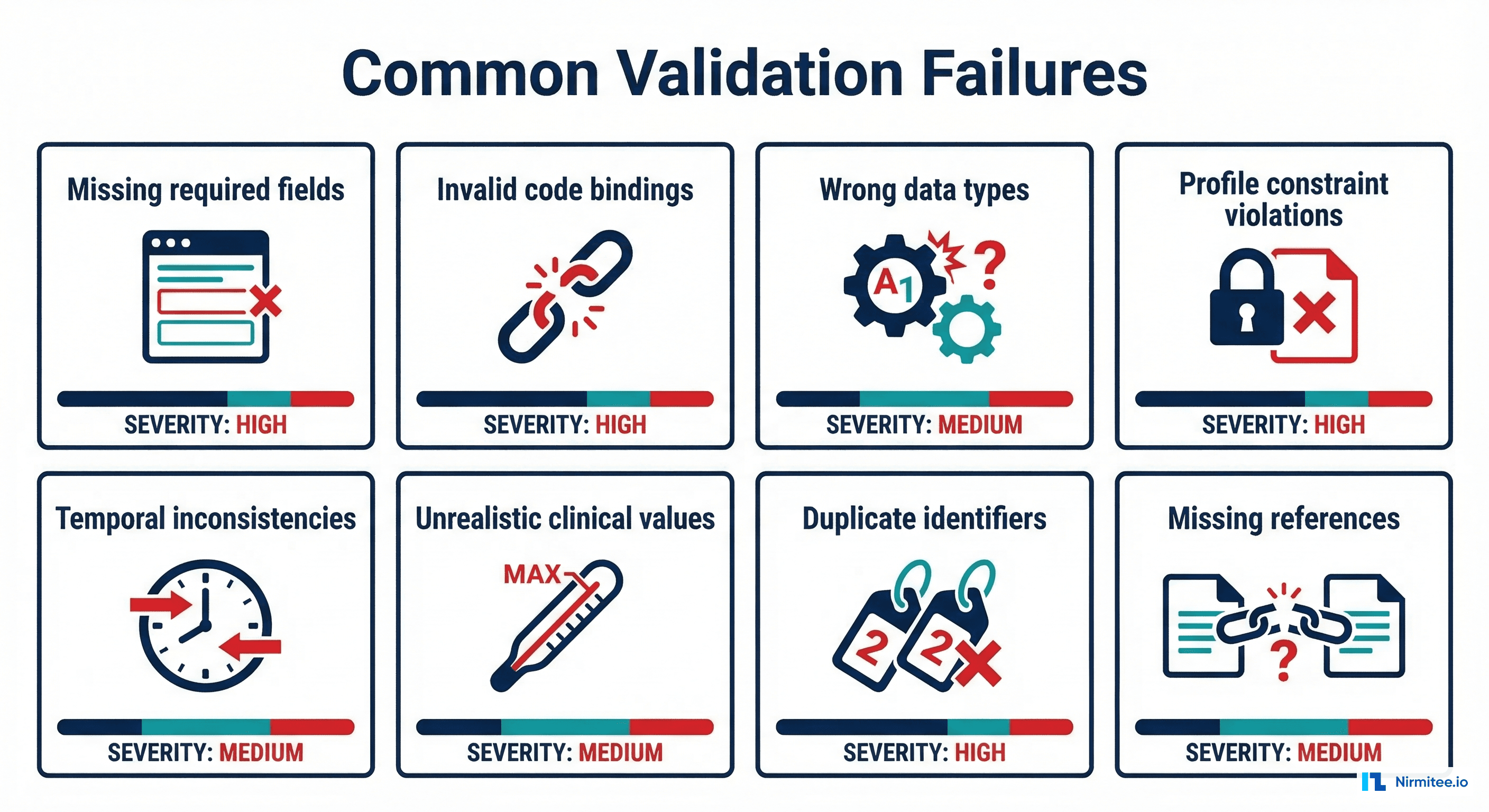
What are the most common FHIR validation failures?
The most common failures are: (1) missing required elements like Observation.status or Observation.code, (2) invalid terminology bindings where codes don't exist in the specified CodeSystem, (3) cardinality violations where arrays exceed max or single values are provided as arrays, (4) incorrect data types (string where date expected), and (5) profile constraint violations where must-support elements are missing or FHIRPath invariants fail.
Should I validate FHIR data on read or write?
Validate on write (create/update). Validating on read is too late — invalid data is already in the system and may have triggered incorrect clinical decisions. Server-side write validation acts as a gatekeeper. However, also validate on inbound data from external sources (payer feeds, device integrations, patient apps) before persisting. Read-time validation is useful only for migration scenarios where you need to assess the quality of existing data.
How do I handle validation in high-throughput scenarios?
For high-throughput scenarios (device data, bulk imports), use a tiered approach: fast structural validation inline (reject malformed data immediately), async profile validation (queue for background processing), and batch clinical validation (run periodic quality reports). Cache terminology lookups aggressively — most code validations are repeated frequently. Consider validating a sample rather than every resource in bulk import scenarios.