Why Accuracy Is the Wrong Metric for Clinical ML
A model that predicts "no sepsis" for every patient in a general ward achieves 97% accuracy—because only 3% of general ward patients develop sepsis. By the standard ML metric of accuracy, this model looks excellent. In clinical reality, it is worse than useless: it misses every single patient who is developing a life-threatening condition. This is the accuracy paradox, and it is the single most common mistake healthcare ML teams make when evaluating their models.
Clinicians do not think in terms of accuracy. They think in terms of clinical questions: "When this alert fires, how often is the patient actually sick?" (positive predictive value). "If the model says a patient is fine, can I trust that?" (negative predictive value). "What percentage of the actually sick patients does this model catch?" (sensitivity). "How many unnecessary work-ups will this model generate?" (1 - specificity). These are the metrics that determine whether a model improves patient care or simply generates noise.
This guide covers every metric that matters for clinical ML evaluation, with complete Python code for calculating each one, visual explanations, and guidance on which metrics to prioritize for different clinical use cases.
The Confusion Matrix: Foundation of Clinical Metrics
Every clinical metric derives from four fundamental counts in the confusion matrix. Understanding these counts in clinical terms—not just mathematical terms—is essential for communicating with clinical stakeholders.

| Cell | ML Term | Clinical Term | Example (Sepsis Prediction) | Impact |
|---|---|---|---|---|
| TP | True Positive | Correctly identified sick patient | Model alerts, patient is developing sepsis | Life saved (early intervention) |
| FP | False Positive | False alarm | Model alerts, patient is stable | Unnecessary blood cultures, alarm fatigue |
| TN | True Negative | Correctly identified stable patient | Model silent, patient is fine | No wasted resources |
| FN | False Negative | Missed diagnosis | Model silent, patient is developing sepsis | Delayed treatment, potential death |
# Complete clinical metrics from confusion matrix
import numpy as np
from sklearn.metrics import confusion_matrix, roc_auc_score
from sklearn.metrics import precision_recall_curve, average_precision_score
from sklearn.metrics import roc_curve, auc, brier_score_loss
from sklearn.calibration import calibration_curve
import matplotlib.pyplot as plt
def clinical_metrics(y_true, y_prob, threshold=0.5):
"""
Calculate all clinically relevant metrics.
Args:
y_true: ground truth labels (0/1)
y_prob: predicted probabilities (0-1)
threshold: classification threshold
Returns:
dict of clinical metrics
"""
y_pred = (y_prob >= threshold).astype(int)
tn, fp, fn, tp = confusion_matrix(y_true, y_pred).ravel()
# Core metrics
sensitivity = tp / (tp + fn) if (tp + fn) > 0 else 0 # Recall
specificity = tn / (tn + fp) if (tn + fp) > 0 else 0
ppv = tp / (tp + fp) if (tp + fp) > 0 else 0 # Precision
npv = tn / (tn + fn) if (tn + fn) > 0 else 0
# Likelihood ratios
lr_positive = sensitivity / (1 - specificity) if specificity < 1 else float('inf')
lr_negative = (1 - sensitivity) / specificity if specificity > 0 else float('inf')
# F-scores
f1 = 2 * ppv * sensitivity / (ppv + sensitivity) if (ppv + sensitivity) > 0 else 0
f2 = 5 * ppv * sensitivity / (4 * ppv + sensitivity) if (4 * ppv + sensitivity) > 0 else 0
# Discrimination metrics
auroc = roc_auc_score(y_true, y_prob)
auprc = average_precision_score(y_true, y_prob)
# Calibration
brier = brier_score_loss(y_true, y_prob)
# NNS — Number Needed to Screen
prevalence = y_true.mean()
nns = 1 / (sensitivity * prevalence) if (sensitivity * prevalence) > 0 else float('inf')
return {
"threshold": threshold,
"sensitivity": round(sensitivity, 4),
"specificity": round(specificity, 4),
"ppv": round(ppv, 4),
"npv": round(npv, 4),
"lr_positive": round(lr_positive, 2),
"lr_negative": round(lr_negative, 4),
"f1_score": round(f1, 4),
"f2_score": round(f2, 4),
"auroc": round(auroc, 4),
"auprc": round(auprc, 4),
"brier_score": round(brier, 4),
"nns": round(nns, 1),
"tp": int(tp), "fp": int(fp),
"tn": int(tn), "fn": int(fn),
"prevalence": round(prevalence, 4)
}Sensitivity and Specificity: The Clinical Tradeoff
Sensitivity (also called recall or true positive rate) answers: "Of all the patients who actually have the condition, what percentage does the model catch?" Specificity answers: "Of all the patients who are healthy, what percentage does the model correctly leave alone?" These two metrics exist in tension—increasing one typically decreases the other via the classification threshold.
The right balance depends entirely on the clinical context. For a sepsis early warning system, high sensitivity (95%+) is essential even at the cost of moderate specificity (60-70%), because missing a septic patient can be fatal while a false alarm only triggers a blood culture. For a cancer screening tool, the balance shifts depending on the downstream cost—if a false positive leads to an invasive biopsy, higher specificity is needed to prevent unnecessary procedures.
| Clinical Scenario | Priority Metric | Target | Rationale |
|---|---|---|---|
| Sepsis early warning | Sensitivity | 95%+ | Missing sepsis can be fatal; false alarms are manageable |
| Readmission risk | Sensitivity | 80%+ | Catching high-risk patients enables intervention |
| Cancer screening | NPV | 99%+ | Patients cleared by screening must be truly safe |
| Drug interaction alerts | Specificity | 90%+ | Excessive alerts cause alert fatigue, leading to overrides |
| Surgical risk prediction | Calibration | Slope ~1.0 | Surgeons need to trust that 30% risk means 30% |
| ICU triage | PPV | 60%+ | ICU beds are expensive; positive predictions must be reliable |
AUROC vs AUPRC: Which Curve Matters More?
AUROC (Area Under the Receiver Operating Characteristic curve) is the most commonly reported discrimination metric in clinical ML papers. It measures how well the model separates positive from negative cases across all possible thresholds. An AUROC of 0.5 means random guessing; 1.0 means perfect separation.
However, AUROC can be misleading for imbalanced clinical datasets—which is nearly every healthcare prediction problem. When the disease prevalence is 2%, a model with AUROC 0.85 might have a PPV of only 15% at a clinically reasonable threshold. AUPRC (Area Under the Precision-Recall Curve) better captures model performance when the positive class is rare, because it focuses on how well the model identifies the minority class without being inflated by the large number of easy-to-classify negative cases.

# Plot both curves side by side
def plot_roc_and_prc(y_true, y_prob, model_name="Model"):
"""Plot ROC and Precision-Recall curves with clinical context."""
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(14, 6))
# ROC Curve
fpr, tpr, roc_thresholds = roc_curve(y_true, y_prob)
roc_auc = auc(fpr, tpr)
ax1.plot(fpr, tpr, color='#2A9D8F', lw=2,
label=f'{model_name} (AUC = {roc_auc:.3f})')
ax1.plot([0, 1], [0, 1], 'k--', lw=1, label='Random (AUC = 0.500)')
ax1.set_xlabel('False Positive Rate (1 - Specificity)')
ax1.set_ylabel('True Positive Rate (Sensitivity)')
ax1.set_title('ROC Curve')
ax1.legend(loc='lower right')
ax1.set_xlim([0, 1])
ax1.set_ylim([0, 1.05])
# Precision-Recall Curve
precision, recall, pr_thresholds = precision_recall_curve(y_true, y_prob)
pr_auc = average_precision_score(y_true, y_prob)
prevalence = y_true.mean()
ax2.plot(recall, precision, color='#1B2B5B', lw=2,
label=f'{model_name} (AUPRC = {pr_auc:.3f})')
ax2.axhline(y=prevalence, color='gray', linestyle='--',
label=f'Random (Prevalence = {prevalence:.3f})')
ax2.set_xlabel('Recall (Sensitivity)')
ax2.set_ylabel('Precision (PPV)')
ax2.set_title('Precision-Recall Curve')
ax2.legend(loc='upper right')
ax2.set_xlim([0, 1])
ax2.set_ylim([0, 1.05])
plt.tight_layout()
plt.savefig('roc_prc_comparison.png', dpi=150)
plt.show()
return {"auroc": roc_auc, "auprc": pr_auc}Calibration: Does 70% Predicted Risk Actually Mean 70%?
A model can have excellent discrimination (high AUROC) but terrible calibration—meaning a predicted risk of 70% might correspond to an actual risk of 40% or 90%. Clinicians making treatment decisions need calibrated risk scores. When a surgeon sees "this patient has a 12% risk of surgical site infection," they need that number to be trustworthy. If the model consistently overestimates risk, patients may receive unnecessary interventions; if it underestimates, patients may be inadequately prepared.
# Calibration analysis with Brier score and calibration plot
def calibration_analysis(y_true, y_prob, n_bins=10, model_name="Model"):
"""Comprehensive calibration analysis."""
# Brier score (lower is better, 0 = perfect)
brier = brier_score_loss(y_true, y_prob)
# Calibration curve
fraction_of_positives, mean_predicted_value = calibration_curve(
y_true, y_prob, n_bins=n_bins, strategy='uniform'
)
# Calibration slope and intercept (logistic calibration)
from sklearn.linear_model import LogisticRegression
cal_model = LogisticRegression()
cal_model.fit(y_prob.reshape(-1, 1), y_true)
cal_slope = cal_model.coef_[0][0]
cal_intercept = cal_model.intercept_[0]
# Expected Calibration Error (ECE)
bin_edges = np.linspace(0, 1, n_bins + 1)
ece = 0
for i in range(n_bins):
mask = (y_prob >= bin_edges[i]) & (y_prob < bin_edges[i+1])
if mask.sum() > 0:
bin_accuracy = y_true[mask].mean()
bin_confidence = y_prob[mask].mean()
bin_weight = mask.sum() / len(y_true)
ece += bin_weight * abs(bin_accuracy - bin_confidence)
# Plot
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(14, 6))
ax1.plot(mean_predicted_value, fraction_of_positives,
's-', color='#2A9D8F', label=model_name)
ax1.plot([0, 1], [0, 1], 'k--', label='Perfectly calibrated')
ax1.set_xlabel('Mean Predicted Probability')
ax1.set_ylabel('Observed Frequency')
ax1.set_title(f'Calibration Plot (ECE = {ece:.3f})')
ax1.legend()
# Prediction distribution
ax2.hist(y_prob[y_true == 0], bins=50, alpha=0.5,
label='Negative', color='#1B2B5B')
ax2.hist(y_prob[y_true == 1], bins=50, alpha=0.5,
label='Positive', color='#2A9D8F')
ax2.set_xlabel('Predicted Probability')
ax2.set_ylabel('Count')
ax2.set_title('Prediction Distribution by Class')
ax2.legend()
plt.tight_layout()
plt.savefig('calibration_analysis.png', dpi=150)
plt.show()
return {
"brier_score": round(brier, 4),
"ece": round(ece, 4),
"calibration_slope": round(cal_slope, 3),
"calibration_intercept": round(cal_intercept, 3)
}A calibration slope of 1.0 and intercept of 0.0 indicates perfect calibration. Slopes greater than 1.0 indicate underconfidence (predicted probabilities are too compressed), while slopes less than 1.0 indicate overconfidence (predicted probabilities are too extreme). Many models benefit from post-hoc calibration using Platt scaling (logistic regression on validation predictions) or isotonic regression. When building clinical ML models for deployment, calibration should be part of your model evaluation pipeline alongside discrimination metrics.
NNT and NNS: Connecting ML to Clinical Workflow
Number Needed to Treat (NNT) and Number Needed to Screen (NNS) translate model performance into operational metrics that clinicians and hospital administrators understand. NNS answers: "How many patients do we need to screen with this model to identify one true positive?" This directly maps to staffing, workflow design, and resource allocation decisions.
def calculate_nnt_nns(y_true, y_prob, threshold, intervention_effectiveness=0.5):
"""
Calculate NNT and NNS from model predictions.
Args:
y_true: ground truth
y_prob: predicted probabilities
threshold: classification threshold
intervention_effectiveness: probability that intervening
on a true positive prevents the outcome (e.g., 0.5 = 50%
of readmissions prevented by the intervention)
"""
y_pred = (y_prob >= threshold).astype(int)
tn, fp, fn, tp = confusion_matrix(y_true, y_pred).ravel()
ppv = tp / (tp + fp) if (tp + fp) > 0 else 0
sensitivity = tp / (tp + fn) if (tp + fn) > 0 else 0
prevalence = y_true.mean()
# NNS: patients screened per true positive found
nns = 1 / ppv if ppv > 0 else float('inf')
# NNT: patients treated per outcome prevented
# NNT = NNS / intervention_effectiveness
nnt = nns / intervention_effectiveness if intervention_effectiveness > 0 else float('inf')
# Workload: total positive screens per day
positive_rate = (tp + fp) / (tp + fp + tn + fn)
return {
"nns": round(nns, 1),
"nnt": round(nnt, 1),
"ppv": round(ppv, 4),
"positive_screen_rate": round(positive_rate, 4),
"interpretation": (
f"For every {round(nns, 0):.0f} patients flagged by the model, "
f"{1} will truly have the condition. "
f"For every {round(nnt, 0):.0f} patients who receive the intervention, "
f"{1} outcome will be prevented."
)
}Fairness Audit: Performance Across Demographics
A model that performs well overall but poorly for specific demographic groups can perpetuate and even amplify healthcare disparities. The fairness audit evaluates model performance stratified by race, age, gender, and insurance status—the four dimensions most commonly associated with healthcare inequity in the United States.

def fairness_audit(y_true, y_prob, demographics: dict, threshold=0.3):
"""
Evaluate model fairness across demographic groups.
Args:
y_true: ground truth labels
y_prob: predicted probabilities
demographics: dict of {attribute_name: array_of_group_labels}
threshold: classification threshold
"""
results = {}
y_pred = (y_prob >= threshold).astype(int)
# Overall metrics
overall_auroc = roc_auc_score(y_true, y_prob)
tn, fp, fn, tp = confusion_matrix(y_true, y_pred).ravel()
overall_sensitivity = tp / (tp + fn)
overall_ppv = tp / (tp + fp) if (tp + fp) > 0 else 0
results["overall"] = {
"auroc": round(overall_auroc, 4),
"sensitivity": round(overall_sensitivity, 4),
"ppv": round(overall_ppv, 4),
"n": len(y_true)
}
for attr_name, groups in demographics.items():
results[attr_name] = {}
unique_groups = np.unique(groups)
for group in unique_groups:
mask = groups == group
if mask.sum() < 50: # Skip tiny groups
continue
group_y_true = y_true[mask]
group_y_prob = y_prob[mask]
group_y_pred = y_pred[mask]
if len(np.unique(group_y_true)) < 2:
continue # Need both classes
group_auroc = roc_auc_score(group_y_true, group_y_prob)
g_tn, g_fp, g_fn, g_tp = confusion_matrix(
group_y_true, group_y_pred
).ravel()
group_sensitivity = g_tp / (g_tp + g_fn) if (g_tp + g_fn) > 0 else 0
group_ppv = g_tp / (g_tp + g_fp) if (g_tp + g_fp) > 0 else 0
group_prevalence = group_y_true.mean()
results[attr_name][str(group)] = {
"auroc": round(group_auroc, 4),
"auroc_gap": round(abs(group_auroc - overall_auroc), 4),
"sensitivity": round(group_sensitivity, 4),
"ppv": round(group_ppv, 4),
"prevalence": round(group_prevalence, 4),
"n": int(mask.sum()),
"flag": group_auroc < overall_auroc - 0.05
}
# Equalized odds check
flagged_groups = []
for attr_name, groups_data in results.items():
if attr_name == "overall":
continue
for group_name, metrics in groups_data.items():
if metrics.get("flag", False):
flagged_groups.append(f"{attr_name}={group_name}")
results["fairness_summary"] = {
"passed": len(flagged_groups) == 0,
"flagged_groups": flagged_groups,
"max_auroc_gap": max(
m.get("auroc_gap", 0)
for attr in results.values() if isinstance(attr, dict)
for m in attr.values() if isinstance(m, dict)
) if len(results) > 1 else 0
}
return resultsThe fairness audit is not optional—it is a clinical and ethical requirement. The American Medical Association, the FDA, and the ONC have all issued guidance calling for demographic stratification of AI model performance. A model that achieves 0.85 AUROC overall but only 0.72 AUROC for Black patients is not a good model—it is a model that will worsen health disparities if deployed without correction. Solutions include collecting more representative training data, applying fairness-aware training objectives, and calibrating models separately by demographic group.
Clinical Decision Curve Analysis
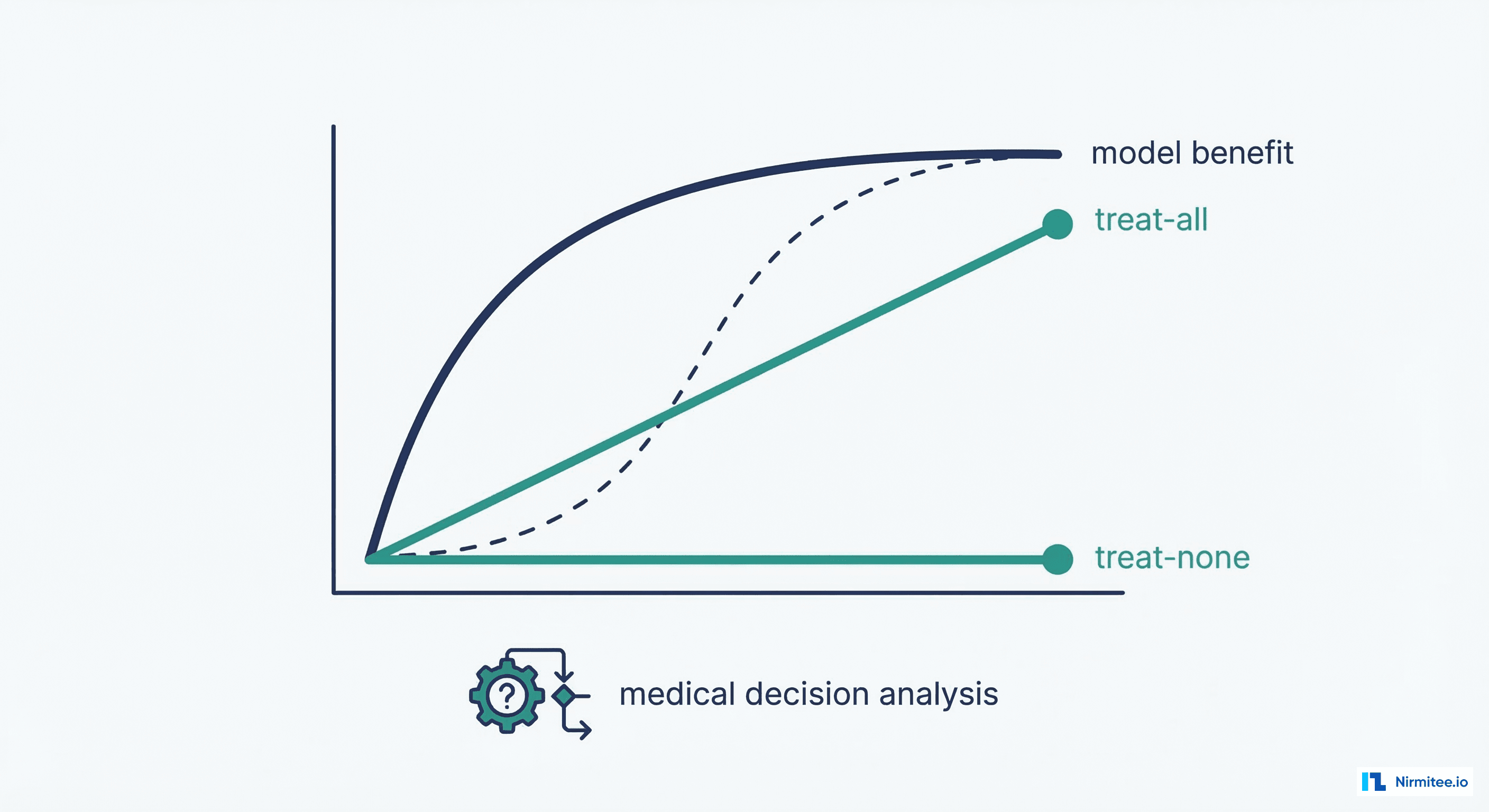
Decision Curve Analysis (DCA) answers the ultimate question: "Is using this model better than the alternatives?" The alternatives are simple strategies that require no model at all: treat everyone (assume all patients are high-risk) and treat no one (assume all patients are low-risk). A model has clinical utility only if its net benefit exceeds both of these strategies across a range of clinically relevant threshold probabilities.
def decision_curve_analysis(y_true, y_prob, thresholds=None, model_name="Model"):
"""
Decision Curve Analysis — does the model improve clinical decisions?
Net benefit = (TP/N) - (FP/N) * (pt / (1 - pt))
where pt = threshold probability
"""
if thresholds is None:
thresholds = np.arange(0.01, 0.99, 0.01)
n = len(y_true)
prevalence = y_true.mean()
net_benefits_model = []
net_benefits_all = []
for pt in thresholds:
# Model net benefit
y_pred = (y_prob >= pt).astype(int)
tp = ((y_pred == 1) & (y_true == 1)).sum()
fp = ((y_pred == 1) & (y_true == 0)).sum()
nb = (tp / n) - (fp / n) * (pt / (1 - pt))
net_benefits_model.append(nb)
# Treat-all strategy
nb_all = prevalence - (1 - prevalence) * (pt / (1 - pt))
net_benefits_all.append(nb_all)
# Plot
plt.figure(figsize=(10, 6))
plt.plot(thresholds, net_benefits_model, color='#2A9D8F', lw=2,
label=model_name)
plt.plot(thresholds, net_benefits_all, color='#1B2B5B', lw=2,
linestyle='--', label='Treat All')
plt.axhline(y=0, color='gray', linestyle=':', label='Treat None')
plt.xlabel('Threshold Probability')
plt.ylabel('Net Benefit')
plt.title('Decision Curve Analysis')
plt.legend()
plt.xlim([0, 0.5])
plt.ylim([-0.05, max(net_benefits_model) * 1.2])
plt.tight_layout()
plt.savefig('decision_curve.png', dpi=150)
plt.show()
# Find useful range (where model exceeds both alternatives)
useful_range = []
for i, pt in enumerate(thresholds):
if net_benefits_model[i] > max(0, net_benefits_all[i]):
useful_range.append(pt)
return {
"useful_threshold_range": (
round(min(useful_range), 2) if useful_range else None,
round(max(useful_range), 2) if useful_range else None
),
"max_net_benefit": round(max(net_benefits_model), 4),
"optimal_threshold": round(
thresholds[np.argmax(net_benefits_model)], 2
)
}Decision curve analysis is particularly important when presenting models to clinical leadership for deployment approval. Showing that a model provides net benefit across a range of clinically relevant thresholds—and quantifying that benefit in terms of avoided unnecessary treatments or detected true cases—is far more persuasive than presenting an AUROC number. This analysis should be part of the clinical review gate in your healthcare ML CI/CD pipeline.
Putting It All Together: Complete Evaluation Report
def generate_clinical_evaluation_report(
y_true, y_prob, demographics=None,
model_name="Readmission Prediction Model v2.1",
threshold=0.3
):
"""Generate a comprehensive clinical evaluation report."""
report = {"model_name": model_name, "threshold": threshold}
# 1. Core clinical metrics
report["clinical_metrics"] = clinical_metrics(y_true, y_prob, threshold)
# 2. Calibration
report["calibration"] = calibration_analysis(y_true, y_prob, model_name=model_name)
# 3. NNT/NNS
report["nnt_nns"] = calculate_nnt_nns(y_true, y_prob, threshold)
# 4. Decision curve
report["decision_curve"] = decision_curve_analysis(y_true, y_prob, model_name=model_name)
# 5. Fairness (if demographics provided)
if demographics:
report["fairness"] = fairness_audit(y_true, y_prob, demographics, threshold)
# 6. Threshold sensitivity analysis
thresholds_to_test = [0.1, 0.2, 0.3, 0.4, 0.5]
report["threshold_analysis"] = {}
for t in thresholds_to_test:
report["threshold_analysis"][str(t)] = clinical_metrics(y_true, y_prob, t)
# Print summary
m = report["clinical_metrics"]
print(f"\n{'='*60}")
print(f"Clinical Evaluation Report: {model_name}")
print(f"{'='*60}")
print(f"Threshold: {threshold}")
print(f"AUROC: {m['auroc']}")
print(f"AUPRC: {m['auprc']}")
print(f"Sensitivity: {m['sensitivity']} (target: >= 0.80)")
print(f"Specificity: {m['specificity']}")
print(f"PPV: {m['ppv']}")
print(f"NPV: {m['npv']}")
print(f"Brier Score: {m['brier_score']}")
print(f"NNS: {report['nnt_nns']['nns']}")
if demographics and report.get('fairness', {}).get('fairness_summary'):
fs = report['fairness']['fairness_summary']
status = 'PASSED' if fs['passed'] else 'FAILED'
print(f"Fairness: {status}")
if fs['flagged_groups']:
print(f" Flagged: {', '.join(fs['flagged_groups'])}")
return reportFrequently Asked Questions
Which single metric should I report to clinical stakeholders?
There is no single metric that captures clinical utility. At minimum, report sensitivity, PPV, and AUROC. Sensitivity tells clinicians how many cases the model catches. PPV tells them how often the alerts are correct. AUROC gives an overall discrimination score. But always present these in the context of the operating threshold and disease prevalence—without that context, the numbers are meaningless.
What AUROC is considered good for clinical ML?
In clinical literature, AUROC 0.70-0.80 is considered acceptable for most prediction tasks, 0.80-0.90 is good, and above 0.90 is excellent but should be viewed with suspicion (may indicate data leakage or overfitting). For comparison, the widely used LACE index for readmission prediction achieves approximately 0.68 AUROC. A new model should significantly exceed existing clinical baselines, not just exceed 0.50.
Why is Brier score important for healthcare?
Brier score measures both discrimination and calibration simultaneously (range 0-1, lower is better). A Brier score of 0.25 is no better than random guessing at 50% prevalence. For healthcare, Brier score matters because it penalizes both incorrect classifications and poorly calibrated probabilities. A model that correctly ranks patients by risk but assigns wildly inaccurate risk percentages will have a poor Brier score despite good AUROC.
How do I handle class imbalance in metric calculation?
Do not upsample or downsample when calculating evaluation metrics—use the natural class distribution. Use AUPRC instead of AUROC as your primary discrimination metric for rare outcomes. Choose a threshold based on clinical needs (not the default 0.5), and use the F2 score instead of F1 when sensitivity is more important than precision. Always report prevalence alongside all metrics so readers can interpret them correctly.
Should I recalibrate my model after deployment?
Yes. Calibration typically degrades over time as patient populations shift. Monitor calibration monthly using the calibration curve and Expected Calibration Error (ECE). When ECE exceeds 0.05 or the calibration slope deviates significantly from 1.0, apply recalibration using Platt scaling (logistic regression) on the most recent validation data. This does not require retraining the model—only adjusting the probability mapping.