The Migration Nobody Warns You About
You have built a custom EHR system. Maybe it started as a handful of PostgreSQL tables tracking patients, encounters, and vitals. Over the years it grew into a sprawling schema with hundreds of tables, custom fields clinicians demanded, and business logic baked into stored procedures. Now you need to migrate to openEHR.
Perhaps your organization needs to participate in a national health data exchange that mandates openEHR. Perhaps you are hitting the ceiling of your custom schema's ability to represent evolving clinical models. Perhaps you have realized that maintaining your own clinical data model is a full-time job that has nothing to do with your core product.
Whatever the reason, this migration is one of the most complex technical projects in health IT. It is not a simple ETL job. You are fundamentally changing how clinical data is modeled, stored, queried, and validated. This guide covers what actually happens when you do it, based on real migrations we have supported at Nirmitee.
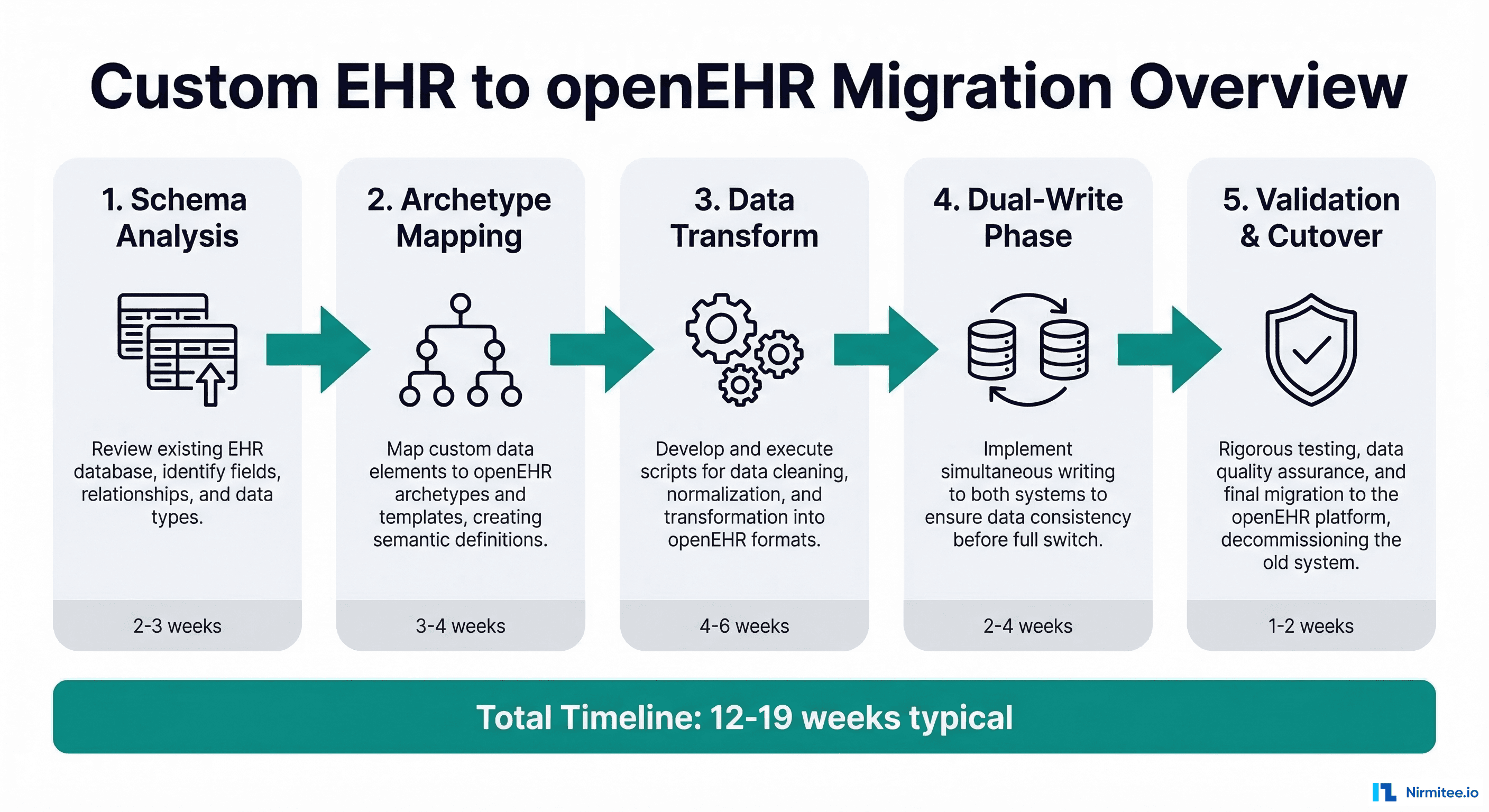
Phase 1: Schema Analysis and Archetype Mapping
Auditing Your Existing Schema
Before touching any openEHR tooling, you need a complete inventory of your current data model. This is not just a list of tables. You need to understand:
- Active vs. dead tables — In any system older than three years, at least 15-20% of tables are no longer written to. Migrating dead data wastes months.
- Data volumes per table — A table with 50 million rows of vital signs requires a different migration strategy than a lookup table with 200 rows.
- Foreign key relationships — Custom schemas often have implicit relationships not enforced by constraints. Interview the developers who built them.
- Business logic in the database — Stored procedures, triggers, views, and materialized views that transform data on read. This logic must be replicated or replaced.
- Custom fields and extensions — The columns clinicians asked for that do not map to any standard. These are the hardest part of the migration.
Run a comprehensive schema audit. Here is a SQL query that generates a useful starting inventory on PostgreSQL:
-- Schema audit: table sizes, row counts, and column inventory
SELECT
t.table_schema,
t.table_name,
pg_size_pretty(pg_total_relation_size(quote_ident(t.table_schema) || '.' || quote_ident(t.table_name))) as total_size,
(SELECT count(*) FROM information_schema.columns c WHERE c.table_name = t.table_name AND c.table_schema = t.table_schema) as column_count,
pg_stat_get_live_tuples(quote_ident(t.table_schema) || '.' || quote_ident(t.table_name))::regclass) as estimated_rows
FROM information_schema.tables t
WHERE t.table_schema NOT IN ('pg_catalog', 'information_schema')
ORDER BY pg_total_relation_size(quote_ident(t.table_schema) || '.' || quote_ident(t.table_name)) DESC;Mapping Tables to Archetypes
This is the core intellectual work of the migration. Every table or group of related tables in your custom schema must be mapped to one or more openEHR archetypes. An archetype is a formal, version-controlled clinical data model that defines the structure, constraints, and terminology bindings for a specific clinical concept.
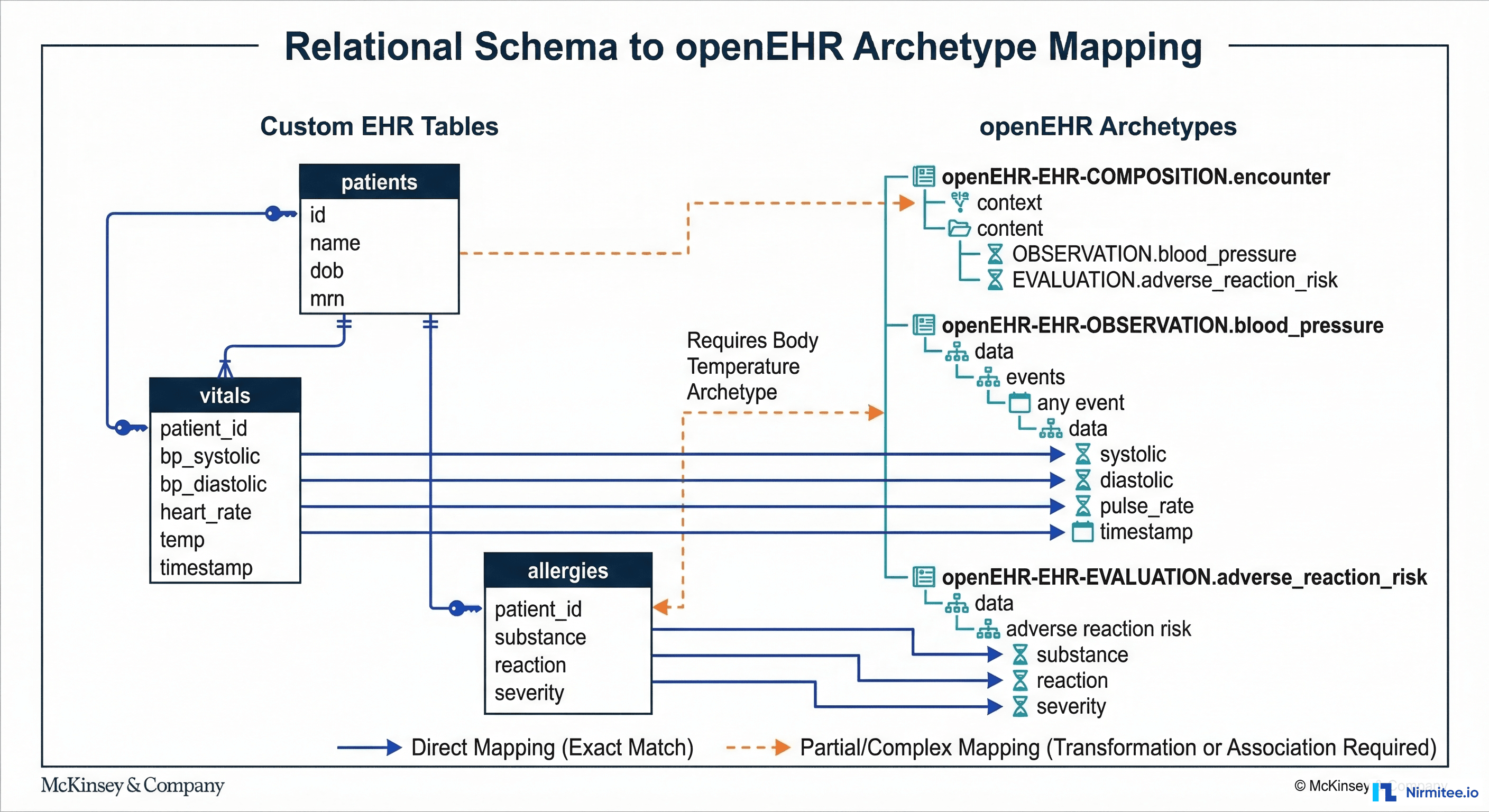
The openEHR Clinical Knowledge Manager (CKM) hosts the international archetype library. For most common clinical concepts, an archetype already exists. Here are the typical mappings:
| Custom EHR Table | openEHR Archetype | Mapping Complexity |
|---|---|---|
| patients / demographics | openEHR-EHR-ADMIN_ENTRY.demographic | Low — direct field mapping |
| vitals / observations | openEHR-EHR-OBSERVATION.blood_pressure, .body_temperature, .pulse | Medium — one table splits into multiple archetypes |
| lab_results | openEHR-EHR-OBSERVATION.laboratory_test_result | Medium — needs CLUSTER.laboratory_test_analyte for each analyte |
| allergies | openEHR-EHR-EVALUATION.adverse_reaction_risk | Low — well-defined archetype |
| medications | openEHR-EHR-INSTRUCTION.medication_order + ACTION.medication | High — order vs. administration split |
| diagnoses / problems | openEHR-EHR-EVALUATION.problem_diagnosis | Medium — terminology mapping needed (ICD-10 to SNOMED CT) |
| clinical_notes | openEHR-EHR-EVALUATION.clinical_synopsis or OBSERVATION.progress_note | High — unstructured text, limited archetype fit |
| custom_assessments | May require custom archetype or GENERIC_ENTRY | Very High — no standard archetype exists |
The critical insight: your custom schema is entity-oriented (one table per concept), while openEHR is composition-oriented (clinical documents containing multiple archetypes). A single patient encounter in your system might pull from 8-10 tables. In openEHR, that same encounter becomes a single Composition containing multiple Observations, Evaluations, and Instructions.
When No Archetype Exists
For roughly 10-20% of your data, you will not find a suitable archetype in CKM. You have three options:
- Create a custom archetype — Follow the openEHR Archetype Design Guidelines. Submit it to CKM for peer review. This takes 4-8 weeks for review and acceptance.
- Use GENERIC_ENTRY — A catch-all archetype for data that does not fit established patterns. Acceptable for migration, but should be replaced with proper archetypes over time.
- Archive without migration — Some data (UI preferences, temporary processing fields, deprecated columns) does not belong in a clinical data repository. Archive it to cold storage and document the decision.
Phase 2: Building the Data Transformation Pipeline
With mappings defined, you need to build the pipeline that actually transforms data from your relational schema into openEHR Compositions and posts them to the Clinical Data Repository (CDR).
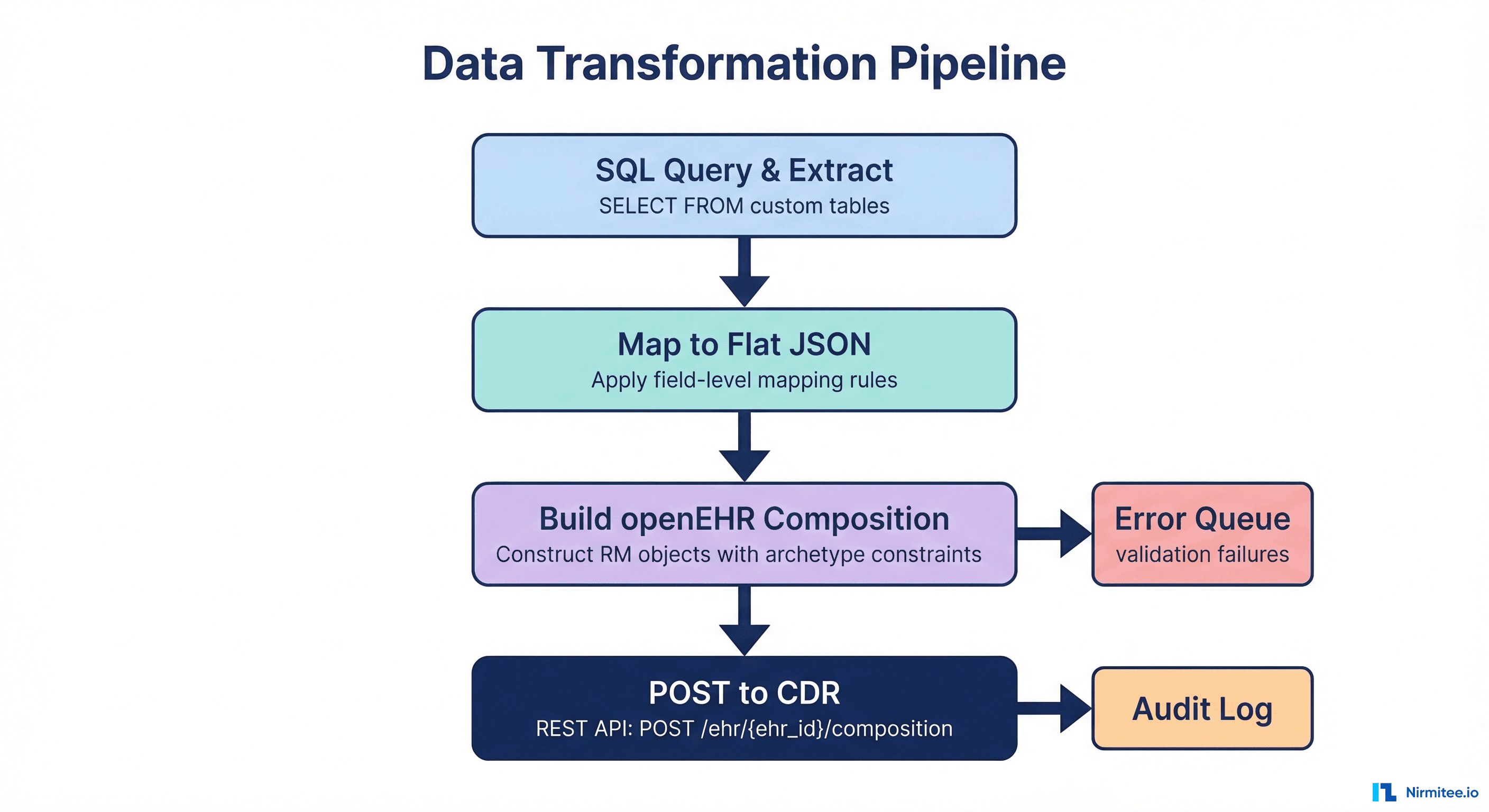
Step 1: Extract from Custom Schema
Write SQL queries that extract data grouped by patient encounter, not by table. This is critical because openEHR Compositions are encounter-centric. A single composition might need data from your vitals, lab_results, medications, and allergies tables for one encounter.
-- Extract encounter-grouped data for transformation
SELECT
e.encounter_id,
e.patient_id,
e.encounter_date,
e.encounter_type,
-- Vitals (pivoted from rows to columns)
MAX(CASE WHEN v.vital_type = 'BP_SYSTOLIC' THEN v.value END) as bp_systolic,
MAX(CASE WHEN v.vital_type = 'BP_DIASTOLIC' THEN v.value END) as bp_diastolic,
MAX(CASE WHEN v.vital_type = 'HEART_RATE' THEN v.value END) as heart_rate,
MAX(CASE WHEN v.vital_type = 'TEMPERATURE' THEN v.value END) as temperature,
-- Allergies (aggregated as JSON array)
COALESCE(
json_agg(DISTINCT jsonb_build_object(
'substance', a.substance,
'reaction', a.reaction,
'severity', a.severity
)) FILTER (WHERE a.id IS NOT NULL),
'[]'::json
) as allergies
FROM encounters e
LEFT JOIN vitals v ON v.encounter_id = e.encounter_id
LEFT JOIN patient_allergies a ON a.patient_id = e.patient_id
WHERE e.encounter_date >= '2020-01-01'
GROUP BY e.encounter_id, e.patient_id, e.encounter_date, e.encounter_type
ORDER BY e.encounter_date;Step 2: Map to Flat JSON (Web Template Format)
openEHR CDRs like EHRbase and Better Platform accept data in a flat JSON format derived from the Operational Template (OPT). Each field is addressed by its AQL path. Here is what a blood pressure mapping looks like:
{
"ctx/language": "en",
"ctx/territory": "US",
"ctx/composer_name": "Migration Script",
"encounter/blood_pressure/any_event/systolic|magnitude": 120,
"encounter/blood_pressure/any_event/systolic|unit": "mm[Hg]",
"encounter/blood_pressure/any_event/diastolic|magnitude": 80,
"encounter/blood_pressure/any_event/diastolic|unit": "mm[Hg]",
"encounter/blood_pressure/any_event/time": "2024-06-15T10:30:00Z",
"encounter/blood_pressure/position": "at1000",
"encounter/body_temperature/any_event/temperature|magnitude": 37.2,
"encounter/body_temperature/any_event/temperature|unit": "Cel",
"encounter/pulse_heart_beat/any_event/rate|magnitude": 72,
"encounter/pulse_heart_beat/any_event/rate|unit": "/min"
}Step 3: POST Compositions to the CDR
Each transformed record becomes a POST request to the CDR's composition endpoint. The CDR validates the composition against the archetype constraints before storing it.
# POST a flat composition to EHRbase
curl -X POST "https://ehrbase.example.com/ehrbase/rest/openehr/v1/ehr/${EHR_ID}/composition" -H "Content-Type: application/json" -H "Accept: application/json" -H "Prefer: return=representation" -d @composition.json
# Expected response: 201 Created with composition UID
# {"_type": "COMPOSITION", "uid": {"value": "8f3a...::ehrbase::1"}}Key considerations for the POST pipeline:
- Rate limiting — Most CDRs handle 50-200 compositions per second. Plan your batch sizes accordingly.
- Idempotency — Track which records have been migrated. If the pipeline fails mid-batch, you need to resume without duplicating data.
- Validation errors — The CDR will reject compositions that violate archetype constraints. Route failures to an error queue for manual review, not into a retry loop.
- EHR creation — Each patient needs an EHR object created in the CDR before compositions can be posted. Batch-create EHRs first using the
/ehrendpoint with the patient's external ID as the subject.
Phase 3: Handling Unmappable Data
Every custom EHR has data that does not cleanly map to openEHR archetypes. Based on migrations we have worked on, expect this distribution:
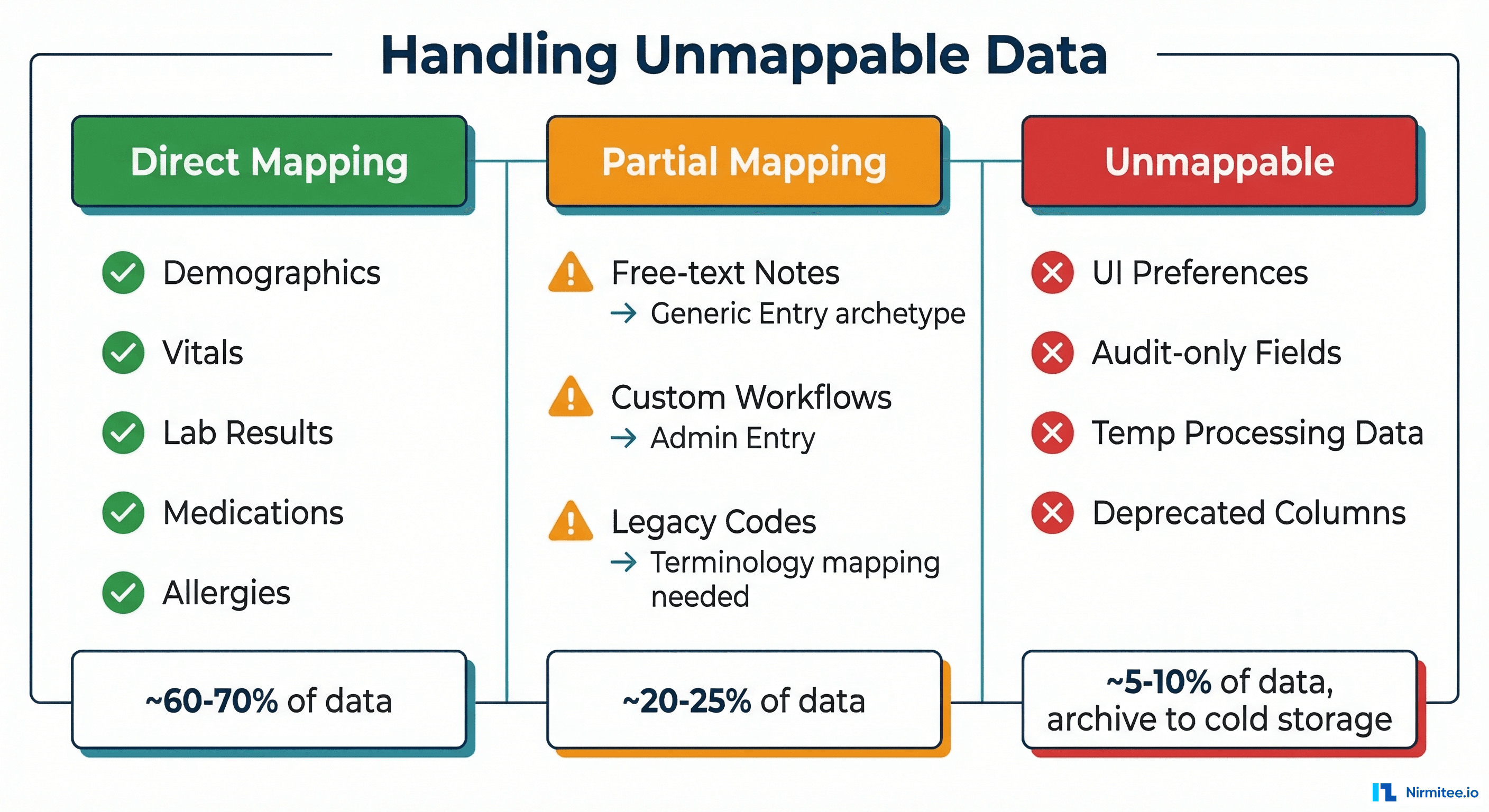
Free-Text Clinical Notes
Unstructured clinical notes are the biggest challenge. A typical EHR has millions of free-text notes that contain clinically valuable information but cannot be decomposed into structured archetype fields without NLP. Options:
- Store as-is in
openEHR-EHR-EVALUATION.clinical_synopsis— preserves the text but gains no structured querying benefit. - NLP extraction — Use clinical NLP (cTAKES, MetaMap, or a fine-tuned LLM) to extract coded concepts from notes and store both the original text and extracted codes. This is expensive and error-prone.
- Hybrid approach — Store the original text in a synopsis archetype, flag notes for future NLP processing. This is the pragmatic choice for most migrations.
Custom Fields and Proprietary Extensions
Fields like workflow_status, billing_flag, custom_risk_score, or department-specific fields created at clinician request. For each one, decide:
- Is this clinical data that should exist in the CDR? If yes, find or create an archetype.
- Is this operational data (workflow, billing, audit)? If yes, it belongs in a separate operational database, not the CDR.
- Is this dead data nobody uses? If yes, archive and document.
Phase 4: Running Dual-Write During Transition
Cutting over from one data store to another on a specific date is a recipe for disaster in healthcare. The safe approach is a dual-write transition where both the old and new systems receive writes simultaneously.
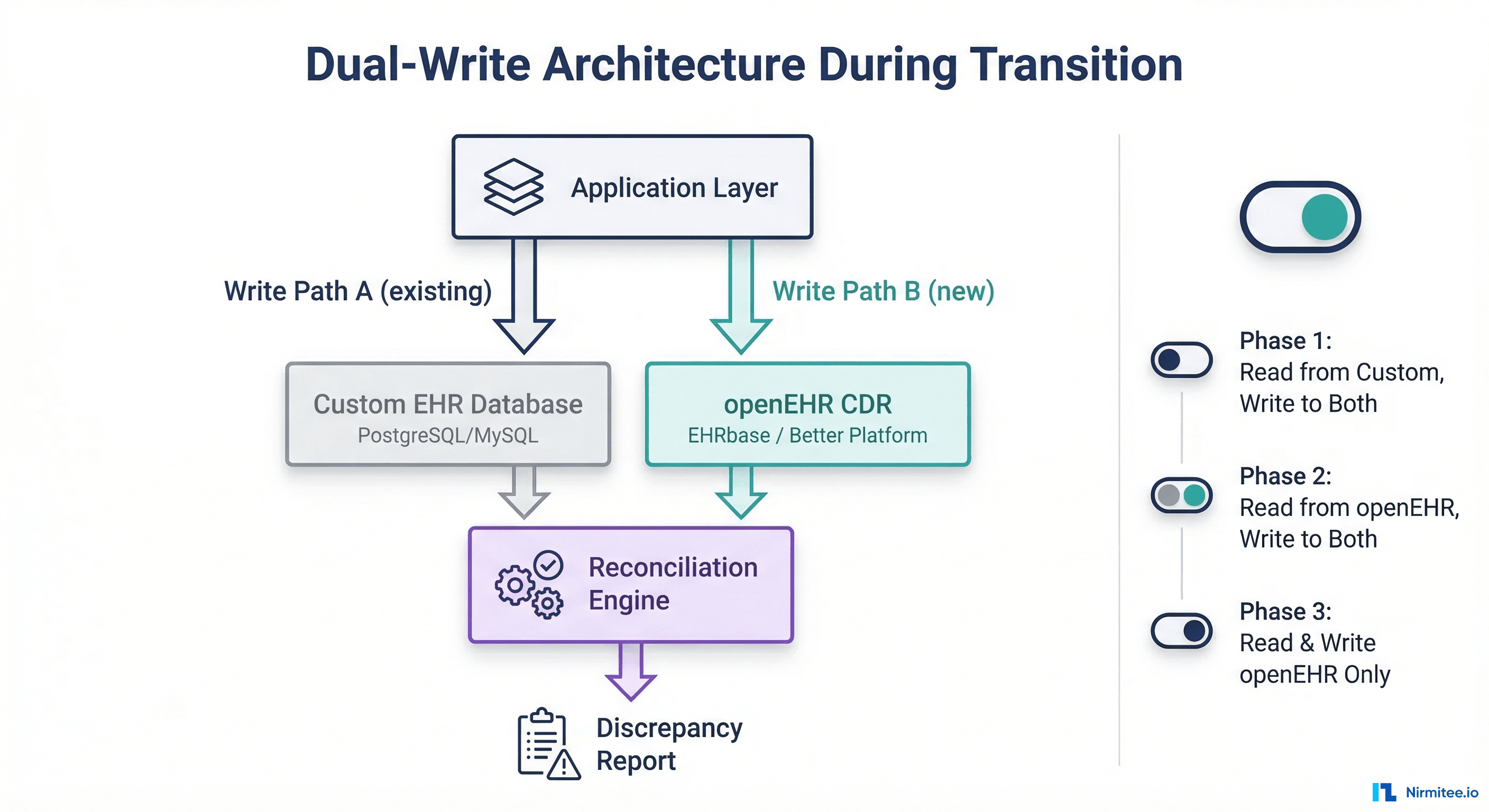
Implementation Architecture
The dual-write layer sits between your application and both databases. Every write operation is duplicated:
class DualWriteService:
def __init__(self, custom_db, openehr_cdr, reconciler):
self.custom_db = custom_db
self.openehr_cdr = openehr_cdr
self.reconciler = reconciler
self.phase = "PHASE_1" # Read Custom, Write Both
def save_encounter(self, encounter_data):
# Always write to custom DB (existing path)
custom_result = self.custom_db.save(encounter_data)
# Transform and write to openEHR CDR
try:
composition = self.transform_to_composition(encounter_data)
openehr_result = self.openehr_cdr.post_composition(composition)
# Queue reconciliation check
self.reconciler.queue_check(
custom_id=custom_result.id,
composition_uid=openehr_result.uid,
timestamp=datetime.utcnow()
)
except OpenEHRValidationError as e:
# Log but do not fail the primary write
logger.error(f"Dual-write openEHR failure: {e}")
self.reconciler.queue_failure(custom_result.id, str(e))
# Read from the appropriate source based on phase
if self.phase == "PHASE_1":
return custom_result
elif self.phase == "PHASE_2":
return self.openehr_cdr.get_composition(openehr_result.uid)
else: # PHASE_3
return self.openehr_cdr.get_composition(openehr_result.uid)The Three Phases
Phase 1: Read from Custom, Write to Both — Your application continues reading from the custom database. The openEHR CDR receives shadow writes. The reconciliation engine compares both stores nightly, flagging discrepancies. Duration: 2-4 weeks.
Phase 2: Read from openEHR, Write to Both — Flip the read path to openEHR. The custom database is now the shadow. If openEHR reads fail or return unexpected results, you can instantly revert to Phase 1. Duration: 2-4 weeks.
Phase 3: Read and Write openEHR Only — Decommission the custom database write path. The custom database becomes read-only for historical queries during a sunset period. Duration: permanent after validation.
Reconciliation Engine
The reconciliation engine is the safety net that makes dual-write viable. It runs nightly batch comparisons:
# Reconciliation query: compare record counts by date
-- Custom DB side
SELECT DATE(encounter_date) as dt, COUNT(*) as custom_count
FROM encounters
WHERE encounter_date >= CURRENT_DATE - INTERVAL '7 days'
GROUP BY DATE(encounter_date);
-- openEHR CDR side (via AQL)
SELECT c/context/start_time/value as dt, COUNT(c) as openehr_count
FROM EHR e CONTAINS COMPOSITION c
WHERE c/context/start_time/value >= '2024-06-01'
GROUP BY c/context/start_time/valueKey metrics to reconcile: record counts per day, field-level checksums on a sample, coded value distribution (are the same SNOMED codes appearing?), and query result equivalence for key clinical reports.
Phase 5: Validation and Go-Live
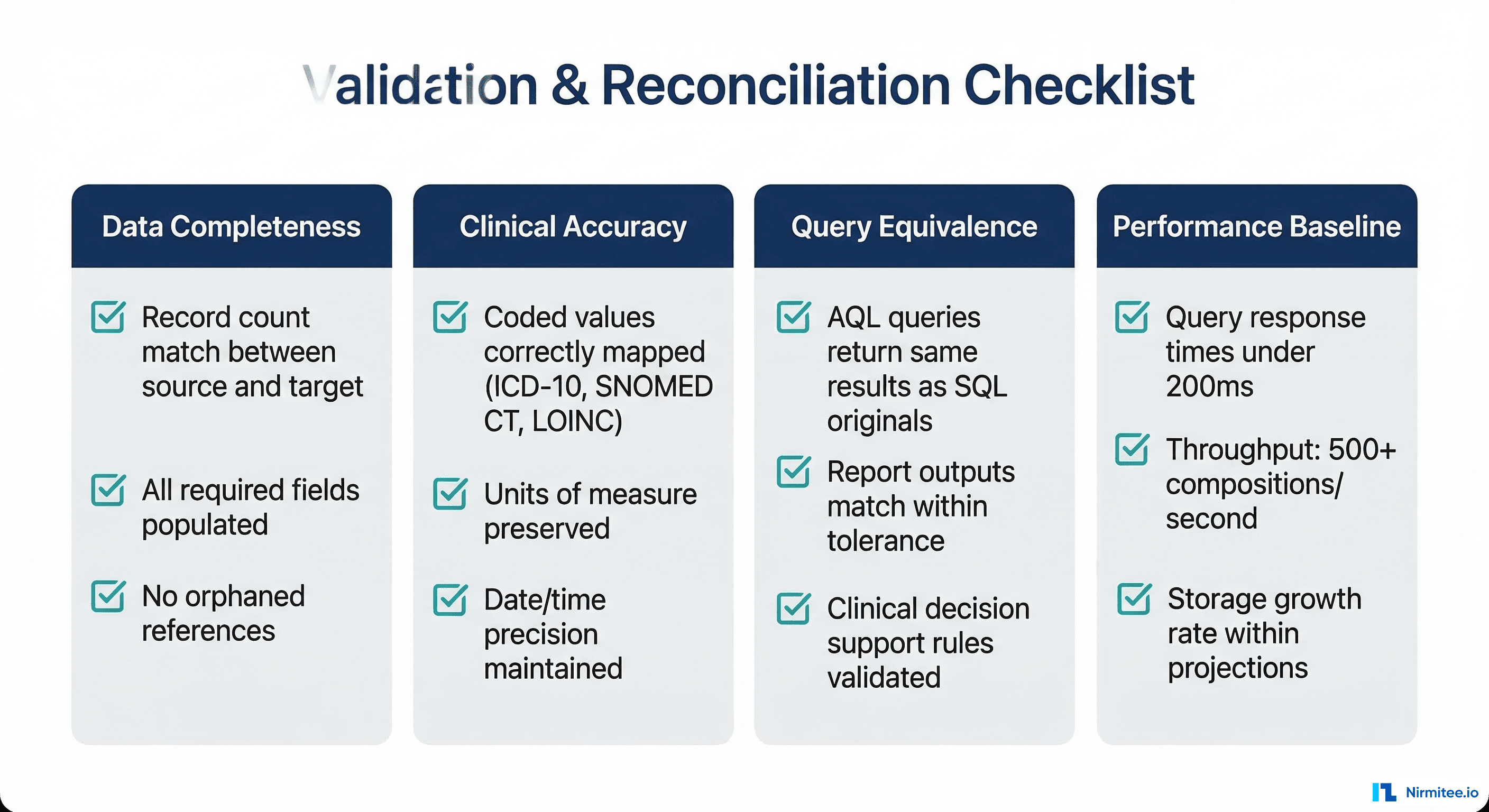
Clinical Validation
Technical validation (record counts, checksums) is necessary but not sufficient. You also need clinical validation from domain experts:
- Chart review — Have clinicians compare patient records in both systems for a sample of 50-100 patients. They check that clinical narratives, medication lists, allergy information, and problem lists are complete and accurate.
- Clinical decision support — If your system has CDS rules (drug interaction checks, allergy alerts, screening reminders), verify they produce identical results against the openEHR data.
- Reporting equivalence — Run all regulatory reports (quality measures, adverse event reports, public health reporting) against both data stores and compare outputs.
Performance Testing
openEHR CDRs have different performance characteristics than custom relational databases. Key benchmarks to establish:
| Operation | Target | Notes |
|---|---|---|
| Single composition POST | Under 100ms | EHRbase: ~50-80ms typical |
| AQL query (single patient) | Under 200ms | Varies by query complexity |
| AQL query (population) | Under 5s | Full-scan queries will be slower than indexed SQL |
| Batch composition POST | 200+ per second | Use async HTTP clients and connection pooling |
| Template upload | Under 2s | One-time operation, not performance-critical |
If AQL query performance does not meet your requirements, consider:
- Materialized views — Some CDRs support materialized AQL views that pre-compute frequently used queries.
- FHIR facade — Expose frequently queried data via a FHIR API backed by a read-optimized secondary store.
- Denormalized read models — CQRS pattern where writes go to the CDR and reads come from a denormalized PostgreSQL or Elasticsearch index.
Timeline, Staffing, and Budget
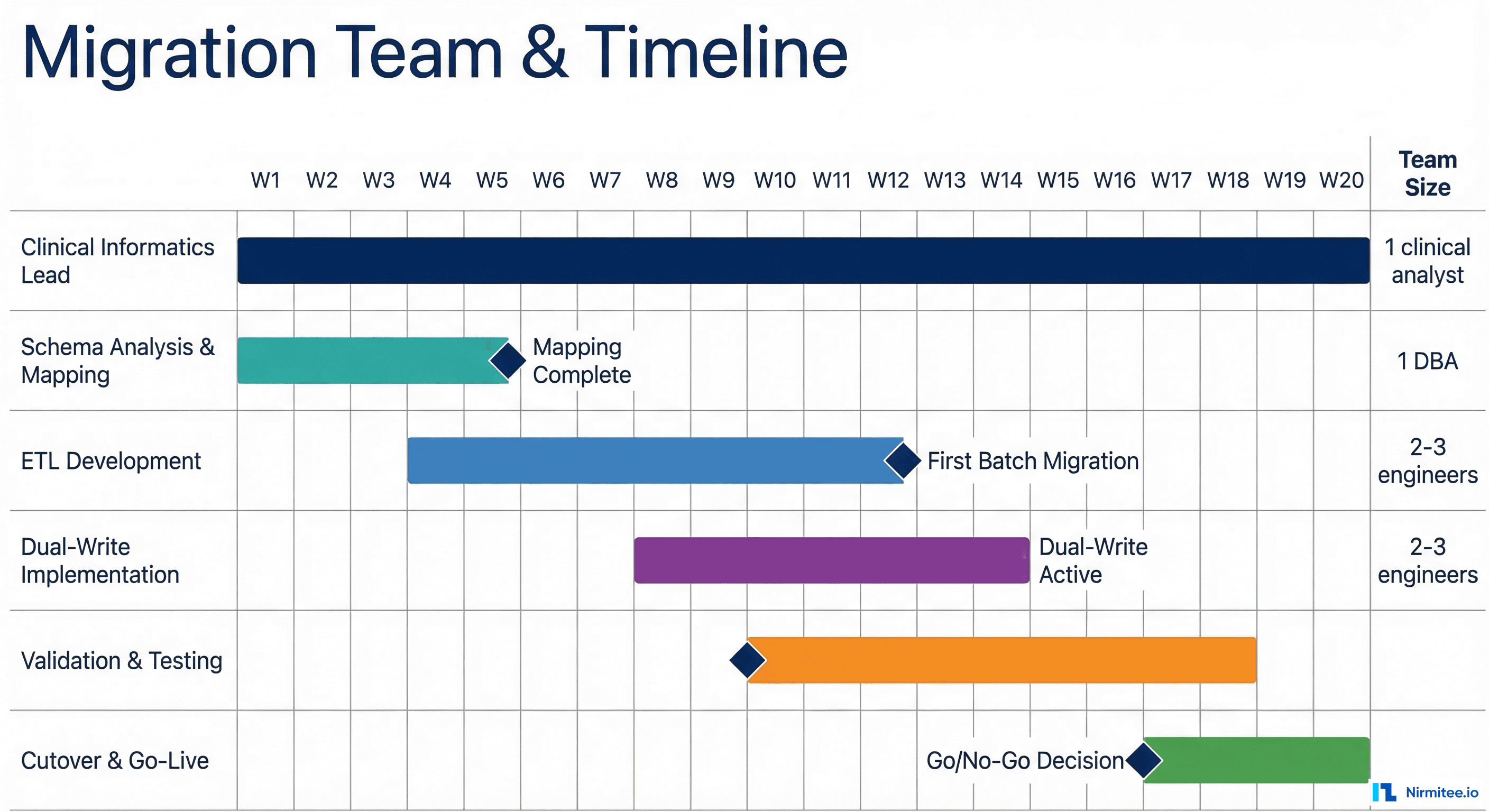
Realistic Timeline
Based on migrations involving 50-200 table custom schemas with 1-10 million patient records:
- Schema analysis and archetype mapping: 3-5 weeks. This is slower than expected because it requires clinical informatics expertise, not just database skills.
- ETL pipeline development: 6-10 weeks. Each archetype mapping requires its own transformation logic, unit tests, and edge case handling.
- Dual-write implementation: 2-4 weeks. The application layer changes are straightforward; the reconciliation engine takes longer.
- Validation and testing: 4-6 weeks. Clinical validation cannot be rushed. Chart reviews and report comparisons take time.
- Cutover and go-live: 1-2 weeks. The actual switch is fast if dual-write has been running successfully.
Total: 16-27 weeks for a medium-complexity migration. Smaller systems (under 50 tables) can be done in 12-16 weeks. Large enterprise EHRs with thousands of tables and complex stored procedure logic can take 6-12 months.
Team Composition
You need a cross-functional team, not just backend engineers:
- Clinical Informaticist (1, full-time) — The single most important role. This person understands both the clinical domain and openEHR modeling. They drive archetype selection, review mappings for clinical accuracy, and lead chart reviews. Without this role, engineers will make incorrect clinical assumptions.
- Backend Engineers (2-3, full-time) — Build the ETL pipeline, dual-write layer, and reconciliation engine. Need experience with your existing tech stack plus openEHR REST APIs.
- DBA / Data Engineer (1, part-time) — Manages the schema audit, optimizes extraction queries, handles data quality issues in the source system.
- QA / Test Engineer (1, part-time becoming full-time) — Builds validation test suites, runs clinical report comparisons, manages the testing phase.
Common Pitfalls
- Underestimating archetype mapping time — Engineers think this is a simple column-to-field mapping. It is not. openEHR archetypes have constraints, terminology bindings, and cardinality rules that your custom schema does not. Budget 2-3x more time than your initial estimate.
- Ignoring terminology mapping — Your custom schema might use local codes (1 = Active, 2 = Inactive). openEHR expects standard terminologies (SNOMED CT, LOINC, ICD-10). Building a terminology crosswalk is a project within the project.
- Skipping dual-write — Going directly from old to new on a cutover date. In healthcare, this is unacceptable risk. Dual-write adds 2-4 weeks but eliminates the possibility of data loss during transition.
- Treating it as a pure IT project — Without clinical informaticist involvement, you will migrate the data but lose clinical meaning. A blood pressure recorded without proper context (body position, cuff size, measurement method) is less useful than the original custom field.
After Migration: What Changes
Once running on openEHR, your relationship with clinical data changes fundamentally:
- Schema evolution becomes governed — New clinical concepts are added by selecting or creating archetypes, not by adding columns. This sounds slower, but it prevents the schema sprawl that made migration necessary in the first place.
- Querying moves to AQL — The Archetype Query Language replaces SQL for clinical queries. AQL is powerful but different. Your team needs training. The openEHR AQL specification is the definitive reference.
- Interoperability becomes native — openEHR data can be exposed via FHIR without complex mapping layers. Most CDRs include built-in FHIR facades.
- Vendor independence — Your data model is now based on an open standard. You can switch CDR vendors (EHRbase, Better, Nedap) without re-modeling your data.
Need expert help with healthcare data integration? Explore our Healthcare Interoperability Solutions to see how we connect systems seamlessly. Talk to our team to get started.
Frequently Asked QuestionsCan we migrate incrementally instead of all at once?
Yes, and you should. Start with the simplest, highest-volume data (vitals, lab results) to build pipeline confidence, then tackle complex concepts (medications, clinical notes) in later phases. Each data domain can follow its own timeline through the five phases described above.
Do we need to stop using our custom EHR during migration?
No. The dual-write architecture allows your existing application to continue operating normally throughout the migration. End users should not notice any change until you are ready for the final read-path cutover.
What happens to our existing reports and analytics?
Reports built on SQL queries need to be rewritten in AQL or fed by a CQRS read model. Budget time for report migration as a separate workstream. Many organizations maintain a SQL-based reporting database alongside the CDR for complex analytical queries.
How do we handle historical data that predates our current schema?
Legacy data imported from even older systems is often the lowest quality. Consider migrating only data from the last 5-7 years at full fidelity, with older data archived in a simplified format or kept in the original system as read-only.
Is openEHR the right choice, or should we consider FHIR as our primary store?
FHIR and openEHR serve different purposes. FHIR is an interoperability standard optimized for data exchange. openEHR is a clinical data persistence standard optimized for storage, querying, and clinical modeling. Most modern health IT architectures use both: openEHR as the system of record, FHIR as the integration layer. See our guide on running SMART apps against an openEHR backend for more detail.
Conclusion
Migrating from a custom EHR schema to openEHR is a significant undertaking, but it is a well-understood problem. The critical success factors are: invest heavily in archetype mapping with clinical expertise, build a robust transformation pipeline with proper error handling, use dual-write to eliminate cutover risk, and validate clinically, not just technically.
The payoff is substantial. You move from maintaining a bespoke clinical data model — a full-time job that grows more expensive every year — to an open, governed, interoperable standard backed by a global community. Your data becomes portable, queryable in standard ways, and ready for the next generation of health IT integrations.
If you are evaluating this migration and need guidance on the archetype mapping phase, reach out to our team. We have supported multiple organizations through this transition and can help you avoid the common pitfalls.