Why Clinical Models Need Compression for Edge Deployment
Your chest X-ray classification model achieves state-of-the-art accuracy with a ResNet-152 architecture: 60 million parameters, 240MB model file, requiring 4GB GPU memory for inference. Now deploy it to a Jetson Nano with 4GB shared RAM at a rural clinic bedside. Or to a smartphone app for dermatology screening in a refugee camp with no internet. Or to an embedded sensor monitoring ICU vital signs with 1MB of flash storage.
The model physically cannot run on these devices. Model compression bridges this gap — reducing model size by 75-96% while preserving the clinical accuracy that matters. According to a 2021 survey on model compression, properly applied compression techniques can reduce model size by 10-100x with less than 2% accuracy degradation on standard benchmarks. The question for healthcare is: how much accuracy loss is acceptable, and for which clinical tasks?
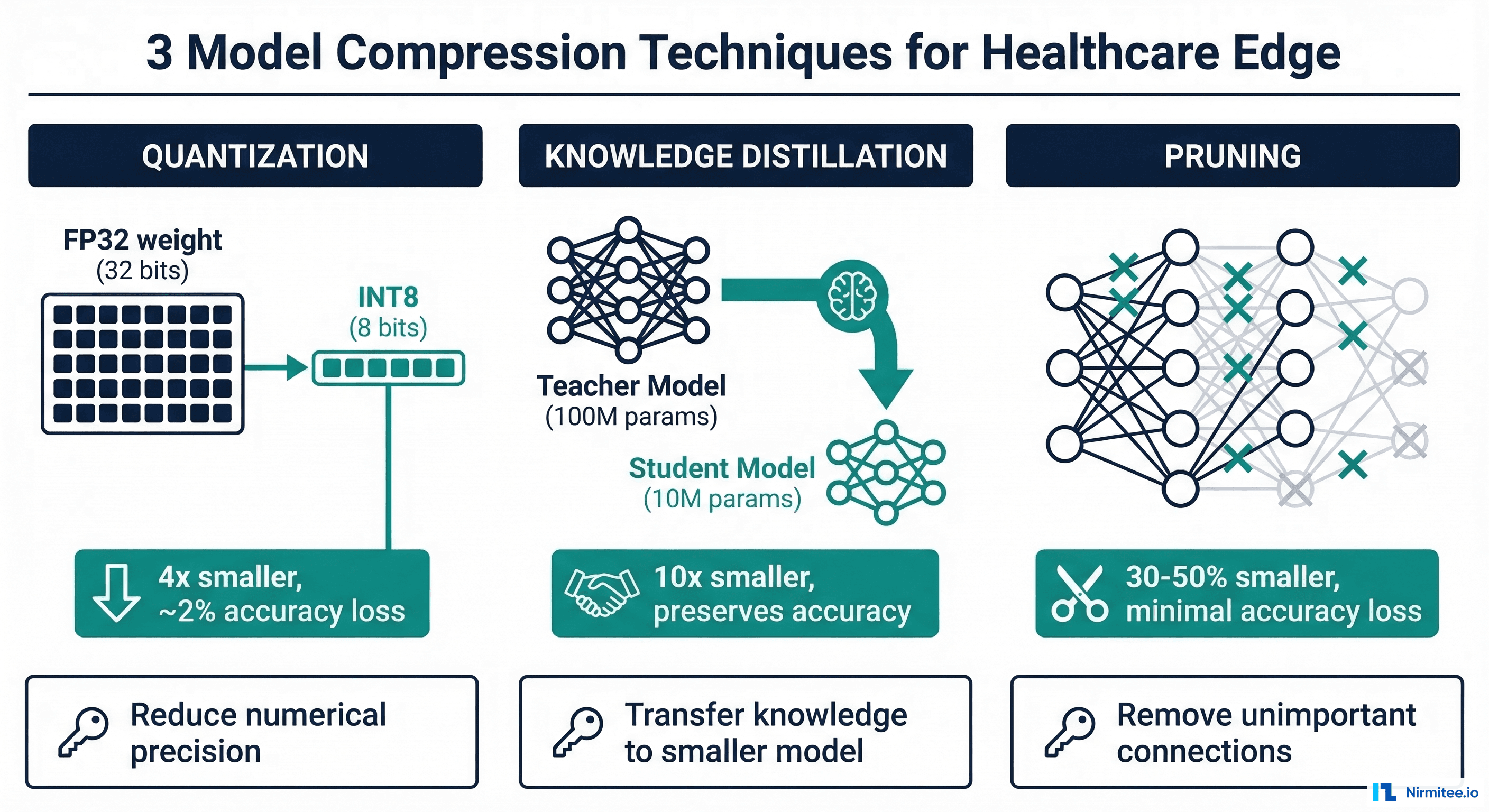
This guide covers the three primary compression techniques — quantization, knowledge distillation, and structured pruning — with healthcare-specific considerations, PyTorch implementation code, and before/after benchmark results. For deploying compressed models to specific runtimes, see our companion guide on ONNX vs TensorRT vs TFLite.
Technique 1: Quantization
Quantization reduces the numerical precision of model weights and activations from 32-bit floating point (FP32) to lower bit-widths — typically FP16 (16-bit) or INT8 (8-bit). This yields immediate benefits: 2-4x smaller model size, 2-4x faster inference on hardware with INT8 support, and reduced memory bandwidth requirements.
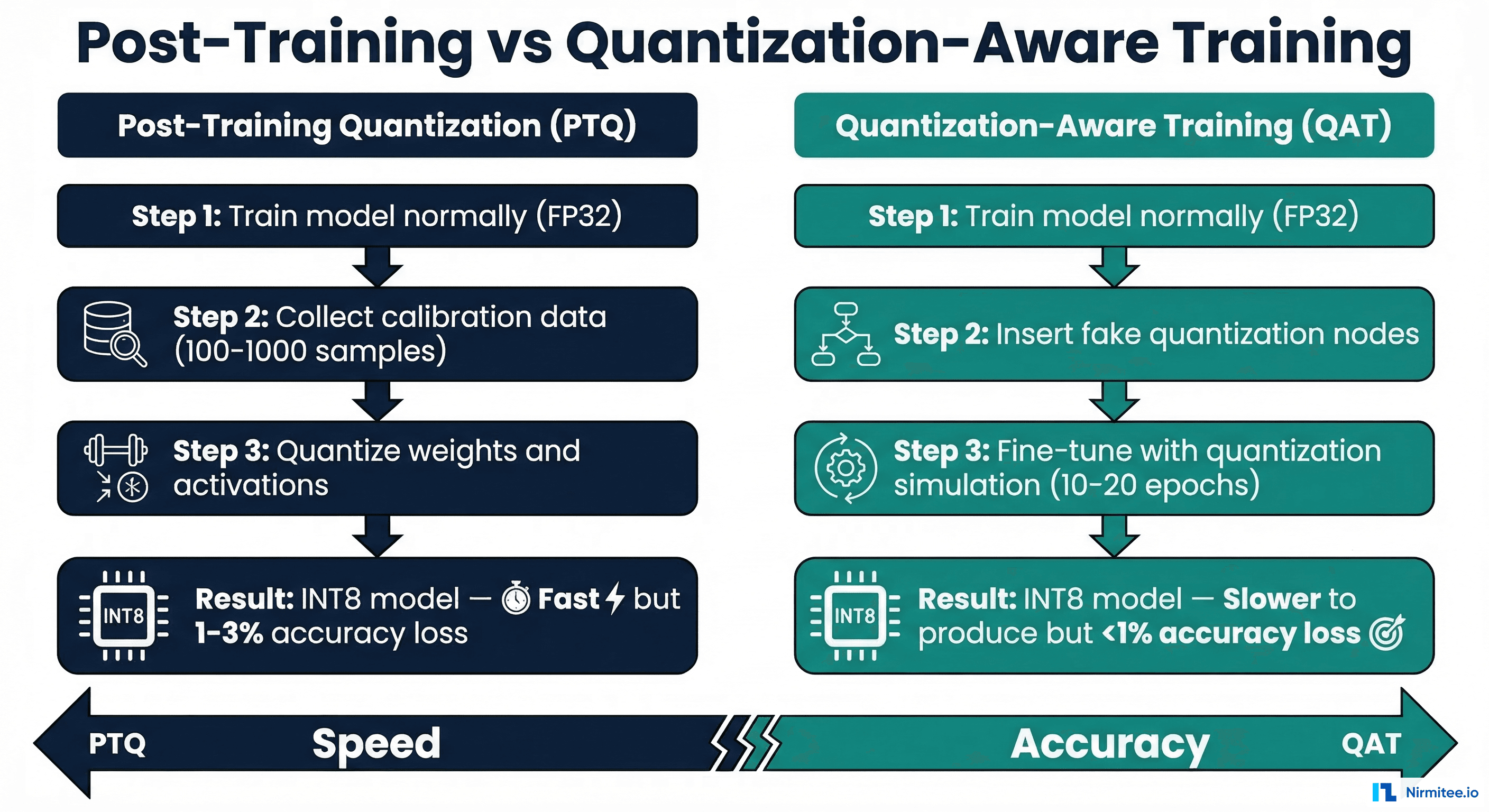
Post-Training Quantization (PTQ)
PTQ is the simplest approach: train your model normally at FP32, then convert weights to INT8 after training using a small calibration dataset. No retraining required.
import torch
from torch.quantization import quantize_dynamic, quantize_fx
from torch.ao.quantization import get_default_qconfig_mapping
import copy
class ClinicalModelQuantizer:
"""Quantize clinical models with accuracy validation."""
def __init__(self, model, calibration_loader,
validation_loader,
accuracy_threshold: float = 0.95):
self.model = model
self.calibration_loader = calibration_loader
self.validation_loader = validation_loader
self.accuracy_threshold = accuracy_threshold
def dynamic_quantization(self):
"""Dynamic quantization — simplest, no calibration needed.
Best for: LSTM, Transformer models (NLP)."""
quantized = quantize_dynamic(
copy.deepcopy(self.model),
{torch.nn.Linear, torch.nn.LSTM},
dtype=torch.qint8
)
return quantized
def static_quantization(self):
"""Static quantization — requires calibration data.
Best for: CNN models (imaging)."""
model_copy = copy.deepcopy(self.model)
model_copy.cpu()
qconfig_mapping = get_default_qconfig_mapping("x86")
model_prepared = quantize_fx.prepare_fx(
model_copy, qconfig_mapping,
example_inputs=torch.randn(1, 3, 224, 224)
)
# Run calibration data through the model
model_prepared.set_mode_to_eval()
with torch.no_grad():
for batch_idx, (data, _) in enumerate(
self.calibration_loader
):
model_prepared(data.cpu())
if batch_idx >= 100:
break
quantized = quantize_fx.convert_fx(model_prepared)
return quantized
def validate_accuracy(self, original_model,
quantized_model) -> dict:
"""Validate that quantization preserves clinical accuracy."""
original_correct = 0
quantized_correct = 0
agreement = 0
total = 0
with torch.no_grad():
for data, target in self.validation_loader:
orig_pred = original_model(data).argmax(dim=1)
quant_pred = quantized_model(data.cpu()).argmax(dim=1)
original_correct += (orig_pred == target).sum().item()
quantized_correct += (
quant_pred == target.cpu()
).sum().item()
agreement += (orig_pred.cpu() == quant_pred).sum().item()
total += target.size(0)
orig_acc = original_correct / total
quant_acc = quantized_correct / total
return {
"original_accuracy": round(orig_acc, 4),
"quantized_accuracy": round(quant_acc, 4),
"accuracy_drop": round(orig_acc - quant_acc, 4),
"accuracy_drop_pct": round(
(orig_acc - quant_acc) / orig_acc * 100, 2
),
"agreement_rate": round(agreement / total, 4),
"meets_threshold": quant_acc >= self.accuracy_threshold,
}Quantization-Aware Training (QAT)
QAT inserts simulated quantization operations during training, allowing the model to learn to compensate for quantization error. It preserves 0.3-1% more accuracy than PTQ, but requires 10-20 additional training epochs.
import torch
from torch.ao.quantization import (
get_default_qat_qconfig_mapping,
prepare_qat_fx,
convert_fx
)
def quantization_aware_training(
model, train_loader, val_loader,
num_epochs: int = 15,
learning_rate: float = 1e-4
):
"""Fine-tune model with quantization-aware training."""
model.train()
qconfig_mapping = get_default_qat_qconfig_mapping("x86")
example_inputs = torch.randn(1, 3, 224, 224)
model_prepared = prepare_qat_fx(
model, qconfig_mapping,
example_inputs=example_inputs
)
optimizer = torch.optim.Adam(
model_prepared.parameters(), lr=learning_rate
)
criterion = torch.nn.CrossEntropyLoss()
for epoch in range(num_epochs):
model_prepared.train()
running_loss = 0.0
for batch_idx, (data, target) in enumerate(train_loader):
optimizer.zero_grad()
output = model_prepared(data)
loss = criterion(output, target)
loss.backward()
optimizer.step()
running_loss += loss.item()
# Freeze quantization parameters after 60% of training
if epoch >= int(num_epochs * 0.6):
model_prepared.apply(
torch.ao.quantization.disable_observer
)
print(f"Epoch {epoch+1}/{num_epochs}, "
f"Loss: {running_loss/len(train_loader):.4f}")
model_prepared.set_mode_to_eval()
quantized_model = convert_fx(model_prepared)
return quantized_modelTechnique 2: Knowledge Distillation
Knowledge distillation trains a small "student" model to mimic the behavior of a large "teacher" model. Instead of learning from hard labels (ground truth), the student learns from the teacher's soft probability distributions — capturing the teacher's nuanced understanding of which classes are similar, which features matter, and what uncertainty looks like.
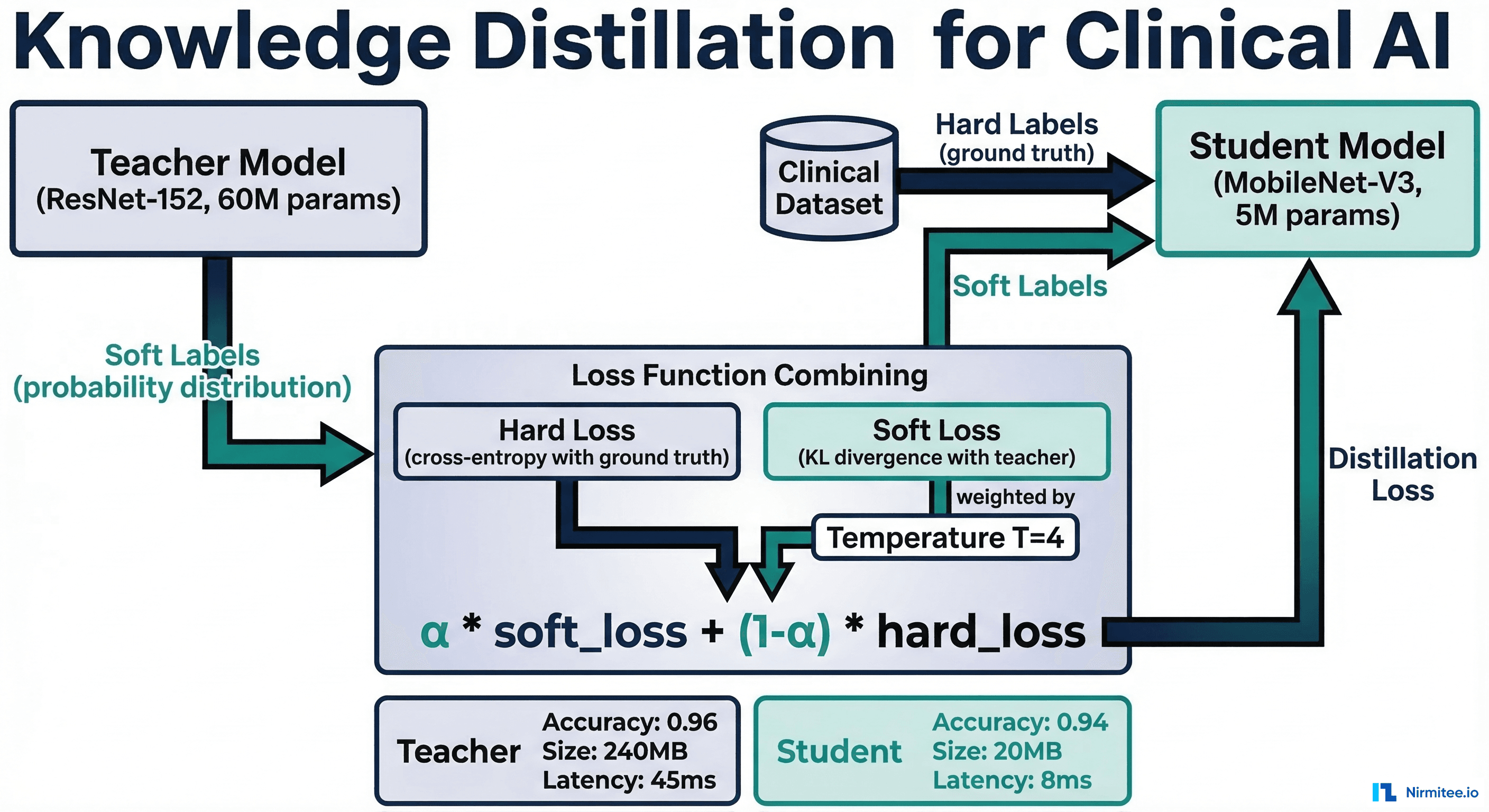
This is particularly valuable in healthcare where data quality and labeling are expensive: the teacher model captures relationships that hard labels miss (e.g., a chest X-ray that is "probably normal but has a subtle finding worth noting" — the soft probability distribution conveys this ambiguity).
import torch
import torch.nn as nn
import torch.nn.functional as F
class ClinicalDistillationTrainer:
"""Knowledge distillation for healthcare models."""
def __init__(self, teacher: nn.Module, student: nn.Module,
temperature: float = 4.0, alpha: float = 0.7):
self.teacher = teacher
self.student = student
self.temperature = temperature
self.alpha = alpha
# Freeze teacher
self.teacher.set_mode_to_eval()
for param in self.teacher.parameters():
param.requires_grad = False
def distillation_loss(self, student_logits, teacher_logits,
targets):
"""Combined distillation + hard label loss."""
T = self.temperature
# Soft loss: KL divergence between student and teacher
soft_loss = F.kl_div(
F.log_softmax(student_logits / T, dim=1),
F.softmax(teacher_logits / T, dim=1),
reduction="batchmean"
) * (T * T)
# Hard loss: standard cross-entropy with ground truth
hard_loss = F.cross_entropy(student_logits, targets)
return self.alpha * soft_loss + (1 - self.alpha) * hard_loss
def train(self, train_loader, val_loader,
num_epochs: int = 50,
lr: float = 1e-3) -> dict:
"""Run distillation training loop."""
optimizer = torch.optim.Adam(
self.student.parameters(), lr=lr
)
scheduler = torch.optim.lr_scheduler.CosineAnnealingLR(
optimizer, T_max=num_epochs
)
best_val_acc = 0.0
for epoch in range(num_epochs):
self.student.train()
epoch_loss = 0.0
for data, targets in train_loader:
with torch.no_grad():
teacher_logits = self.teacher(data)
student_logits = self.student(data)
loss = self.distillation_loss(
student_logits, teacher_logits, targets
)
optimizer.zero_grad()
loss.backward()
optimizer.step()
epoch_loss += loss.item()
scheduler.step()
val_acc = self._validate(val_loader)
if val_acc > best_val_acc:
best_val_acc = val_acc
torch.save(self.student.state_dict(),
"best_student.pt")
print(f"Epoch {epoch+1}/{num_epochs} "
f"Loss: {epoch_loss/len(train_loader):.4f} "
f"Val Acc: {val_acc:.4f}")
return {
"best_val_accuracy": best_val_acc,
"teacher_params": sum(
p.numel() for p in self.teacher.parameters()
),
"student_params": sum(
p.numel() for p in self.student.parameters()
),
}
def _validate(self, val_loader) -> float:
self.student.set_mode_to_eval()
correct = total = 0
with torch.no_grad():
for data, targets in val_loader:
preds = self.student(data).argmax(dim=1)
correct += (preds == targets).sum().item()
total += targets.size(0)
return correct / totalTechnique 3: Structured Pruning
Structured pruning removes entire filters, channels, or attention heads from a neural network — not individual weights. This produces genuinely smaller models that run faster on standard hardware without requiring specialized sparse-computation support.
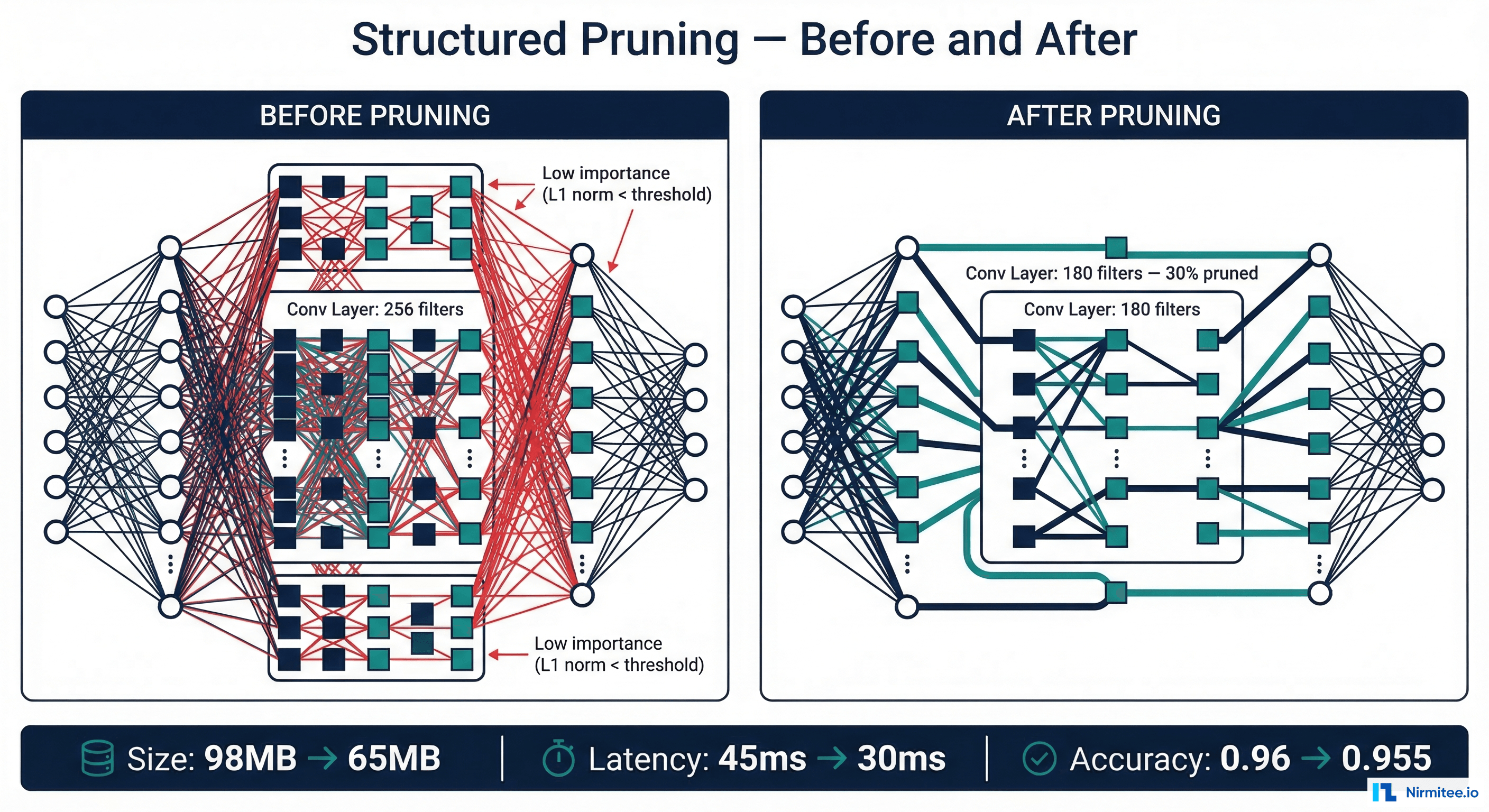
import torch
import torch.nn.utils.prune as prune
import torch.nn as nn
from typing import List, Tuple
class ClinicalModelPruner:
"""Structured pruning for healthcare CNN models."""
def __init__(self, model: nn.Module,
prune_ratio: float = 0.3):
self.model = model
self.prune_ratio = prune_ratio
def get_prunable_layers(self) -> List[Tuple]:
"""Identify convolutional layers for pruning."""
layers = []
for name, module in self.model.named_modules():
if isinstance(module, nn.Conv2d):
layers.append((module, "weight"))
return layers
def prune_l1_structured(self) -> nn.Module:
"""Prune filters with lowest L1 norm."""
layers = self.get_prunable_layers()
for module, param_name in layers:
prune.ln_structured(
module, param_name,
amount=self.prune_ratio,
n=1, # L1 norm
dim=0 # Prune output channels (filters)
)
return self.model
def compute_sparsity(self) -> dict:
"""Report pruning statistics."""
total_params = 0
zero_params = 0
for param in self.model.parameters():
total_params += param.numel()
zero_params += (param == 0).sum().item()
return {
"total_parameters": total_params,
"zero_parameters": zero_params,
"sparsity": round(zero_params / total_params, 4),
}
def iterative_prune_with_finetuning(
self, train_loader, val_loader,
prune_steps: int = 5,
finetune_epochs: int = 3,
lr: float = 1e-4
) -> dict:
"""Gradually prune and fine-tune to preserve accuracy."""
step_ratio = self.prune_ratio / prune_steps
results = []
for step in range(prune_steps):
layers = self.get_prunable_layers()
for module, param_name in layers:
prune.ln_structured(
module, param_name,
amount=step_ratio,
n=1, dim=0
)
optimizer = torch.optim.Adam(
self.model.parameters(), lr=lr
)
criterion = nn.CrossEntropyLoss()
for epoch in range(finetune_epochs):
self.model.train()
for data, target in train_loader:
optimizer.zero_grad()
output = self.model(data)
loss = criterion(output, target)
loss.backward()
optimizer.step()
sparsity = self.compute_sparsity()
print(f"Step {step+1}/{prune_steps}: "
f"Sparsity={sparsity['sparsity']:.2%}")
results.append(sparsity)
return resultsHealthcare-Specific Considerations
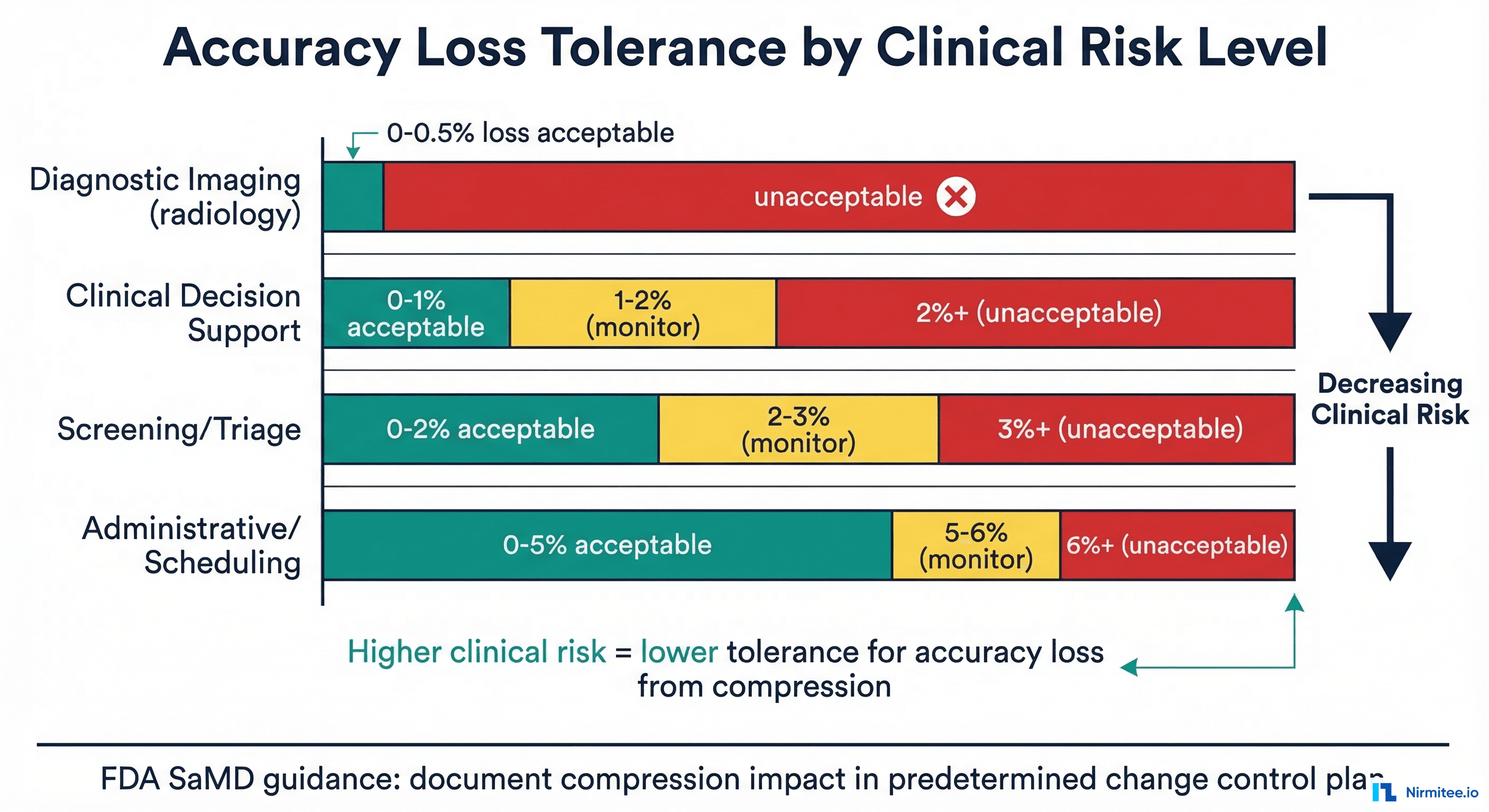
The acceptable accuracy loss from compression depends entirely on the clinical context. A screening tool that triages cases for radiologist review can tolerate more accuracy loss than a diagnostic tool that directly influences treatment decisions. This risk-based approach aligns with FDA SaMD guidance on predetermined change control plans.
| Clinical Risk Level | Example | Max Accuracy Loss | Recommended Technique |
|---|---|---|---|
| Critical (diagnostic) | Diabetic retinopathy grading | <0.5% | QAT only — never PTQ alone |
| High (treatment-affecting) | Drug interaction detection | <1% | QAT + iterative pruning with validation |
| Medium (screening/triage) | Chest X-ray triage | <2% | Distillation + QAT |
| Low (informational) | Chart summarization | <5% | Aggressive distillation + INT4 quantization |
Beyond accuracy, compression can disproportionately affect fairness across demographic subgroups. Research from MIT has shown that quantization can increase the accuracy gap between majority and minority groups by up to 2x. Always validate fairness metrics after compression using the monitoring approaches described in our guide on model monitoring for healthcare AI.
Benchmark Results: Before and After Compression
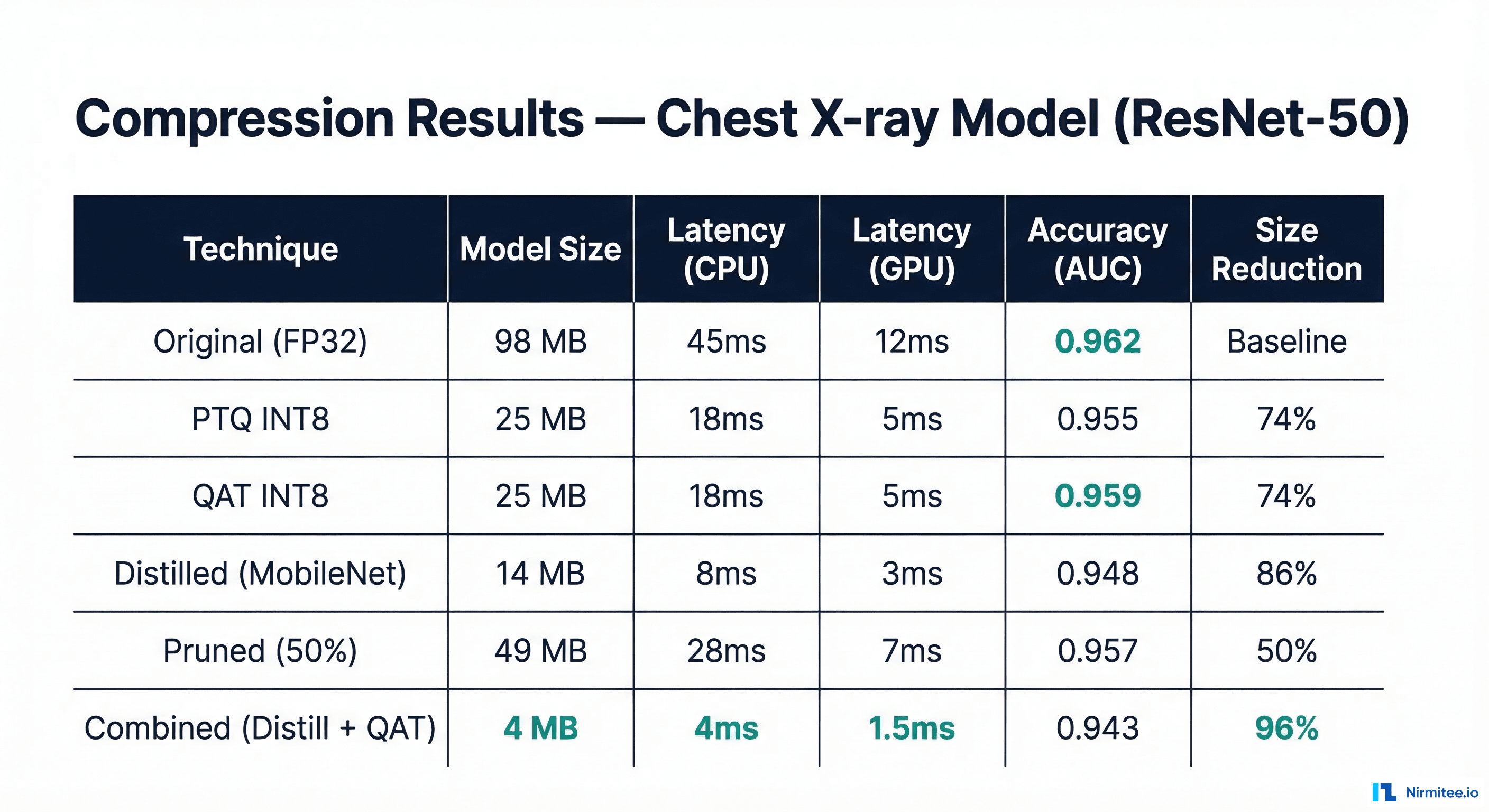
| Technique | Model Size | CPU Latency | GPU Latency | Accuracy (AUC) | Size Reduction |
|---|---|---|---|---|---|
| Original (FP32) | 98 MB | 45ms | 12ms | 0.962 | Baseline |
| PTQ INT8 | 25 MB | 18ms | 5ms | 0.955 (-0.7%) | 74% |
| QAT INT8 | 25 MB | 18ms | 5ms | 0.959 (-0.3%) | 74% |
| Distilled (MobileNet-V3) | 14 MB | 8ms | 3ms | 0.948 (-1.5%) | 86% |
| Pruned 50% | 49 MB | 28ms | 7ms | 0.957 (-0.5%) | 50% |
| Distill + QAT (Combined) | 4 MB | 4ms | 1.5ms | 0.943 (-2.0%) | 96% |
The Compression Pipeline
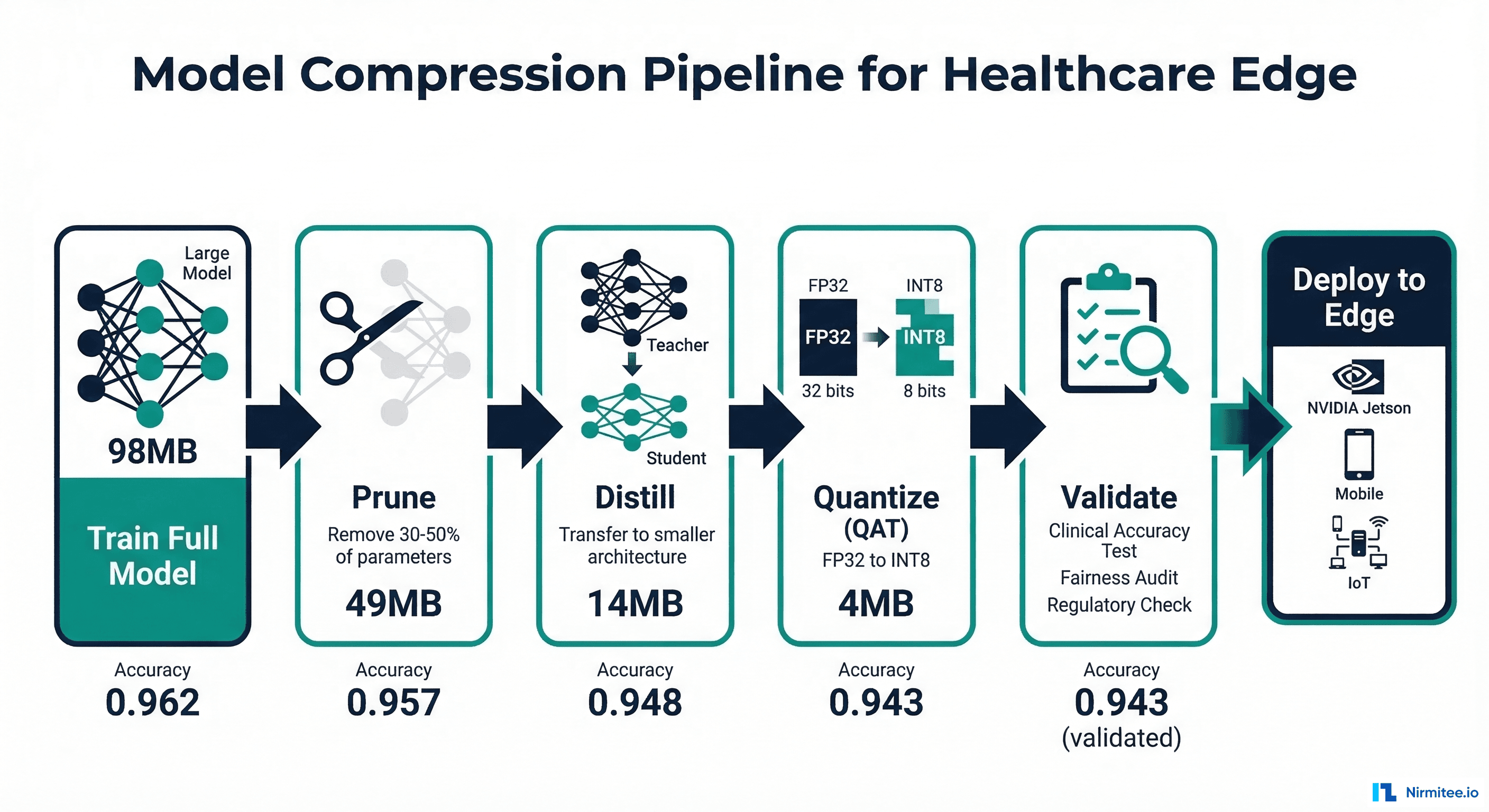
The techniques can be combined for maximum compression. The recommended order is: prune first (remove structural redundancy), then distill (transfer knowledge to smaller architecture), then quantize (reduce precision). Each step builds on the previous, and accuracy is validated at each gate before proceeding. For deploying the final compressed model, see our guide on choosing the right inference runtime.
Target Edge Hardware
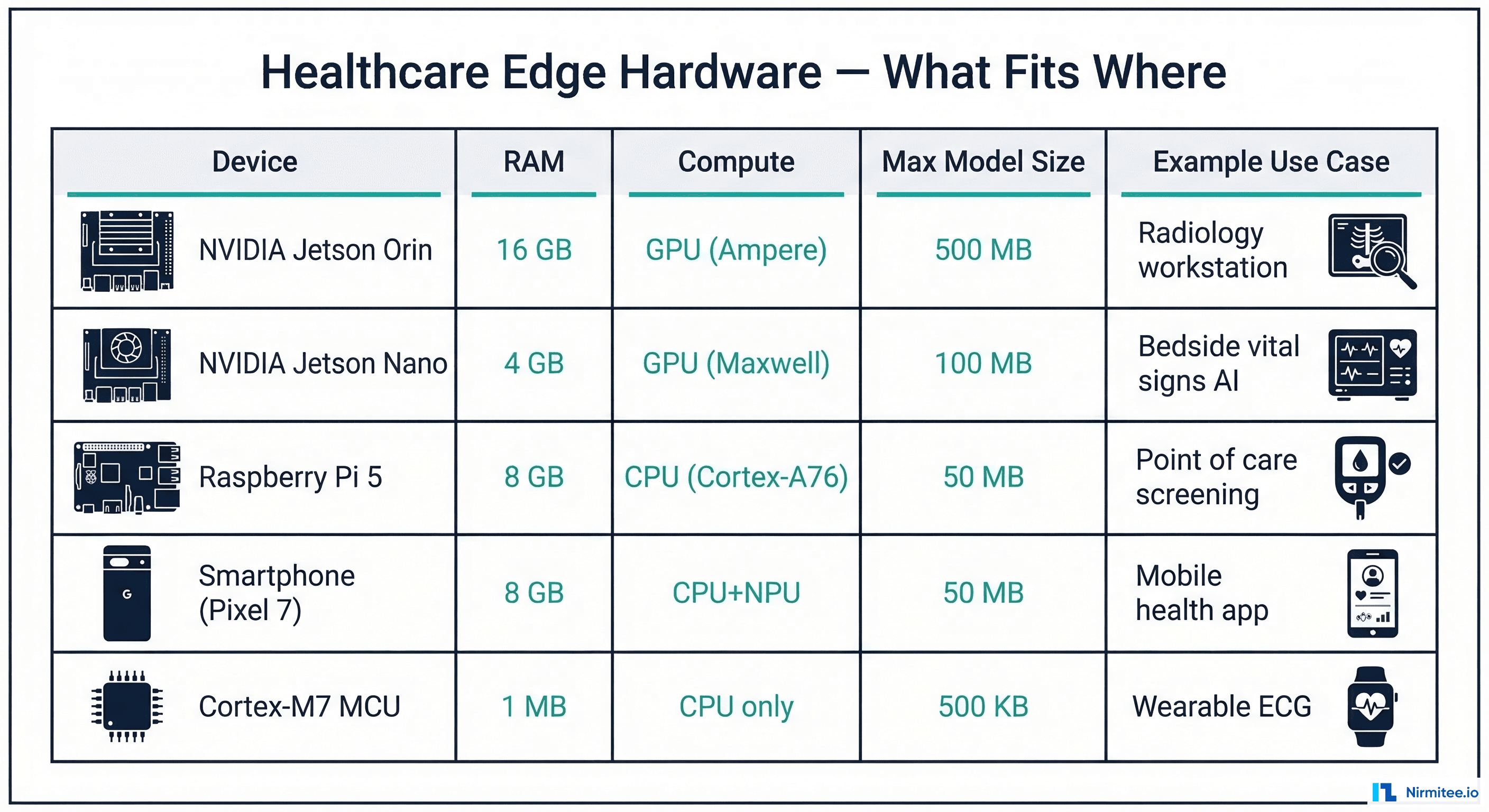
| Device | RAM | Compute | Max Model Size | Best Compression | Healthcare Use Case |
|---|---|---|---|---|---|
| NVIDIA Jetson Orin | 16 GB | GPU (Ampere) | 500 MB | TensorRT FP16 | Radiology workstation, surgical navigation |
| NVIDIA Jetson Nano | 4 GB | GPU (Maxwell) | 100 MB | TensorRT INT8 | Bedside vital signs AI, point-of-care |
| Raspberry Pi 5 | 8 GB | CPU (Cortex-A76) | 50 MB | ONNX INT8 | Remote clinic screening |
| Smartphone | 6-8 GB | CPU + NPU | 50 MB | TFLite INT8 | Mobile dermatology, medication tracking |
| Cortex-M7 MCU | 1 MB | CPU only | 500 KB | TFLite Micro INT8 | Wearable ECG, glucose monitor |
Frequently Asked Questions
Should I quantize, distill, or prune first?
The recommended order is: prune (remove structural redundancy while the model is still at full precision), then distill (transfer to a smaller architecture), then quantize (reduce precision). Pruning at full precision is more stable because gradient information is intact. Distillation after pruning captures the pruned model's behavior. Quantization is always the final step because it can be applied to any model architecture.
How do I validate that a compressed model is safe for clinical use?
Run the same validation suite you used for the original model: accuracy on your test set, fairness metrics across demographic subgroups, calibration analysis, and stress testing with edge cases. Additionally, compare the compressed model's disagreements with the original — any systematic pattern (e.g., the compressed model consistently misses subtle findings) is a safety concern. Use the shadow deployment strategy to validate the compressed model against the original in production before promoting it.
Does compression affect model fairness?
Yes. Research has shown that quantization and pruning can disproportionately degrade performance on underrepresented subgroups. The reason: minority-group features often have lower activation magnitudes, which are more likely to be quantized to zero or pruned as "unimportant." Always measure fairness metrics before and after compression, and consider fairness-aware compression techniques that constrain the optimization to maintain equitable performance.
What is the smallest model that can run clinical NLP tasks?
DistilBERT (66M parameters, ~250MB FP32) is the practical minimum for clinical NLP tasks requiring contextual understanding. Distilled further to TinyBERT (14M parameters, ~60MB), it can run on mobile devices. For keyword-based clinical NLP (ICD coding, simple entity extraction), simpler models like TF-IDF + logistic regression can run at ~1MB. For transformer models on microcontrollers, consider specialized architectures like EdgeBERT or MobileBERT.
Can I compress a model after it has received FDA clearance?
This depends on your predetermined change control plan. If your clearance documentation specifies the exact model architecture and precision, compression would constitute a modification requiring a new 510(k) or De Novo submission. If your plan includes compression as a "predetermined change" with validation criteria, you can compress and validate without resubmission. The 2024 FDA guidance on predetermined change control plans specifically addresses this scenario.
How much compression is "too much" for clinical safety?
There is no universal answer — it depends on clinical risk level. For diagnostic models (e.g., detecting cancerous lesions), even 0.5% accuracy drop may be unacceptable if it translates to missed diagnoses. For screening tools, 2% accuracy loss might be acceptable if it enables deployment to devices that otherwise could not run the model at all, bringing AI-assisted screening to populations that currently have no access. The decision should involve clinical stakeholders, not just ML engineers. Document the trade-off analysis as part of your SaMD quality management system.