A growing number of healthcare CIOs now cite AI as a primary driver for evaluating openEHR. Not just interoperability. Not just regulatory compliance. Not just vendor independence. AI.
That shift surprises people who think of openEHR as "just another clinical data standard." But the CIOs who have lived through the reality of building ML pipelines on traditional EHR data understand something the rest of the industry is still learning: the architecture of your clinical data repository determines whether AI is a six-month science project or a two-week engineering task.
This blog explains exactly why openEHR's archetype-based modeling produces data that is structurally, semantically, and operationally ready for machine learning — and shows you the concrete engineering patterns that make it work.
The 80% Problem: Why Clinical AI Projects Stall
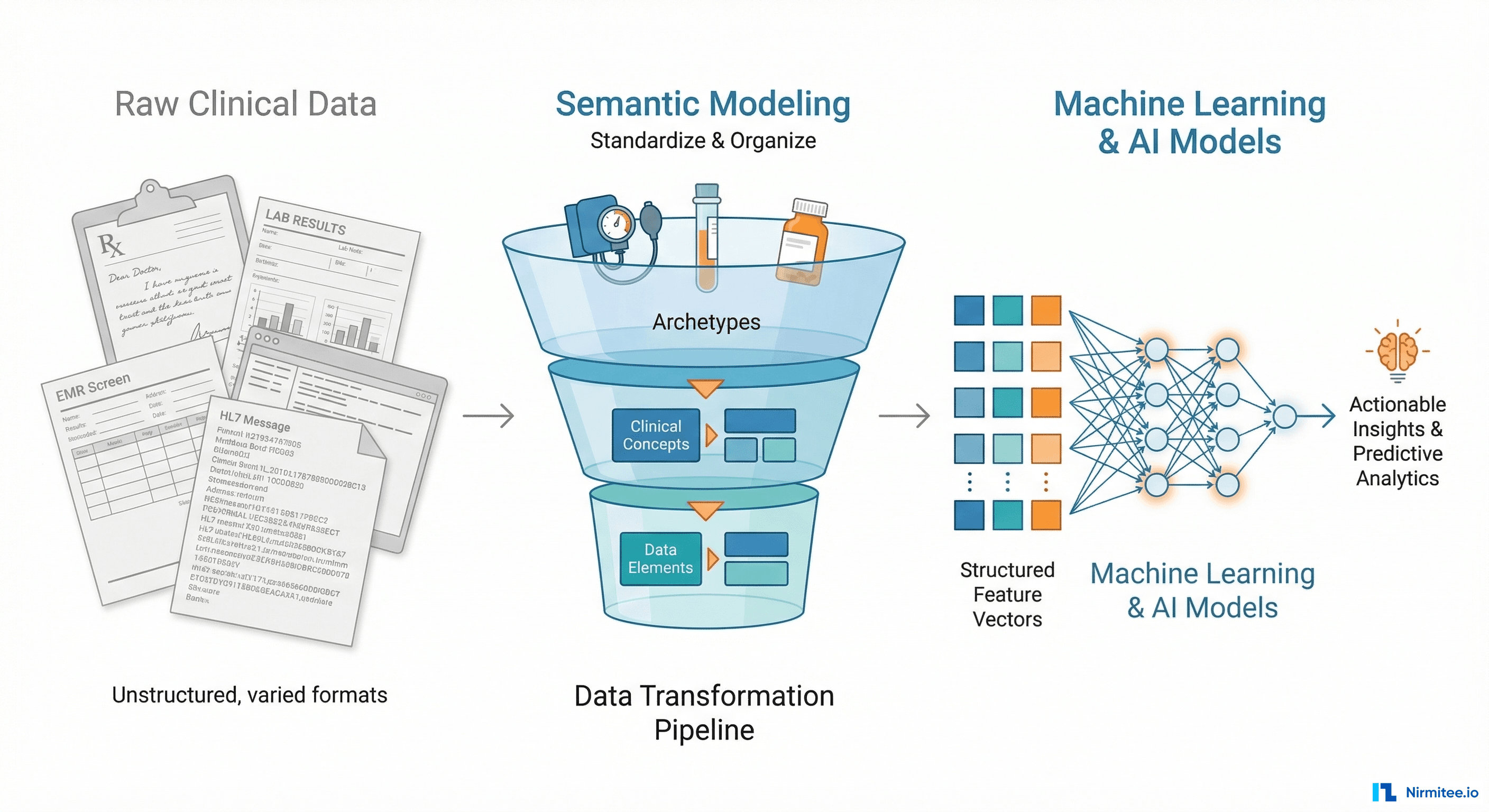
Every data scientist who has worked in healthcare knows the number: 80% of an ML project's time goes to data preparation, not model building. In clinical settings, that number is often higher.
Here is what a typical ML data preparation pipeline looks like when you are working with a conventional EHR system:
- Data extraction — Reverse-engineering proprietary database schemas, writing custom ETL jobs for each EHR vendor, navigating access restrictions and audit requirements. Timeline: 4-6 weeks.
- Schema mapping — The same clinical concept (blood pressure, for instance) is stored differently across Epic, Cerner, MEDITECH, and every other system. You need a mapping layer. Timeline: 3-4 weeks.
- Semantic normalization — Even after mapping, the data lacks consistent meaning. One system stores blood pressure in mmHg, another in kPa. One records position as "sitting," another as code "33586001." You write conversion logic for every variation. Timeline: 6-8 weeks.
- Quality cleaning — Missing values, duplicate records, impossible physiological values (systolic of 900?), inconsistent timestamps. Each requires domain-specific rules. Timeline: 4-6 weeks.
- Feature engineering — Finally, you can start creating the features your model actually needs. But because the upstream data was messy, your feature pipelines are fragile and break when data patterns shift. Timeline: 6-8 weeks.
Total: 23-32 weeks before you train your first model. And when you want to deploy the same model at a second hospital with a different EHR vendor, you start most of this work over.
This is not a tooling problem. It is an architecture problem. The data was never structured for computational consumption. It was structured for human documentation and billing workflows.
What Makes openEHR Different: Archetypes as Computable Contracts
openEHR introduces a concept that changes everything for AI: the archetype. An archetype is not just a data format. It is a computable contract that defines the complete semantic structure of a clinical concept.
Take the blood pressure archetype (openEHR-EHR-OBSERVATION.blood_pressure.v2). It defines:
- Every data point — systolic, diastolic, mean arterial pressure, pulse pressure, with exact data types (DV_QUANTITY with units)
- Context metadata — patient position (sitting, standing, lying), cuff size, measurement location (left arm, right arm), device used
- Temporal structure — when the measurement was taken, whether it is a single reading or an average of multiple readings, the protocol followed
- Value constraints — systolic must be between 0-1000 mmHg, position must come from a defined value set
- Terminology bindings — each element maps to SNOMED CT, LOINC, or other standard terminologies
This is not metadata bolted on after the fact. It is the structure of the data itself. Every blood pressure recorded in any openEHR system worldwide conforms to this same archetype. The systolic value is always at the same path, with the same type, the same units, the same constraints.
For a data scientist, this means: zero schema discovery, zero semantic mapping, zero unit conversion. You know exactly where the data is, what it means, and how to use it — before you write a single line of code.
The Two-Level Modeling Advantage
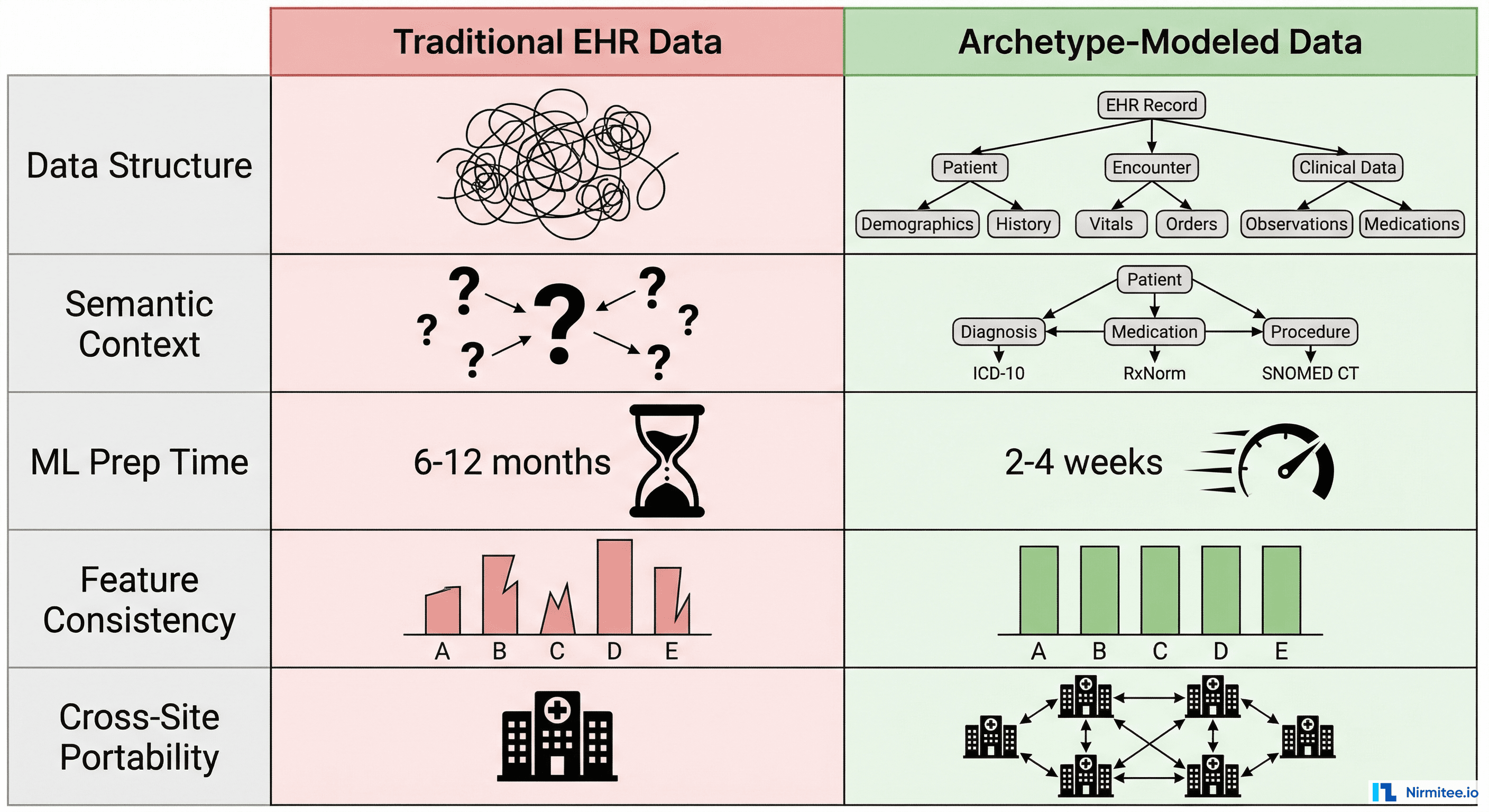
openEHR separates the reference model (how data is stored technically) from the archetype model (what clinical concepts mean). This two-level approach creates three properties that are uniquely valuable for AI:
- Semantic stability — Archetypes evolve slowly through international clinical consensus. A blood pressure archetype from 2019 is structurally compatible with one from 2026. Your ML pipelines do not break when the platform upgrades.
- Cross-site consistency — Every openEHR deployment uses the same archetypes from the international Clinical Knowledge Manager (CKM). A model trained on data from Hospital A can process data from Hospital B without re-engineering.
- Type safety at the data layer — Archetype constraints prevent invalid data from entering the system. You do not need to write defensive code to handle impossible values or type mismatches in your feature pipeline.

From Archetypes to Features: The Engineering Pipeline
Understanding why openEHR data is AI-ready in theory is one thing. Building the actual pipeline is another. Here is how the archetype-to-feature engineering pipeline works in practice.
Step 1: AQL — The Query Language Designed for Clinical Semantics
openEHR includes its own query language, AQL (Archetype Query Language), specifically designed to query clinical data using archetype paths. Unlike SQL, AQL queries are portable across any openEHR system regardless of the underlying database technology.
A typical AQL query for extracting blood pressure features:
SELECT
e/ehr_id/value as patient_id,
o/data[at0001]/events[at0006]/time/value as timestamp,
o/data[at0001]/events[at0006]/data[at0003]/items[at0004]/value/magnitude
as systolic,
o/data[at0001]/events[at0006]/data[at0003]/items[at0005]/value/magnitude
as diastolic,
o/state[at0007]/items[at1030]/value/defining_code/code_string
as position
FROM EHR e
CONTAINS OBSERVATION o[
openEHR-EHR-OBSERVATION.blood_pressure.v2
]
WHERE
o/data[at0001]/events[at0006]/time/value > '2025-01-01'This single query, running against any openEHR CDR worldwide, returns a clean, typed, columnar dataset with consistent semantics. No joins across proprietary tables. No vendor-specific SQL. No post-processing to interpret what the columns mean.
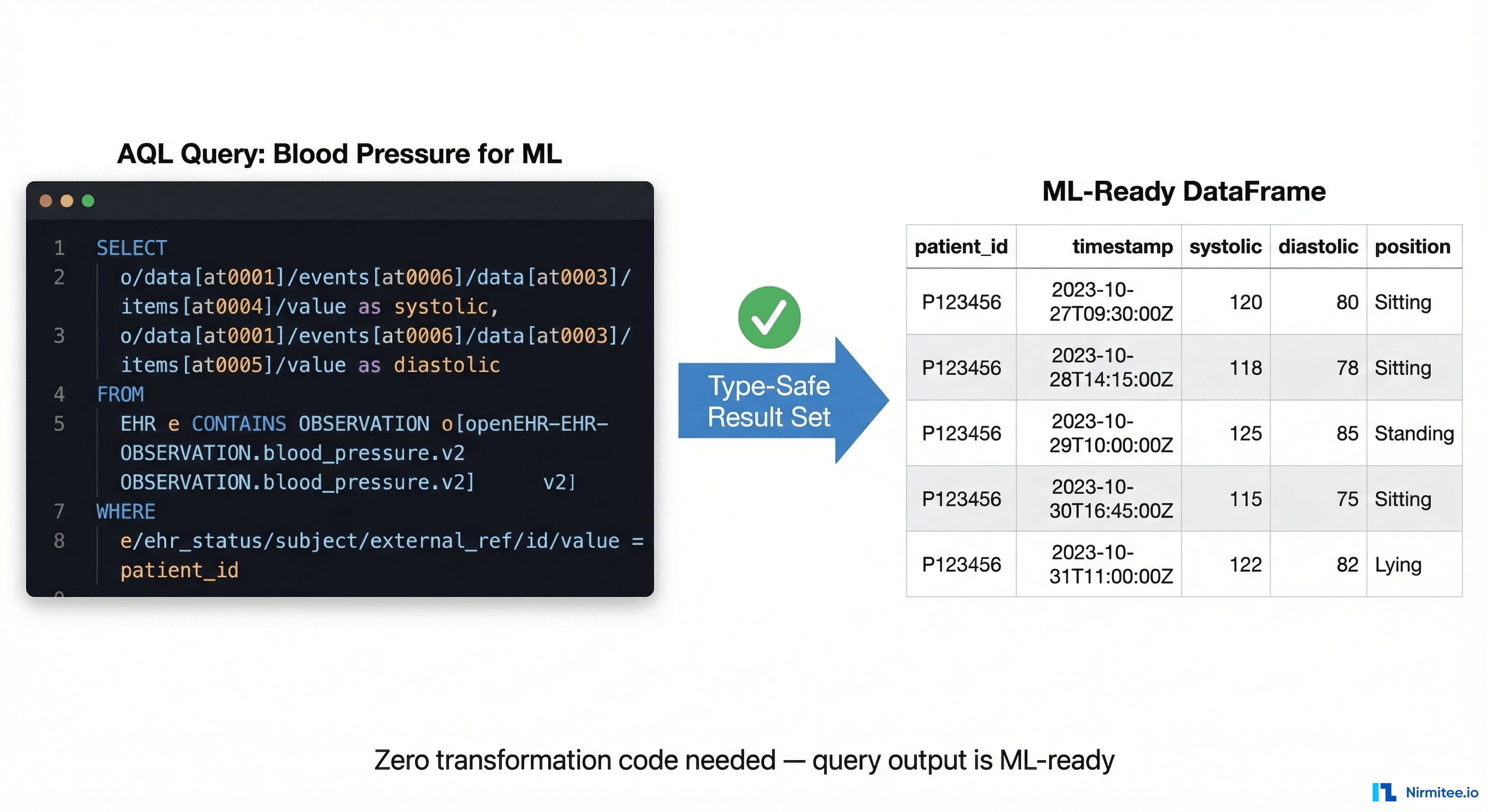
Step 2: Type-Safe Feature Extraction
Because archetypes define exact data types and constraints, feature extraction becomes deterministic. Here is a Python pattern for building features from openEHR query results:
from datetime import datetime, timedelta
import numpy as np
class BloodPressureFeatureExtractor:
def __init__(self, aql_client):
self.client = aql_client
def extract_patient_features(self, patient_id, window_days=90):
results = self.client.query(
archetype='openEHR-EHR-OBSERVATION.blood_pressure.v2',
patient_id=patient_id,
since=datetime.now() - timedelta(days=window_days)
)
# Archetypes guarantee structure, but still guard empty cases
if not results:
return self._empty_features()
systolic_values = [r.systolic for r in results]
diastolic_values = [r.diastolic for r in results]
timestamps = [r.timestamp for r in results]
pulse_pressures = [
s - d for s, d in zip(systolic_values, diastolic_values)
]
return {
'systolic_mean': np.mean(systolic_values),
'systolic_std': np.std(systolic_values),
'systolic_trend': self._linear_trend(
timestamps,
systolic_values
),
'diastolic_mean': np.mean(diastolic_values),
'pulse_pressure_mean': np.mean(pulse_pressures),
'measurement_frequency': len(results) / window_days,
'position_variance': self._position_entropy(results),
}
def _empty_features(self):
return {
'systolic_mean': None,
'systolic_std': None,
'systolic_trend': None,
'diastolic_mean': None,
'pulse_pressure_mean': None,
'measurement_frequency': 0,
'position_variance': None,
}Notice what is missing: no null-checking for unexpected types, no unit conversion, no validation that systolic is actually a number. The archetype guarantees all of this at the data layer. Your feature code is pure domain logic.
Step 3: Multi-Archetype Feature Composition
Real clinical ML models combine features from multiple archetypes. openEHR makes this composable because every archetype follows the same structural patterns:
class ClinicalFeatureComposer:
def __init__(self, aql_client):
self.aql_client = aql_client
self.extractors = {
'blood_pressure': BloodPressureFeatureExtractor,
'lab_result': LabResultFeatureExtractor,
'medication': MedicationFeatureExtractor,
'body_weight': BodyWeightFeatureExtractor,
}
def compose_features(self, patient_id):
features = {}
for name, extractor_cls in self.extractors.items():
extractor = extractor_cls(self.aql_client)
extracted = extractor.extract_patient_features(
patient_id
)
# Namespace features to avoid collisions
namespaced = {
f"{name}_{k}": v
for k, v in extracted.items()
}
features.update(namespaced)
return features # Flat dict → ready for pandas/sklearnEach extractor follows the same pattern: AQL query → typed results → domain features. Adding a new clinical concept to your model means writing one new extractor class, not re-engineering your entire data pipeline.
Quantifying the Advantage: Traditional vs. openEHR ML Pipelines
The effort reduction is not incremental. It is structural.
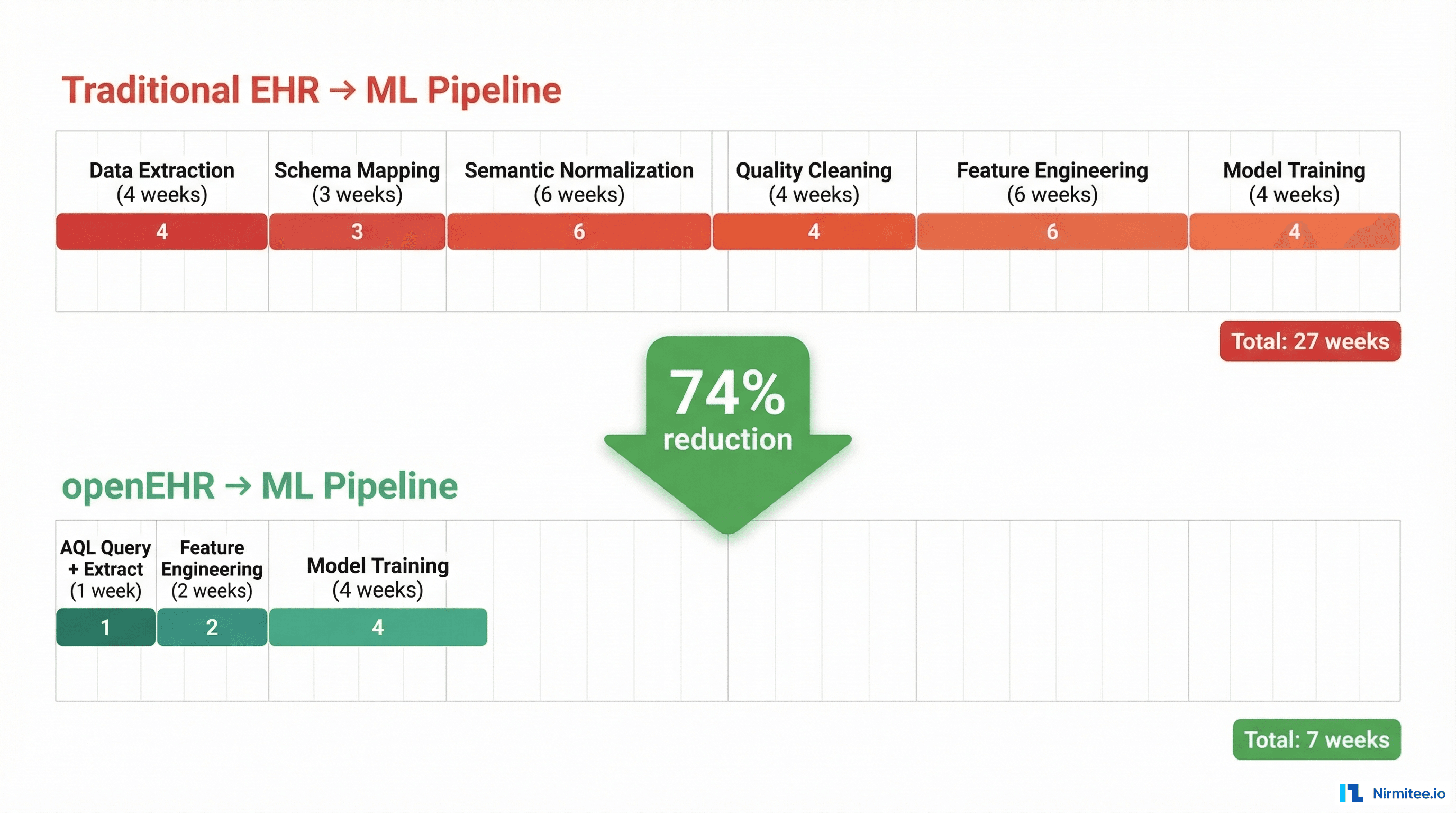
With a traditional EHR, the pipeline from raw data to trained model takes 23-32 weeks. With openEHR, the same pipeline takes 5-7 weeks. That is a 74% reduction in time-to-model.
But the bigger advantage is not the first model — it is the second, third, and tenth. Because openEHR feature extractors are portable across sites, deploying a validated model at a new hospital requires days, not months. The AQL queries work unchanged. The feature extractors produce identical output. Only the connection string changes.
Where the Time Savings Come From
| Pipeline Stage | Traditional EHR | openEHR | Why |
|---|---|---|---|
| Data extraction | 4-6 weeks | 1-2 days | AQL replaces vendor-specific ETL |
| Schema mapping | 3-4 weeks | 0 | Archetypes are the schema |
| Semantic normalization | 6-8 weeks | 0 | Terminology bindings built in |
| Quality cleaning | 4-6 weeks | 2-3 days | Archetype constraints prevent bad data |
| Feature engineering | 6-8 weeks | 2-3 weeks | Type-safe, no defensive coding |
| Model training | 4 weeks | 4 weeks | Same (model-dependent) |
The savings are concentrated in the data preparation stages — exactly where traditional healthcare AI projects waste the most time and money.
Five AI Use Cases That openEHR Unlocks
When your clinical data is AI-ready by default, use cases that were previously impractical become engineering exercises. Here are five high-impact applications that organizations are building on openEHR foundations.

1. Hypertension Risk Prediction
The blood pressure archetype provides longitudinal systolic/diastolic measurements with position context and measurement protocol metadata. Combined with the body weight and lab result archetypes, you can build a hypertension risk model that accounts for measurement conditions — not just raw numbers.
Key features enabled by archetypes: systolic trend over 6 months, measurement position consistency, pulse pressure variability, BMI trajectory correlation. Structured archetype data provides the consistent feature inputs that hypertension risk models need — longitudinal systolic trends, position-aware measurements, and pulse pressure variability — without the months of data cleaning that proprietary EHR data requires.
2. Sepsis Early Warning
Sepsis detection requires combining vital signs, lab results, and medication orders in real-time. With traditional EHRs, integrating these three data streams from different subsystems is the primary engineering challenge. With openEHR, each is a well-defined archetype queryable through a single AQL interface.
The critical advantage: temporal alignment. Archetype events have standardized timestamp structures, so correlating a temperature spike with a WBC count and an antibiotic order is a query, not an integration project. The temporal alignment that archetypes provide — correlating vital sign spikes with lab results and medication orders through a single query interface — is exactly what early warning systems need.
3. Drug Interaction Detection at Scale
The medication order archetype captures drug identity (with terminology bindings to RxNorm/dm+d), dose, route, frequency, and duration with full semantic precision. Building a drug interaction model means querying active medications per patient — a single AQL call — and running them against an interaction knowledge base.
The medication archetype delivers drug identity, dose, route, frequency, and duration in a structured format that interaction engines can consume directly — no ETL overhead to normalize vendor-specific medication schemas.
4. Automated Clinical Coding
Clinical compositions in openEHR bundle related observations, evaluations, and instructions into coherent clinical documents. Each element within a composition carries terminology bindings (SNOMED CT, ICD-10) that serve as strong signals for automated coding models.
Rather than training NLP models on free-text clinical notes, the structured archetype data with built-in terminology bindings (SNOMED CT, ICD-10) provides direct input for automated coding models — reducing reliance on free-text interpretation and improving coding consistency.
5. Disease Progression Modeling
The problem list archetype tracks diagnoses with onset dates, severity, and clinical status over time. Combined with longitudinal lab results and medication history, you can model disease trajectories — how conditions evolve, what interventions change the trajectory, and when to escalate care.
The archetype structure makes 10+ year longitudinal analysis feasible because the semantic contracts hold across time. A problem list entry from 2016 has the same structure as one from 2026. Your progression model does not need version-aware parsing logic.
Building the AI-Ready Clinical Data Platform
Knowing that openEHR produces AI-ready data is the starting point. Operationalizing it requires a platform architecture that connects the clinical data repository to your ML infrastructure.
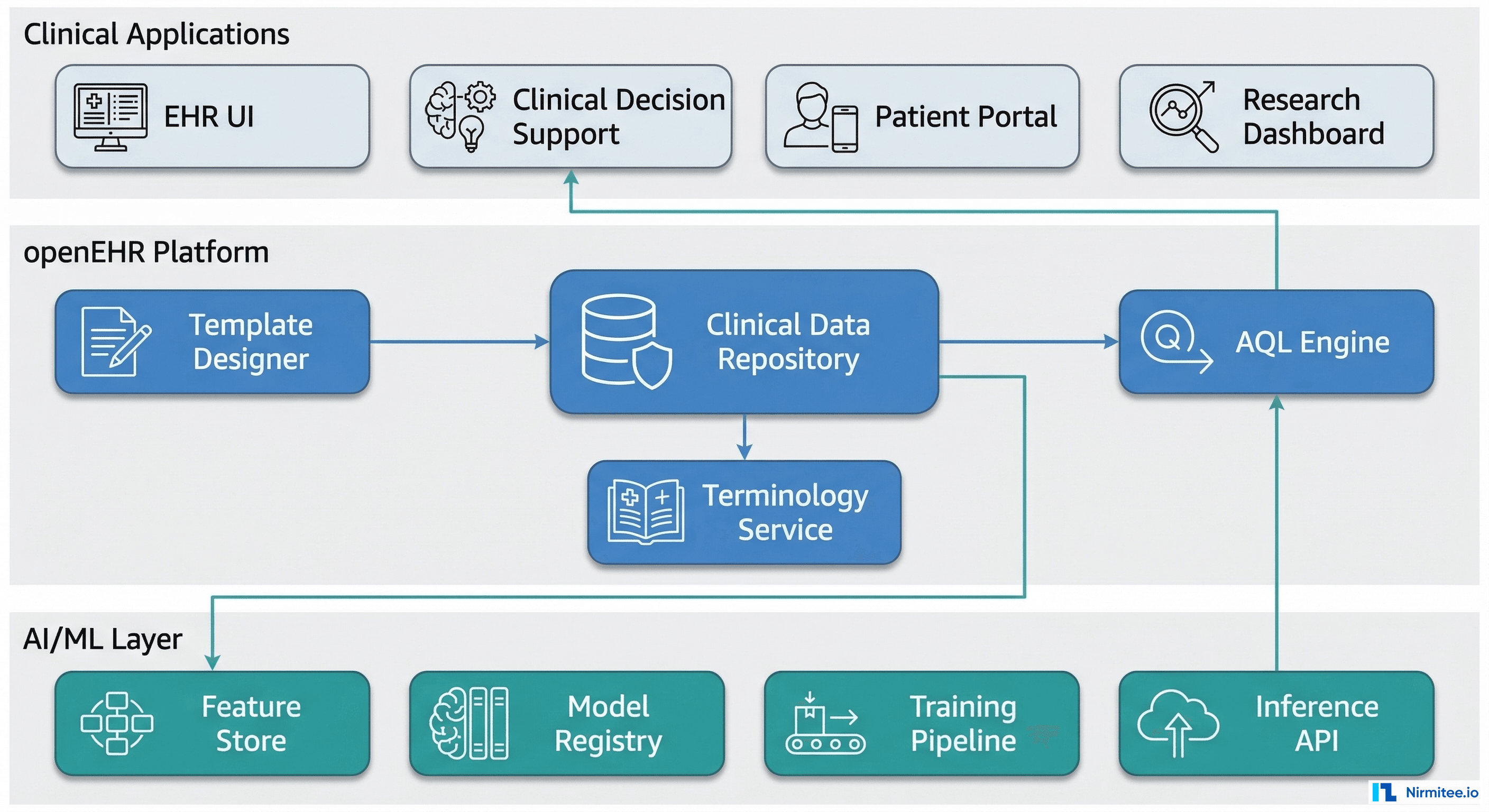
The Three-Layer Architecture
Layer 1 — Clinical Applications: EHR interfaces, clinical decision support, patient portals, and research dashboards. These are the consumers of both raw clinical data and AI-generated insights.
Layer 2 — openEHR Platform: The Clinical Data Repository (CDR) at the center, flanked by the Template Designer (for configuring data capture) and the AQL Engine (for data retrieval). A Terminology Service provides SNOMED CT, LOINC, and other vocabulary resolution.
Layer 3 — AI/ML Infrastructure: A Feature Store caches pre-computed features extracted via AQL. A Model Registry tracks trained models and their archetype dependencies. A Training Pipeline orchestrates model development. An Inference API serves predictions back to clinical applications.
The Feature Store Pattern
The Feature Store is the critical integration point. It bridges openEHR's clinical semantics with ML's need for fast, pre-computed numerical features:
class OpenEHRFeatureStore:
def __init__(self, cdr_client, cache):
self.cdr = cdr_client
self.cache = cache
self.extractors = {}
def get_features(self, patient_id, feature_set, freshness='1h'):
cache_key = f"{patient_id}:{feature_set}:{freshness}"
cached = self.cache.get(cache_key)
if cached:
return cached
# AQL queries are archetype-path based — portable across CDRs
extractor = self.extractors[feature_set]
features = extractor.extract(patient_id)
self.cache.set(
cache_key,
features,
ttl=freshness
)
return features
def refresh_cohort(self, patient_ids, feature_set):
# Batch AQL query for cohort-level feature extraction
return self.cdr.batch_query(
archetype_paths=self.extractors[feature_set].paths,
patient_ids=patient_ids
)This pattern gives you sub-millisecond feature serving for real-time inference while maintaining the semantic guarantees of the archetype layer.
Common Objections (And Why They Do Not Hold)
"We can normalize data in a data warehouse instead"
You can. And organizations do. But a data warehouse normalizes after the fact — it is a remediation layer on top of semantically inconsistent source data. openEHR normalizes at the point of capture. The data warehouse approach means maintaining transformation logic for every source system, every schema change, every terminology update. With openEHR, the archetype is the normalization.
"FHIR resources also have defined structures"
FHIR resources define the shape of data for exchange, not for clinical depth. A FHIR Observation resource for blood pressure does not mandate patient position, measurement protocol, or cuff size. It can carry them as extensions, but extensions are implementation-specific — which means you are back to per-site mapping. openEHR archetypes mandate the full clinical model, including context that matters for AI feature quality.
"Our data scientists can handle messy data"
They can. And they spend 80% of their time doing it. The question is not whether they can handle it, but whether the organization can afford the cost. At typical health system data science team rates, 20 weeks of data preparation across a team of 3-4 people represents $250,000-$400,000 per model. openEHR reduces that to $60,000-$100,000. Multiply by the number of models in your AI roadmap.
Getting Started: Three Practical Steps
Step 1 — Audit your AI use case backlog. Identify which planned ML models depend on clinical data that spans multiple source systems or requires complex semantic mapping. These are the use cases where openEHR will deliver the largest ROI.
Step 2 — Pilot with a single archetype domain. Start with a well-defined clinical domain — vital signs or lab results — where international archetypes are mature and your organization has clear AI use cases. Build one feature extractor, train one model, and measure the pipeline effort against your traditional approach.
Step 3 — Design the Feature Store bridge. Plan how your openEHR CDR will feed pre-computed features to your ML infrastructure. The AQL Engine → Feature Store → Model Serving pattern is well-proven and can be implemented incrementally alongside existing data warehouse pipelines.
The organizations that will lead clinical AI in the next five years are not the ones with the most data scientists. They are the ones whose data architecture makes those data scientists productive from day one. openEHR is that architecture.
Share
Related Posts
NABH Asset Management Compliance: A Complete Guide for Indian Hospitals
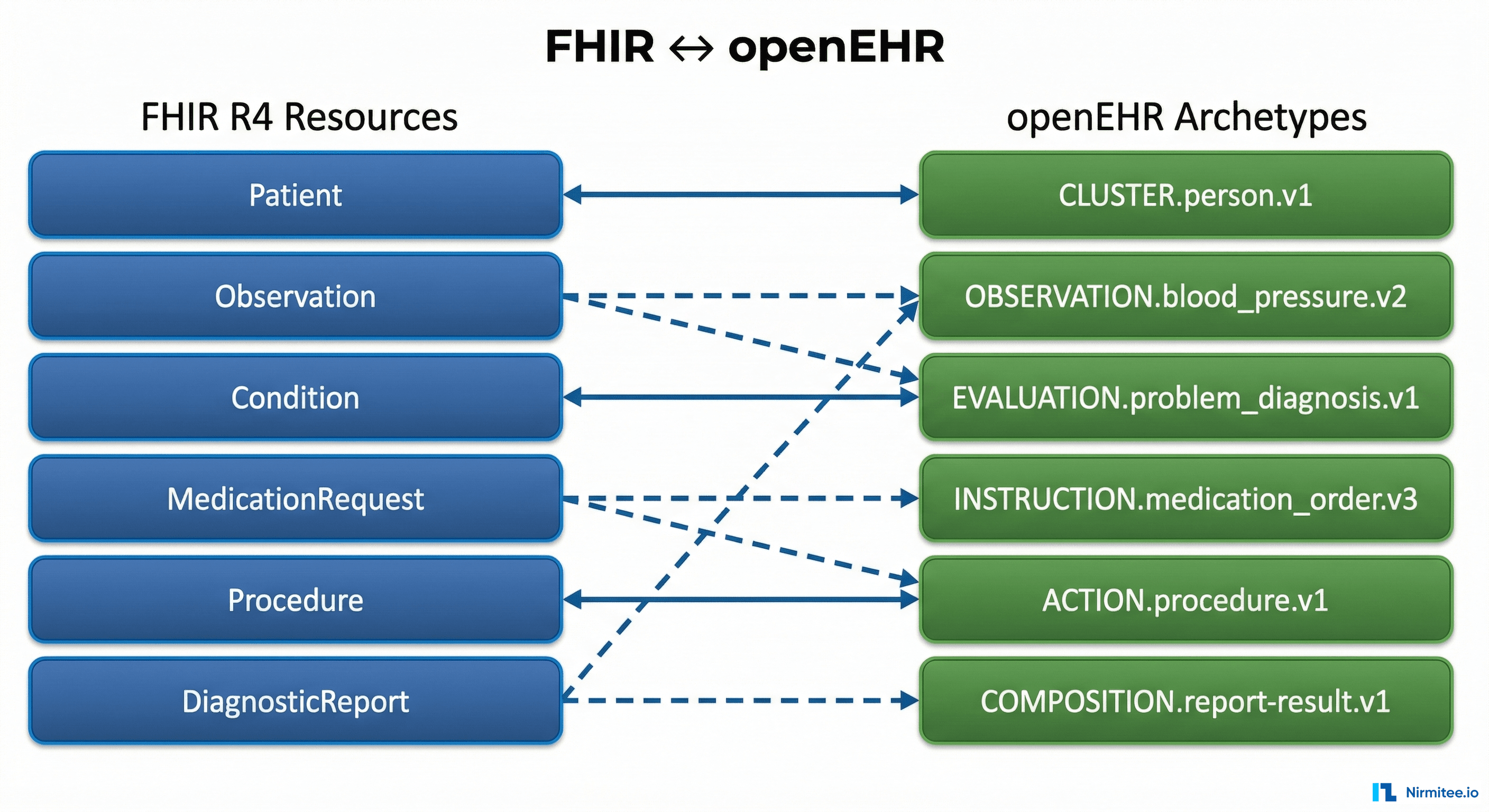
The Complete FHIR-to-openEHR Resource Mapping Matrix (2026 Reference)