Every developer who starts working with an openEHR Clinical Data Repository goes through the same two weeks of pain. The specification is comprehensive. The documentation is scattered. The examples assume you already know things you do not know yet. And the gap between "I understand the concepts" and "I can POST a composition that the CDR actually accepts" is wider than anyone warns you about.
This guide bridges that gap. It covers every REST API operation you will use in a real implementation — with actual curl commands, real JSON payloads, real error messages, and the explanations of why things fail that the official docs leave out. Bookmark this. You will come back to it.
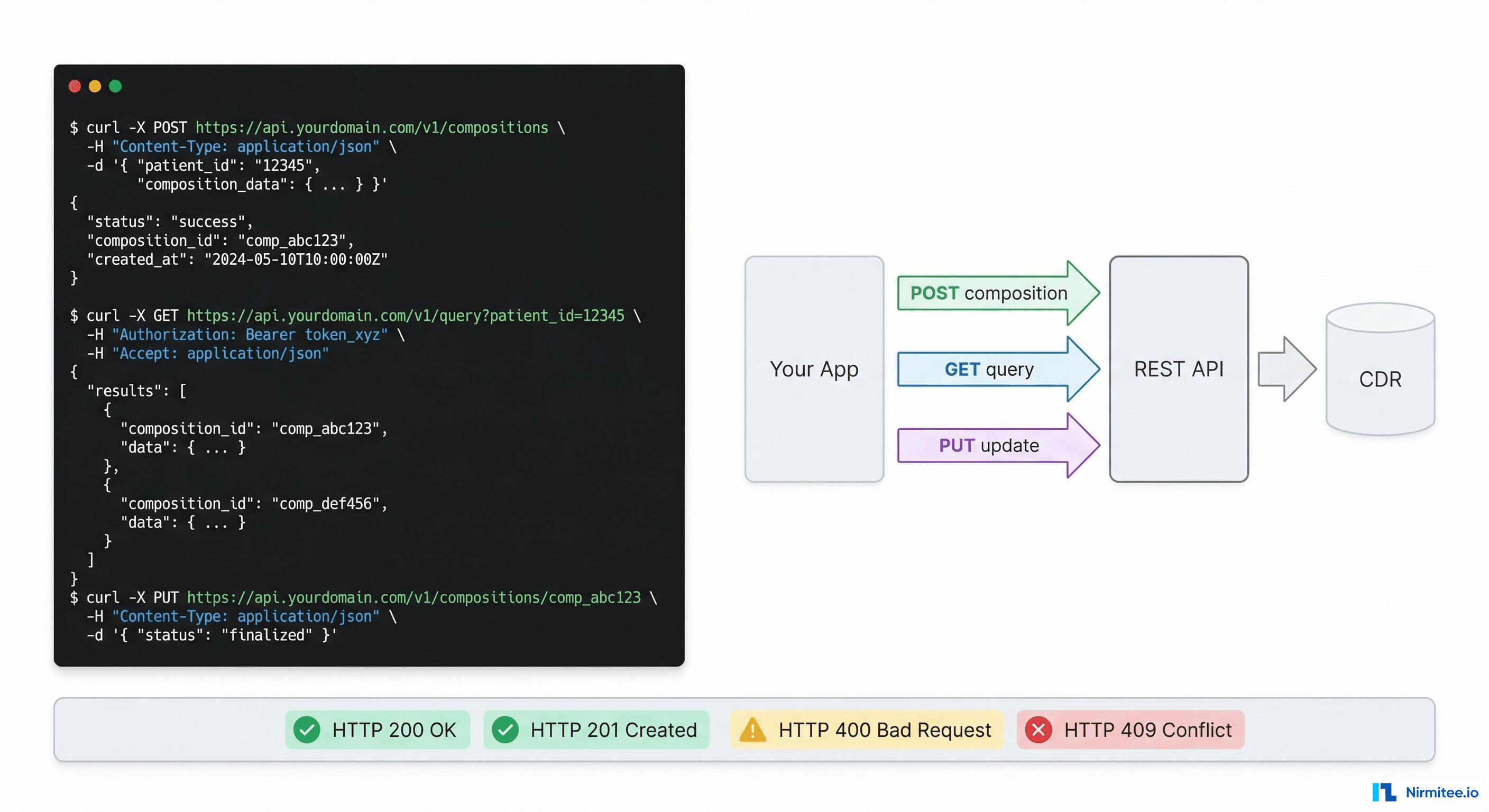
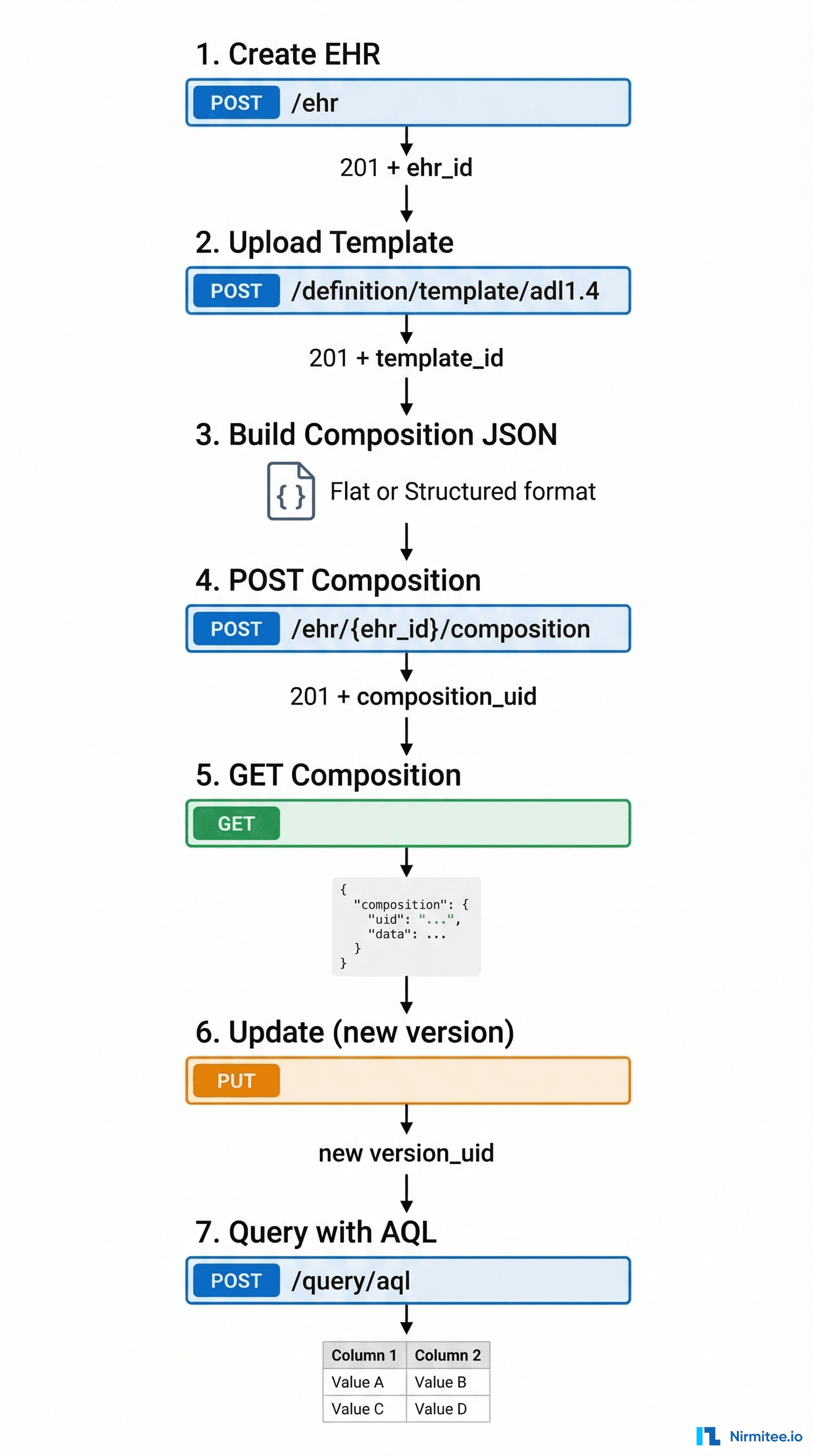
The Complete Lifecycle: Seven API Calls That Do Everything
An openEHR integration has exactly seven operations that matter. Everything else is a variation of these. Master these and you can build any clinical data application.
Step 1: Create an EHR
Before you can store any clinical data, you need an EHR (Electronic Health Record) container for the patient. Think of it as creating a patient folder — the folder itself has no clinical data, but every composition will go inside it.
curl -X POST 'http://localhost:8080/ehrbase/rest/openehr/v1/ehr' \
-H 'Content-Type: application/json' \
-H 'Accept: application/json' \
-H 'Prefer: return=representation' \
-d '{
"ehr_status": {
"subject": {
"external_ref": {
"id": {
"value": "patient-12345",
"scheme": "hospital-mpi"
},
"namespace": "hospital.com",
"type": "PERSON"
}
},
"is_modifiable": true,
"is_queryable": true
}
}'Response (201 Created):
{
"ehr_id": {
"value": "7d44b88c-4199-4bad-97dc-d78268e01398"
},
"ehr_status": {
...
},
"system_id": {
"value": "ehrbase.org"
},
"time_created": {
"value": "2026-03-12T10:00:00.000Z"
}
}What to save: The ehr_id.value — you will need this for every subsequent API call for this patient. Store it in your user/patient table alongside whatever patient identifier your system uses.
The gotcha: If you POST again with the same subject external_ref, most CDRs will return 409 Conflict because the EHR already exists. Use GET /ehr?subject_id=patient-12345&subject_namespace=hospital.com to look up existing EHRs first.
Alternative — create with known ID:
curl -X PUT 'http://localhost:8080/ehrbase/rest/openehr/v1/ehr/7d44b88c-4199-4bad-97dc-d78268e01398' \
-H 'Content-Type: application/json' \
-d '{"ehr_status": {...}}'Using PUT with a specific ehr_id lets you control the identifier — useful when migrating from another system where you want to preserve IDs.
Step 2: Upload a Template
Templates tell the CDR what kinds of clinical documents it should accept. You must upload at least one template before you can POST compositions. Think of it as registering a form definition before accepting form submissions.
curl -X POST 'http://localhost:8080/ehrbase/rest/openehr/v1/definition/template/adl1.4' \
-H 'Content-Type: application/xml' \
-H 'Accept: application/json' \
-d @vital-signs-template.optResponse (201 Created):
{
"template_id": "Vital Signs",
"concept": "Vital Signs",
"archetype_id": "openEHR-EHR-COMPOSITION.encounter.v1"
}Templates are OPT files (Operational Template, XML format). You create these using a template designer tool, not by hand. The CDR parses the template and uses it to validate every composition you submit.
The gotcha: Upload the template once. If you try to upload the same template_id again, some CDRs return 409 Conflict, others silently update. Check your CDR's behavior. Use GET /definition/template/adl1.4 to list all uploaded templates.
Pro tip — get the web template:
curl 'http://localhost:8080/ehrbase/rest/openehr/v1/definition/template/adl1.4/Vital%20Signs' \
-H 'Accept: application/openehr.wt+json'The web template response gives you the flat paths and tree structure you need to build composition JSON. This is the single most useful endpoint for front-end developers — it tells you exactly which fields exist, their types, their constraints, and their paths.
Step 3: Build a Composition
This is where most teams get stuck. The CDR accepts compositions in two formats: structured (canonical) and flat (simplified). Choose wisely — it affects your entire integration architecture.
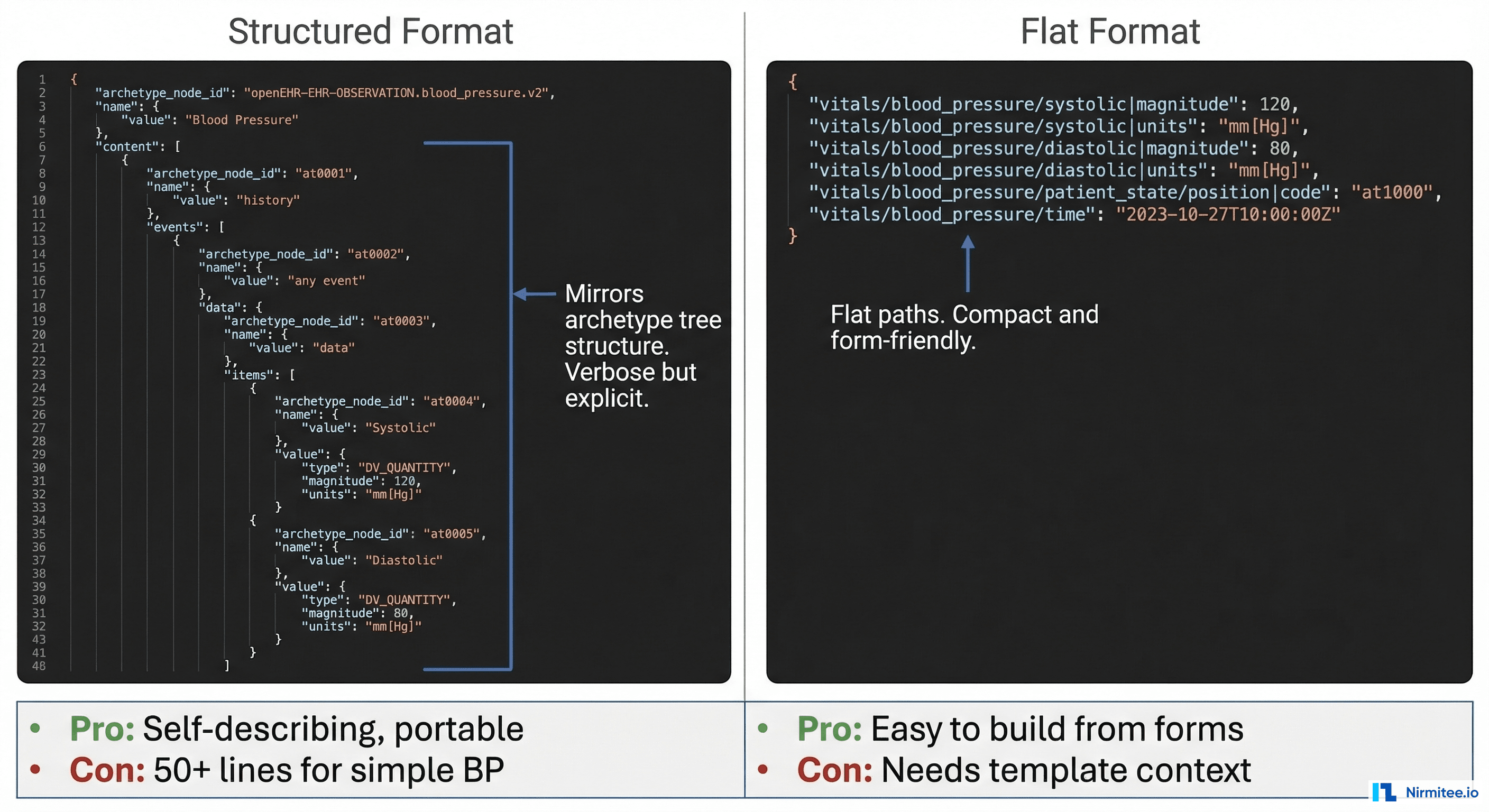
Structured Format (Canonical)
Mirrors the archetype tree structure exactly. Self-describing — you can interpret the JSON without knowing which template produced it. But it is verbose: a simple blood pressure reading becomes 50+ lines of JSON.
{
"_type": "COMPOSITION",
"archetype_node_id": "openEHR-EHR-COMPOSITION.encounter.v1",
"name": {
"value": "Vital Signs"
},
"language": {
"terminology_id": {
"value": "ISO_639-1"
},
"code_string": "en"
},
"territory": {
"terminology_id": {
"value": "ISO_3166-1"
},
"code_string": "US"
},
"category": {
"value": "event",
"defining_code": {
"terminology_id": {
"value": "openehr"
},
"code_string": "433"
}
},
"composer": {
"_type": "PARTY_IDENTIFIED",
"name": "Dr. Smith"
},
"content": [
{
"_type": "OBSERVATION",
"archetype_node_id": "openEHR-EHR-OBSERVATION.blood_pressure.v2",
"name": {
"value": "Blood pressure"
},
"language": {
"terminology_id": {
"value": "ISO_639-1"
},
"code_string": "en"
},
"encoding": {
"terminology_id": {
"value": "IANA_character-sets"
},
"code_string": "UTF-8"
},
"subject": {},
"data": {
"archetype_node_id": "at0001",
"name": {
"value": "History"
},
"origin": {
"value": "2026-03-12T10:30:00Z"
},
"events": [
{
"archetype_node_id": "at0006",
"name": {
"value": "Any event"
},
"time": {
"value": "2026-03-12T10:30:00Z"
},
"data": {
"archetype_node_id": "at0003",
"name": {
"value": "blood pressure"
},
"items": [
{
"archetype_node_id": "at0004",
"name": {
"value": "Systolic"
},
"value": {
"_type": "DV_QUANTITY",
"magnitude": 120.0,
"units": "mm[Hg]"
}
},
{
"archetype_node_id": "at0005",
"name": {
"value": "Diastolic"
},
"value": {
"_type": "DV_QUANTITY",
"magnitude": 80.0,
"units": "mm[Hg]"
}
}
]
}
}
]
}
}
]
}Flat Format (Simplified)
Uses dot-separated paths as keys. Compact, easy to generate from form data, and what most web/mobile apps actually use in practice.
{
"ctx/language": "en",
"ctx/territory": "US",
"ctx/composer_name": "Dr. Smith",
"vital_signs/blood_pressure/any_event:0/systolic|magnitude": 120.0,
"vital_signs/blood_pressure/any_event:0/systolic|unit": "mm[Hg]",
"vital_signs/blood_pressure/any_event:0/diastolic|magnitude": 80.0,
"vital_signs/blood_pressure/any_event:0/diastolic|unit": "mm[Hg]",
"vital_signs/blood_pressure/any_event:0/time": "2026-03-12T10:30:00Z"
}Practical recommendation: Use flat format for write operations (forms → CDR) and structured format for read operations (CDR → analytics). Flat is dramatically easier to build from UI form data. Structured is better for data processing pipelines that need self-describing documents.
Step 4: POST a Composition
This is the core write operation. You are storing a clinical document for a patient.
Structured format:
curl -X POST 'http://localhost:8080/ehrbase/rest/openehr/v1/ehr/7d44b88c-4199-4bad-97dc-d78268e01398/composition' \
-H 'Content-Type: application/json' \
-H 'Accept: application/json' \
-H 'Prefer: return=representation' \
-d @composition.jsonFlat format (note the different content type):
curl -X POST 'http://localhost:8080/ehrbase/rest/ecis/v1/composition?ehrId=7d44b88c-...&templateId=Vital%20Signs&format=FLAT' \
-H 'Content-Type: application/json' \
-d @flat-composition.jsonNote: The /ecis/v1/ endpoint shown above is EHRbase-specific, not part of the openEHR REST specification. Better Platform and Ocean have their own flat format APIs with different endpoint paths. The flat format concept is the same across CDRs; only the URL differs.
Response (201 Created):
{
"uid": {
"value": "8849182c-82ad-4088-a07f-48ead4180515::ehrbase.org::1"
},
"archetype_node_id": "openEHR-EHR-COMPOSITION.encounter.v1",
"name": {
"value": "Vital Signs"
},
...
}What to save: The uid.value — this is the version_uid. It has three parts separated by ::: the object ID, the creating system ID, and the version number. You need the full version_uid for updates.
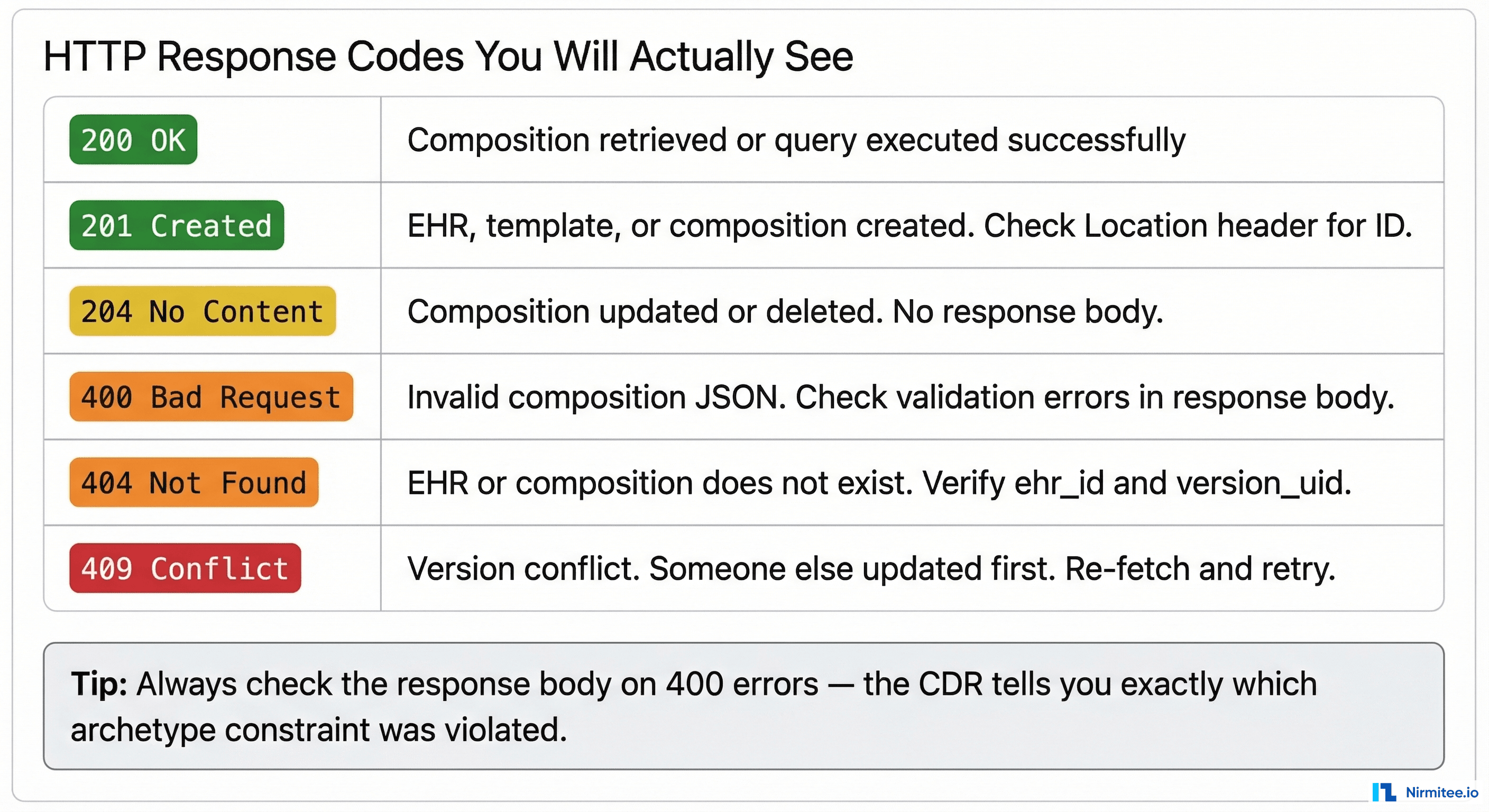
The gotcha that wastes the most time: If the CDR returns 400 Bad Request, the response body contains validation errors. Read them carefully. Common causes:
- Missing mandatory fields defined in the template
- Wrong data type (sending a string where DV_QUANTITY is expected)
- Wrong units (sending "mmHg" instead of "mm[Hg]" — the UCUM format matters)
- Archetype node ID mismatch between your JSON and the template
- Missing
_typefield on structured format objects

Step 5: GET a Composition
Retrieving a specific composition by its version_uid:
curl 'http://localhost:8080/ehrbase/rest/openehr/v1/ehr/7d44b88c-.../composition/8849182c-82ad-4088-a07f-48ead4180515::ehrbase.org::1' \
-H 'Accept: application/json'This returns the full structured composition. To get a specific version, use the full version_uid with the version number. To get the latest version, use just the object ID (without ::system::version):
# Latest version curl '.../composition/8849182c-82ad-4088-a07f-48ead4180515'
# Specific version curl '.../composition/8849182c-82ad-4088-a07f-48ead4180515::ehrbase.org::2'Pro tip: Add Accept: application/openehr.wt+json header to get flat format back — much easier to process in front-end applications.
Step 6: Update a Composition (Versioning)
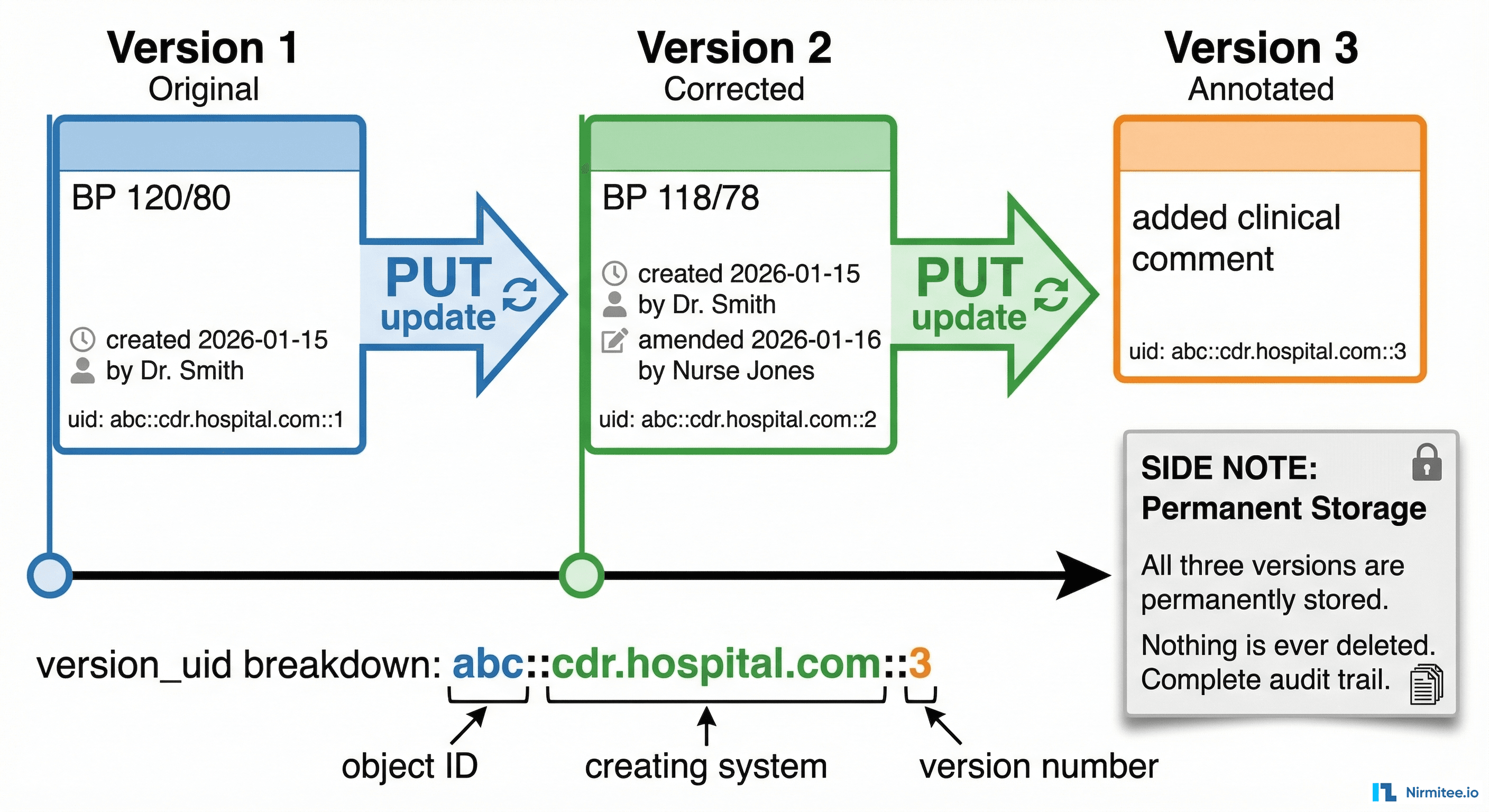
openEHR never overwrites data. When you "update" a composition, you create a new version. The previous version is permanently preserved. This is legally required in most healthcare jurisdictions and is one of openEHR's strongest features.
curl -X PUT 'http://localhost:8080/ehrbase/rest/openehr/v1/ehr/7d44b88c-.../composition/8849182c-82ad-4088-a07f-48ead4180515' \
-H 'Content-Type: application/json' \
-H 'Accept: application/json' \
-H 'If-Match: 8849182c-82ad-4088-a07f-48ead4180515::ehrbase.org::1' \
-H 'Prefer: return=representation' \
-d @updated-composition.jsonCritical: The If-Match header must contain the current version_uid. This is optimistic concurrency control — if someone else updated the composition between your GET and your PUT, you get 409 Conflict and must re-fetch, merge, and retry.
Response (200 OK):
{
"uid": {
"value": "8849182c-82ad-4088-a07f-48ead4180515::ehrbase.org::2"
}
}Notice the version number incremented from ::1 to ::2. The old version at ::1 is still retrievable. Nothing was deleted or overwritten.
The version_uid format explained:
8849182c-82ad-4088-a07f-48ead4180515— the permanent object ID (never changes across versions)ehrbase.org— the system that created this version2— the version number (increments with each update)
Deletion works the same way:
curl -X DELETE '.../composition/8849182c-...' \
-H 'If-Match: 8849182c-...::ehrbase.org::2'This does not physically remove the data. It creates a "deleted" version marker. The previous versions are still in the CDR and can be retrieved. In clinical data, nothing ever truly disappears — and your integration code should expect this.
Step 7: Query with AQL
AQL queries let you search across all compositions for all patients. This is how you build dashboards, reports, clinical decision support, and analytics.
curl -X POST 'http://localhost:8080/ehrbase/rest/openehr/v1/query/aql' \
-H 'Content-Type: application/json' \
-H 'Accept: application/json' \
-d '{
"q": "SELECT e/ehr_id/value as patient_id, o/data[at0001]/events[at0006]/data[at0003]/items[at0004]/value/magnitude as systolic, o/data[at0001]/events[at0006]/data[at0003]/items[at0005]/value/magnitude as diastolic, o/data[at0001]/events[at0006]/time/value as measured_at FROM EHR e CONTAINS OBSERVATION o[openEHR-EHR-OBSERVATION.blood_pressure.v2] WHERE o/data[at0001]/events[at0006]/data[at0003]/items[at0004]/value/magnitude > 140 ORDER BY o/data[at0001]/events[at0006]/time/value DESC LIMIT 100"
}'Response:
{
"rows": [
[
"7d44b88c-...",
165.0,
95.0,
"2026-03-12T10:30:00Z"
],
[
"a1b2c3d4-...",
158.0,
92.0,
"2026-03-11T14:15:00Z"
],
...
],
"columns": [
{
"name": "patient_id",
"path": "/ehr_id/value"
},
{
"name": "systolic",
"path": "/data[
at0001
]/events[
at0006
]/data[
at0003
]/items[
at0004
]/value/magnitude"
},
{
"name": "diastolic",
"path": "/data[
at0001
]/events[
at0006
]/data[
at0003
]/items[
at0005
]/value/magnitude"
},
{
"name": "measured_at",
"path": "/data[
at0001
]/events[
at0006
]/time/value"
}
]
}The response is tabular — rows and columns, like SQL. You can feed this directly into pandas, a charting library, or any analytics tool.
Parameterized queries (prevent AQL injection):
{
"q": "SELECT ... FROM EHR e CONTAINS OBSERVATION o[openEHR-EHR-OBSERVATION.blood_pressure.v2] WHERE e/ehr_id/value = $patient_id AND o/data[at0001]/events[at0006]/time/value > $since",
"query_parameters": {
"patient_id": "7d44b88c-...",
"since": "2025-06-01"
}
}Always use parameterized queries in production. Never concatenate user input into AQL strings — the same injection risks as SQL apply here.
The Seven Mistakes Every Team Makes in Week One
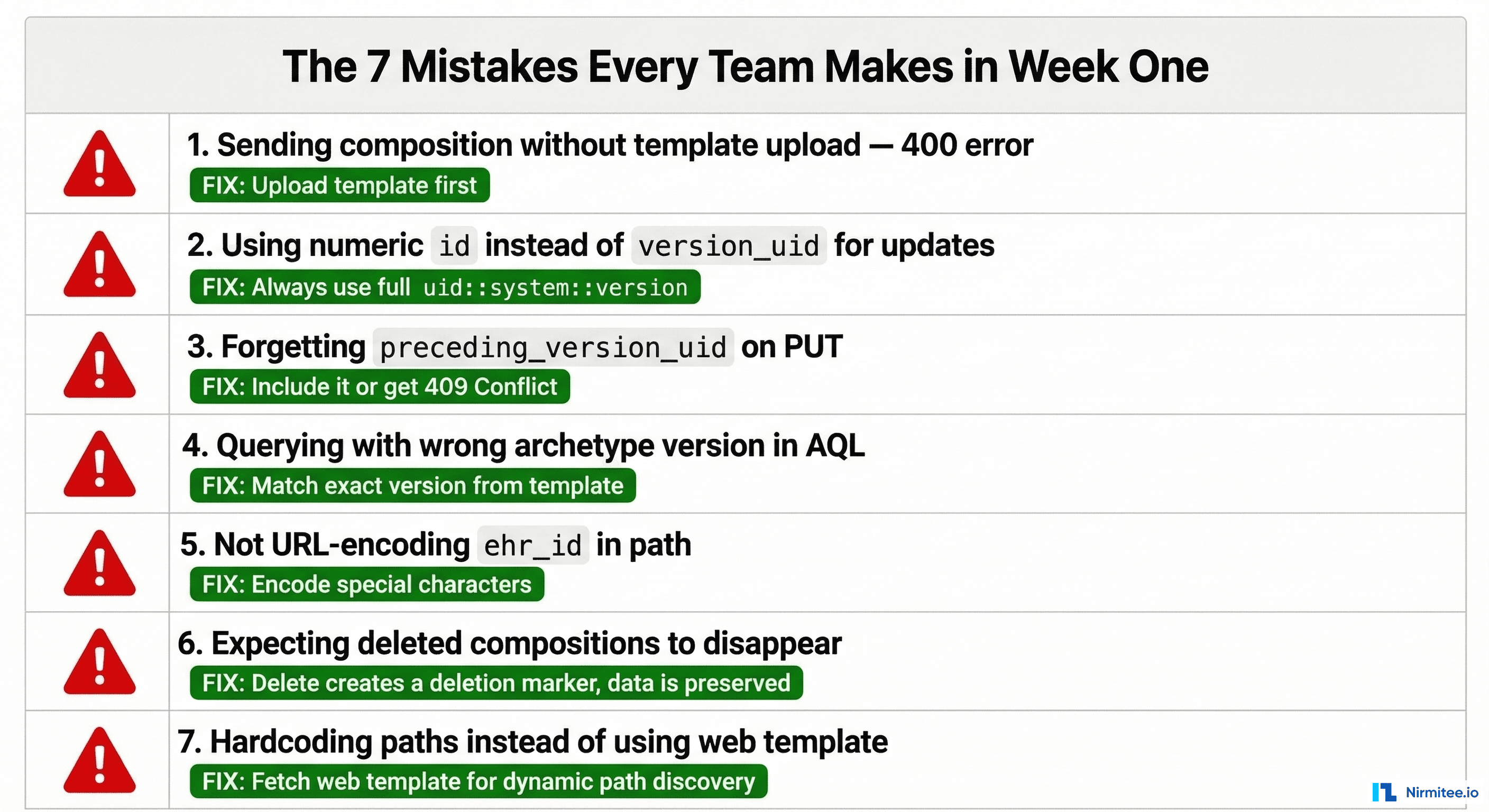

Mistake 1: Posting compositions without uploading the template first
The CDR validates every composition against its template. No template → 400 Bad Request with an unhelpful error message. Always verify your template is uploaded: GET /definition/template/adl1.4/{template_id}.
Mistake 2: Using the numeric database ID instead of version_uid
Some CDRs expose internal numeric IDs in list responses. Never use these for API calls. The openEHR standard uses the version_uid format: object_id::system_id::version. If your code references a numeric ID, something is wrong.
Mistake 3: Forgetting the If-Match header on PUT
Updates require optimistic locking via the If-Match header containing the current version_uid. Without it, some CDRs return 400, others 409. Pattern: always GET the latest version, extract the version_uid, include it in If-Match on your PUT.
Mistake 4: Wrong archetype version in AQL queries
If your template uses blood_pressure.v2 but your AQL query references blood_pressure.v1, you get zero results with no error. The CDR does not warn you — it just finds no matching data. Always check the archetype version in your template matches your query.
Mistake 5: Wrong unit format
openEHR uses UCUM (Unified Code for Units of Measure) notation. Common traps: mm[Hg] not mmHg. kg/m2 not kg/m^2. Cel not °C. Get the exact unit string from the web template response — do not guess.
Mistake 6: Expecting DELETE to remove data
DELETE creates a deletion marker version. The data is preserved and retrievable. If your application logic depends on data being physically gone after delete, you need to redesign. openEHR is an append-only system by design.
Mistake 7: Hardcoding archetype paths instead of discovering them
Fetch the web template (GET /definition/template/adl1.4/{id} with Accept: application/openehr.wt+json) and use the paths it returns. This makes your code resilient to template changes and lets you build dynamic forms from template metadata.
Production Integration Pattern
In production, you do not call the CDR directly from your front-end. Here is the architecture pattern that works.
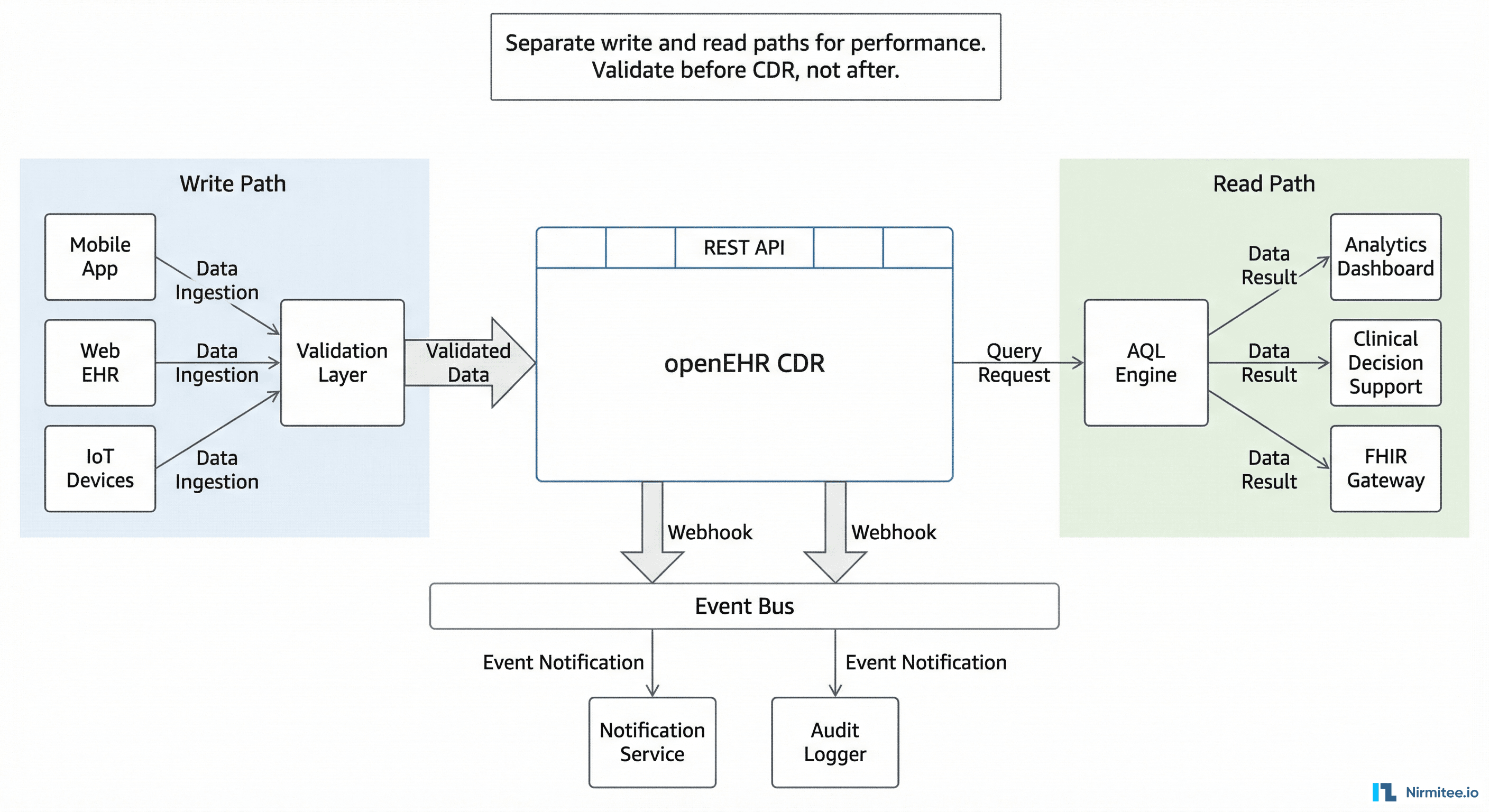
Write Path
Front-end form → your backend API → validation layer → CDR. The validation layer converts form data to flat-format composition JSON, validates business rules that the CDR does not know about (e.g., "nurses cannot submit medication orders"), and handles authentication. Never expose the CDR REST API directly to client applications.
Read Path
CDR → AQL Engine → your backend API → consumer applications. For dashboards and analytics, pre-build common AQL queries as named stored queries: POST /query/{query_name}. This avoids AQL construction in front-end code and lets you optimize queries centrally.
Event Path
Many CDRs support webhooks or event triggers when compositions are created or updated. Use these for real-time notifications (new lab result → alert the ordering physician), audit logging, and downstream synchronization (new diagnosis → update the problem list in your FHIR server).
Key Architectural Decisions
- Validate before the CDR, not after. CDR validation errors are correct but not user-friendly. Validate in your backend and return human-readable errors to the UI.
- Cache web templates. Fetch once at startup, refresh on template change. Do not call
GET /definition/templateon every form render. - Use flat format for writes, structured for reads. Your front-end team will thank you.
- Index your AQL queries. The CDR's AQL engine performs best with archetype-specific queries. Avoid broad
CONTAINS COMPOSITIONqueries without archetype filters in production. - Handle version conflicts gracefully. In clinical workflows, two nurses can record vital signs for the same patient simultaneously. Your UI should detect
409 Conflictand merge or prompt, not crash.
Quick Reference: Every Endpoint You Need
| Operation | Method | Endpoint | Key Headers |
|---|---|---|---|
| Create EHR | POST | /ehr | Prefer: return=representation |
| Get EHR by subject | GET | /ehr?subject_id=X&subject_namespace=Y | — |
| Upload template | POST | /definition/template/adl1.4 | Content-Type: application/xml |
| Get web template | GET | /definition/template/adl1.4/{id} | Accept: application/openehr.wt+json |
| Create composition | POST | /ehr/{ehr_id}/composition | Prefer: return=representation |
| Get composition | GET | /ehr/{ehr_id}/composition/{version_uid} | — |
| Update composition | PUT | /ehr/{ehr_id}/composition/{object_id} | If-Match: {current_version_uid} |
| Delete composition | DELETE | /ehr/{ehr_id}/composition/{object_id} | If-Match: {current_version_uid} |
| Execute AQL query | POST | /query/aql | Content-Type: application/json |
| List stored queries | GET | /query | — |
| Store named query | PUT | /query/{query_name} | Content-Type: application/json |
All endpoints are relative to the base URL of your CDR's openEHR REST API (e.g., http://localhost:8080/ehrbase/rest/openehr/v1). The base path varies by CDR vendor — check your CDR's documentation for the correct prefix.
From Here: What to Build First
If you are starting a new openEHR integration, build in this order:
- Template upload and web template fetch. Automate this in your CI/CD pipeline. Templates should be version-controlled and deployed like database migrations.
- Single composition write. Get one vital signs composition through the CDR with flat format. This proves your end-to-end path works.
- Composition read-back. Verify what you wrote matches what you read. This catches serialization bugs early.
- AQL query for the data you wrote. Confirm the data is queryable and the archetype paths match.
- Version update cycle. Create → update → verify both versions exist. This proves your version_uid handling works.
- Error handling. Intentionally submit invalid compositions and verify your code handles 400 and 409 responses gracefully.
Do not try to build the full clinical application first. Get the API integration solid with one template, one composition type, and one AQL query. Then expand. The API surface is the same for every clinical concept — once blood pressure works, adding lab results, medications, and diagnoses is the same pattern with different templates and paths.
The teams that ship fastest are the ones that master the seven operations above before they write a single line of UI code. The API is your foundation. Build it right and everything above it moves faster.
Share
Related Posts
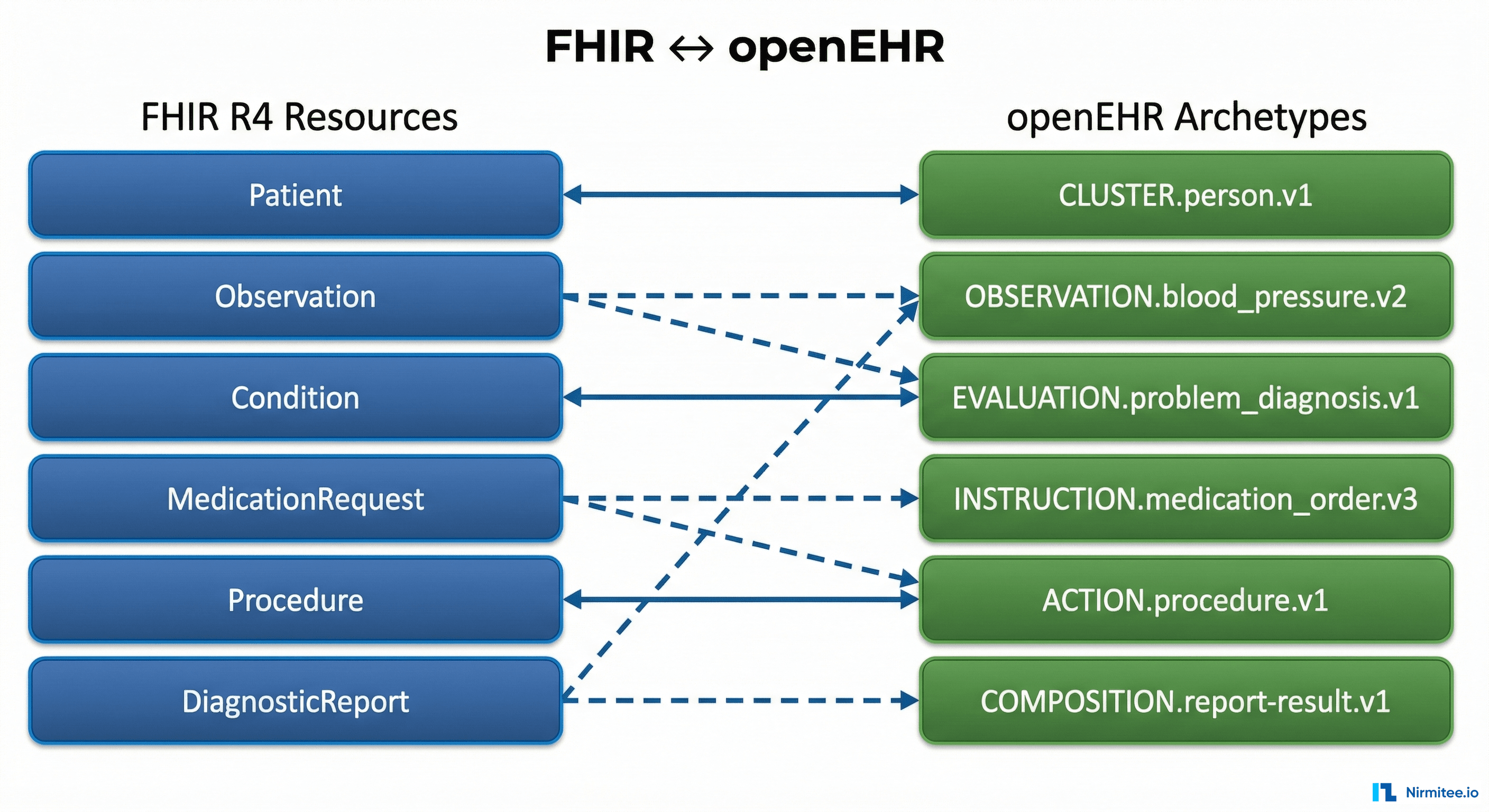
The Complete FHIR-to-openEHR Resource Mapping Matrix (2026 Reference)
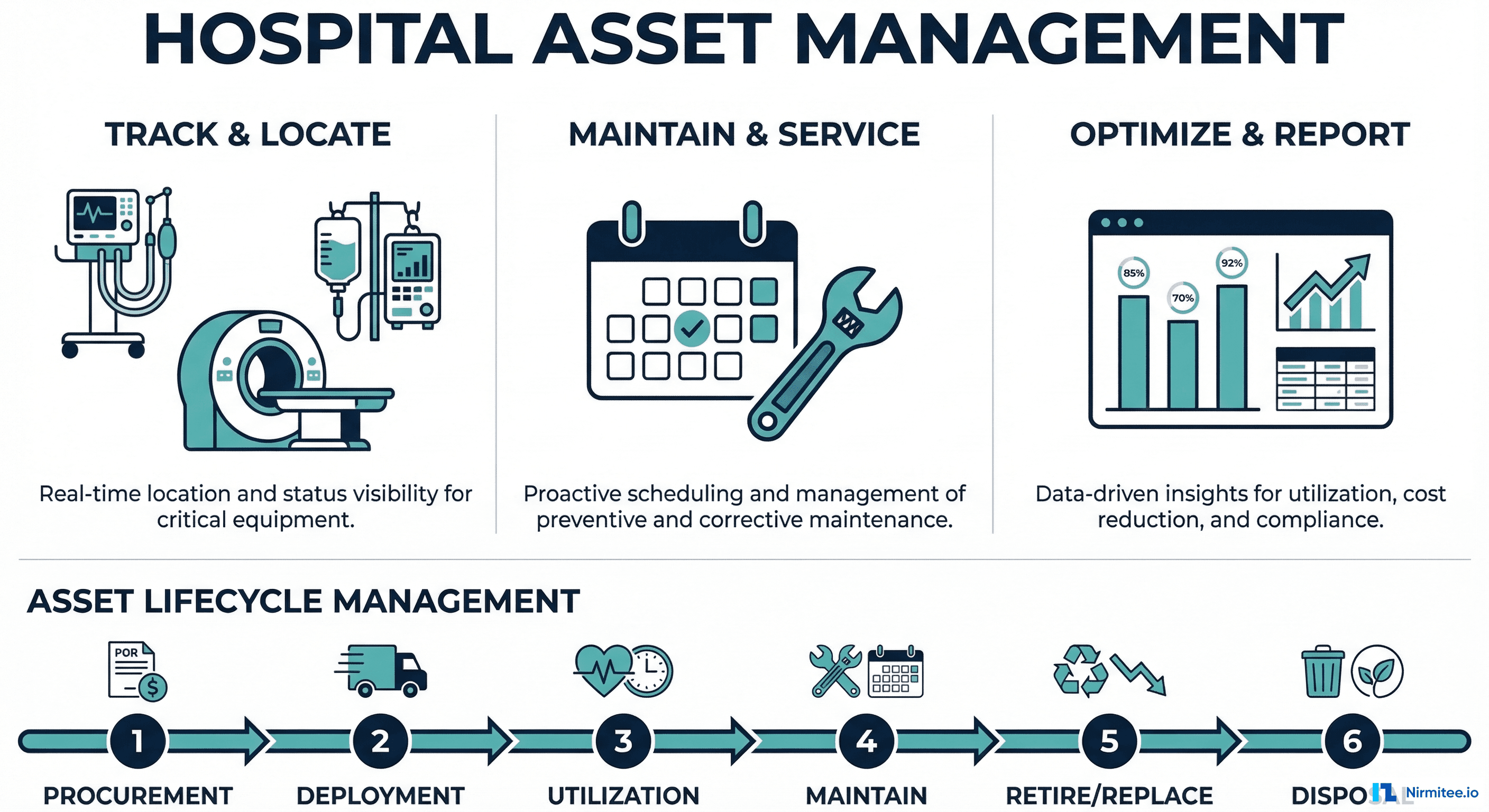
The Complete Guide to Hospital Asset Management in 2026