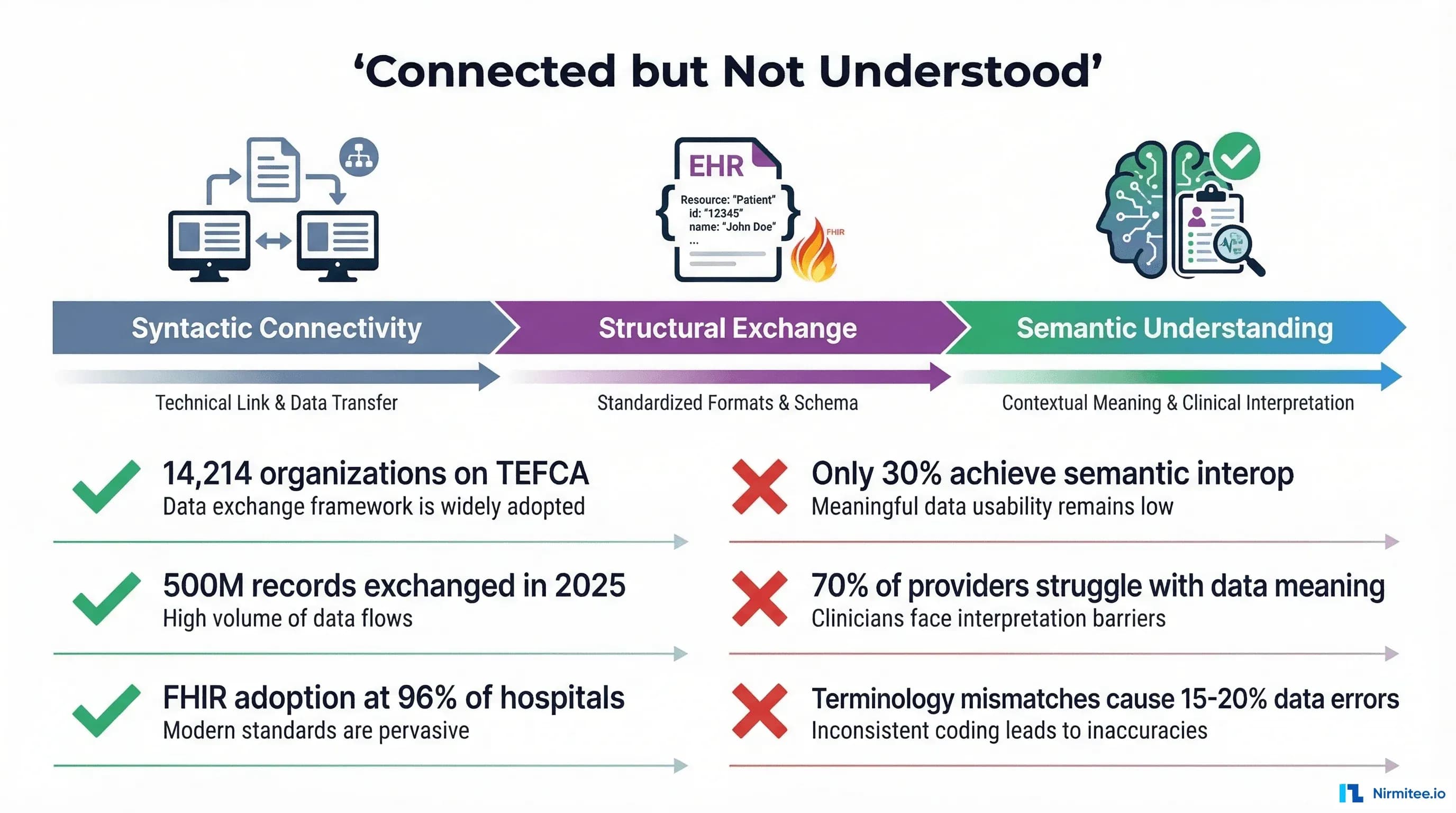
Healthcare spent the last decade solving the wrong problem. We connected the systems. We built the pipes. TEFCA now links 14,214 organizations across the United States, and nearly 500 million health records flowed through that network in 2025 alone. FHIR adoption has reached 96% of certified EHR systems. From a connectivity standpoint, the healthcare interoperability problem looks solved.
It is not.
The systems are connected, but they do not understand each other. A patient's diabetes diagnosis coded as E11.9 in one EHR, 44054006 in another, and stored as free text "Type II DM" in a third are, from the perspective of most receiving systems, three different conditions. A hemoglobin A1c result of 7.2% from Lab A and 55 mmol/mol from Lab B are the same measurement in different units, but without semantic alignment, they cannot be compared, trended, or used for clinical decision support. An allergy to "penicillin" entered as free text in one hospital and coded as an amoxicillin sensitivity in another can — and does — lead to missed cross-reactivity alerts.
This is the semantic interoperability problem: the gap between exchanging data and understanding it. It is the deepest unsolved problem in healthcare IT, and it is the reason that FHIR, TEFCA, and HL7 alone cannot deliver on the promise of interoperability. This article examines what semantic interoperability actually means, why current standards fall short, where real-world failures occur, and what approaches — from terminology services to AI — are beginning to close the gap.
The Connectivity vs. Understanding Gap
The numbers look impressive on paper. TEFCA, the Trusted Exchange Framework and Common Agreement, has grown from approximately 10 million records exchanged in January 2025 to nearly 500 million by early 2026. The Sequoia Project, serving as the Recognized Coordinating Entity (RCE), now oversees a network of Qualified Health Information Networks (QHINs) that together represent over 79,000 unique organizational connections. Health Gorilla, one of the inaugural QHINs, saw its query volume explode from 200,000 to 66 million per month.
But connectivity and understanding are different things. A systematic review published in the Journal of Medical Internet Research found that achieving semantic interoperability of electronic health records remains an open research challenge, with most implementations achieving only syntactic or structural interoperability. Nearly 70% of healthcare providers still struggle with seamless data exchange across platforms — not because the data cannot be sent, but because the receiving system cannot reliably interpret what was sent.
The ONC's HTI-5 Proposed Rule (December 2025) acknowledged this gap explicitly, proposing to shift away from legacy C-CDA standards toward FHIR-first interoperability and explicitly advancing a foundation for AI-enabled interoperability solutions. The regulatory infrastructure recognizes that exchanging documents is not the same as exchanging meaning.
Consider the difference in practical terms. A patient moves from Hospital A (Epic) to Hospital B (Oracle Health). Hospital A sends a C-CDA document via TEFCA. Hospital B receives it — foundational interoperability achieved. The receiving system parses the XML structure — structural interoperability achieved. But the problem list uses SNOMED CT codes that Hospital B's system maps differently. The medication list includes NDC codes that do not match Hospital B's formulary. The lab results use LOINC codes, but with different reference ranges and different units. The data is there, but it cannot be used for clinical decision-making without manual review by a clinician — which defeats the purpose of interoperability entirely.
What Semantic Interoperability Actually Means
The Healthcare Information and Management Systems Society (HIMSS) defines four levels of interoperability, each building on the last. Understanding these levels is essential because most healthcare IT investment — and most regulatory attention — focuses on the first two levels while the real clinical value lives in the third and fourth.
Level 1: Foundational Interoperability
The ability to send data from one system to another. This is the transport layer: TCP/IP, HTTPS, VPN tunnels, SFTP transfers. The sending system transmits bits, and the receiving system receives them. There is no requirement that the receiving system can interpret the data. This level is essentially solved — if two systems are on a network, they can exchange data. TEFCA operates at this level for network connectivity.
Level 2: Structural Interoperability
The ability to parse and process the received data at the field level. The receiving system knows that a particular element is a "diagnosis," another is a "medication," and another is a "lab result." Standards like FHIR, HL7 v2, C-CDA, and X12 EDI operate primarily at this level. FHIR resources define the structure: a Condition resource has a code field, a clinicalStatus field, and an onsetDateTime field. The receiving system knows where to find each piece of information.
Level 3: Semantic Interoperability
The ability to understand the meaning of the exchanged data in the same way. This is where healthcare breaks down. Semantic interoperability requires that when System A sends "Type 2 diabetes mellitus" coded as SNOMED CT 44054006, System B not only receives and parses the code, but understands it as the same clinical concept, maps it to its own internal representation, and can use it for clinical decision support, population health queries, quality measurement, and research. This requires shared terminologies, consistent terminology binding, and bidirectional concept mapping — none of which are guaranteed by FHIR or any transport standard.
Level 4: Organizational Interoperability
The ability to use shared data within governance, policy, and workflow frameworks. Even if two systems semantically understand each other's data, organizational interoperability asks: Is this organization authorized to receive this data? Does the patient consent to this exchange? What are the data use agreements? What happens when there is a conflict? TEFCA's Common Agreement addresses some of these questions, but organizational interoperability remains the least mature level in practice.
The critical insight is that most healthcare IT investment — the billions spent on EHR implementations, HIE networks, and FHIR API development — has focused on Levels 1 and 2. Level 3 is where clinical value is created, and it remains largely unresolved at scale.
The Terminology Tower of Babel
Healthcare has no shortage of coding systems. It has too many. Each was designed for a different purpose, by a different organization, with a different level of granularity, and they do not align cleanly. Understanding why is essential to understanding the semantic interoperability problem.
SNOMED CT: The Clinical Reference Terminology
SNOMED CT (Systematized Nomenclature of Medicine — Clinical Terms) is the most comprehensive clinical terminology in the world, with over 350,000 concepts organized in a polyhierarchical structure. Each concept is defined by a unique set of attribute-value pairs that distinguish it from every other concept. It is designed for capturing clinical meaning — the reason a clinician documents a finding, procedure, or diagnosis. SNOMED CT is the terminology you should use for problem lists, clinical observations, and clinical documentation.
But SNOMED CT is not used for billing. It is not used for lab orders. It is not used for drug identification. And critically, mapping SNOMED CT to other systems is not a one-to-one exercise.
ICD-10-CM: The Classification System
ICD-10-CM (International Classification of Diseases, 10th Revision, Clinical Modification) contains approximately 72,000 codes designed primarily for diagnosis classification, epidemiology, and billing. ICD-10 is a classification system, not a reference terminology — it groups similar conditions into categories for statistical and reimbursement purposes. The distinction matters enormously: SNOMED CT describes clinical meaning with fine granularity, while ICD-10-CM classifies conditions into billing-relevant buckets.
The NLM maintains an official SNOMED CT to ICD-10-CM map, but the mapping is inherently lossy. A single SNOMED CT concept may map to multiple ICD-10-CM codes depending on patient context (age, gender, comorbidities). The SNOMED CT concept "Sleep apnea" can map to either "Primary sleep apnea of newborn" or "Sleep apnea, unspecified" depending on the patient's age. Research shows that coding variability between professionals averages a Jaccard coefficient of only 0.53 — meaning on average, only about half of the codes from two independent coders agree.
LOINC: Laboratory and Clinical Observations
LOINC (Logical Observation Identifiers Names and Codes) contains over 98,000 codes for laboratory tests, clinical observations, and survey instruments. LOINC identifies what was measured (Glucose, Hemoglobin A1c, Blood Pressure), but not the result itself or the reference range. Two labs can both use LOINC code 2345-7 for serum glucose, but report results in mg/dL versus mmol/L with different reference ranges, making comparison impossible without unit normalization.
RxNorm: Medication Normalization
RxNorm provides normalized names and codes for clinical drugs in the United States. It bridges the gap between drug databases, EHR formularies, and pharmacy systems. RxNorm operates at multiple levels of specificity: ingredient (metformin), clinical drug (metformin 500mg oral tablet), and branded drug (Glucophage 500mg tablet). Cross-referencing RxNorm with the National Drug Code (NDC) directory — which identifies specific manufacturer packaged products — adds another layer of complexity, as NDC codes are version-specific and manufacturer-dependent.
CPT and HCPCS: Procedures and Billing
Current Procedural Terminology (CPT) codes, maintained by the AMA, and Healthcare Common Procedure Coding System (HCPCS) codes identify medical procedures and services for billing. These codes overlap with SNOMED CT procedure concepts but use entirely different hierarchies and groupings. A surgical procedure documented clinically in SNOMED CT must be re-coded in CPT/HCPCS for billing — a translation that is frequently inconsistent across organizations.
Why They Clash
The fundamental problem is that these systems were designed for different purposes and different audiences. SNOMED CT optimizes for clinical precision. ICD-10 optimizes for classification and billing. LOINC optimizes for lab test identification. RxNorm optimizes for drug normalization. CPT optimizes for procedure billing. When the same clinical reality — a patient with Type 2 diabetes taking metformin whose hemoglobin A1c was measured — must be represented across all six systems, semantic alignment requires mapping between systems that were never designed to map to each other.
Real Examples of Semantic Failure
Theory is abstract. These are concrete examples of how semantic interoperability failures affect patient care, clinical research, and health system operations. Every one of these scenarios occurs routinely in US healthcare today.
"Diabetes" in Three Systems Means Three Different Things
Hospital A uses Epic and records a diagnosis of "Type 2 diabetes mellitus with diabetic chronic kidney disease" using ICD-10-CM code E11.22. Hospital B uses Oracle Health and captures the same patient's condition as SNOMED CT code 44054006 ("Diabetes mellitus type 2") with a separate SNOMED code for the kidney complication. Hospital C, a community health center still running an older system, stores "DM Type II" as free text with a legacy ICD-9 code of 250.00.
When these records are aggregated for a population health query — "How many patients in our network have Type 2 diabetes with renal complications?" — the answer depends entirely on how the query system handles the semantic mismatch. Without a robust terminology mapping layer, Hospital A's patients are captured (ICD-10 code explicitly includes renal complication), Hospital B's patients may or may not be captured (depending on whether the separate SNOMED code for kidney disease is linked), and Hospital C's patients are invisible (free text plus legacy code contains no complication specificity).
Lab Results with Incompatible Reference Ranges
Lab A reports a serum creatinine of 1.2 mg/dL with a reference range of 0.7-1.3 mg/dL (normal). Lab B reports the same patient's creatinine as 106 μmol/L with a reference range of 62-115 μmol/L (also normal). Both use LOINC code 2160-0 for serum creatinine, but one reports in conventional US units and the other in SI units. A clinical decision support system that does not normalize units will either generate false alerts (flagging the SI value as abnormally high when compared against the conventional reference range) or miss genuine abnormalities.
This problem extends beyond units. Different labs use different methodologies (enzymatic vs. Jaffe for creatinine), different instruments with different analytical sensitivities, and different reference populations to establish normal ranges. Two "normal" results from two labs may represent meaningfully different clinical states. LOINC standardizes what was measured, not how it was measured or what the result means in context.
Medication Lists That Cannot Be Reconciled
Medication reconciliation — the process of creating an accurate list of all medications a patient is taking — is one of the most error-prone processes in healthcare. A patient fills prescriptions at CVS (which uses NDC codes), receives inpatient medications at a hospital (which uses RxNorm mapped to its formulary), and has a medication list in their primary care EHR (which may use a proprietary drug database). "Metformin 500mg twice daily" at the pharmacy, "Glucophage 500mg BID" in the hospital, and "metformin hydrochloride 500mg tablet" in the EHR are all the same medication — but without RxNorm normalization to the clinical drug level, automated reconciliation fails.
The consequences are clinical. Duplicate medications, missed drug interactions, and therapeutic substitution errors all stem from the inability to semantically align medication representations across systems.
Allergy Lists That Conflict Between EHRs
An AHRQ Patient Safety Network case study documented a patient with a known gelatin allergy whose EHR allergy entry did not trigger an alert when a gelatin-containing medication was prescribed — because the allergy was recorded as free text rather than as a coded, cross-referenced concept. A 2025 study in JAMIA on allergy reconciliation found that allergy information is often spread across the electronic health record, and allergy lists are frequently inaccurate or incomplete, with reactions entered as free text due to insufficiently comprehensive terminologies for encoding diverse reactions.
The semantic challenge is particularly acute with drug allergies, where the difference between "penicillin allergy" (a drug class allergy), "amoxicillin sensitivity" (a specific drug), and "hives from amoxicillin" (a drug + reaction) has direct clinical implications for cross-reactivity checking. Without semantic normalization, these entries are just strings — and strings cannot protect patients.
Why FHIR Does Not Solve Semantic Interoperability
This is a point that needs to be stated clearly because the industry narrative often conflates FHIR adoption with interoperability achievement: FHIR is a structural standard, not a semantic one.
FHIR (Fast Healthcare Interoperability Resources) defines resources — Condition, Observation, MedicationRequest, AllergyIntolerance — and specifies what fields each resource contains. A FHIR Condition resource has a code element where you put the diagnosis code, a clinicalStatus element for active/resolved/inactive, and a verificationStatus for confirmed/provisional/refuted. This is structural interoperability: the receiving system knows where to find each piece of information.
But FHIR does not mandate which terminology you use for the code element. US Core profiles bind certain elements to specific value sets — US Core Condition requires codes from SNOMED CT or ICD-10-CM — but the binding strength is often "extensible" or "preferred," not "required." This means systems can still use local codes, proprietary terminologies, or codes from value sets that the receiving system does not support.
A systematic mapping review published in JMIR (2024) examining FHIR and semantic issues found that even though FHIR allows the use of clinical terminologies like SNOMED CT and LOINC, users still enter information as text or use local terminologies. Selecting an appropriate concept from SNOMED CT's 350,000+ concepts is not a trivial task for a busy clinician, and many EHR interfaces make it easier to type free text than to search a terminology browser.
As one analysis put it directly: "FHIR alone is not enough. Terminology servers and normalization enable interoperability." FHIR provides the envelope. Semantic interoperability requires what goes inside the envelope to be mutually understood — and that requires terminology services, concept mapping, and normalization pipelines that sit on top of FHIR, not within it.
Approaches That Actually Work
Despite the scale of the problem, there are proven approaches to achieving semantic interoperability in healthcare. None of them are simple, none of them are cheap, and none of them work in isolation. But organizations that invest in these capabilities gain a meaningful advantage in data quality, clinical decision support, and research readiness.
Terminology Services: FHIR $translate, $lookup, and ConceptMap
FHIR's terminology services specification defines a set of operations that, when backed by a capable terminology server, provide the infrastructure for semantic translation. The key operations are:
- $translate: Converts a code from one system to another using a ConceptMap resource. For example, translating SNOMED CT 44054006 to ICD-10-CM E11.9. The operation returns the mapped code plus a relationship qualifier (equivalent, broader, narrower, inexact) that tells the consuming system how much semantic precision was preserved in the translation.
- $lookup: Returns metadata about a code — display name, properties, designations, parent/child relationships — without requiring the consuming system to have a local copy of the entire code system.
- $validate-code: Checks whether a given code is a member of a specified ValueSet, enabling systems to verify that received data conforms to expected terminology bindings.
- $expand: Returns all codes in a ValueSet, useful for building pick lists, validating incoming data, and understanding the full scope of a terminology binding.
Production-grade terminology servers like Ontoserver (CSIRO), Snowstorm (SNOMED International), and TerminologyHub provide these operations at scale. Google Cloud Healthcare API and AWS HealthLake also offer FHIR-native terminology services. The critical architectural decision is whether to deploy a terminology server as a shared service within your health system or consume it as a managed API — both approaches work, but the shared service model provides better control over mapping quality and custom extensions.
Clinical NLP for Concept Normalization
A significant portion of clinical data exists as unstructured text — progress notes, discharge summaries, pathology reports, radiology interpretations. Terminology services work on coded data, but they cannot normalize what was never coded in the first place. Clinical Natural Language Processing (NLP) bridges this gap by extracting clinical concepts from free text and mapping them to standard terminologies.
Mature NLP pipelines for clinical concept extraction include:
- Apache cTAKES: Open-source NLP system developed at Mayo Clinic for extracting clinical information from unstructured text. Uses a pipeline of sentence detection, tokenization, POS tagging, and concept lookup against UMLS.
- Amazon Comprehend Medical: Managed service that extracts medical entities (conditions, medications, dosages, procedures) and maps them to ICD-10-CM, RxNorm, and SNOMED CT codes.
- Google Cloud Healthcare Natural Language API: Extracts medical concepts and relationships from clinical text, with mapping to standard ontologies.
- MedCAT (Medical Concept Annotation Tool): Open-source tool that performs named entity recognition and linking to UMLS/SNOMED CT, with the ability to learn from local annotation data.
The key challenge in clinical NLP is not extraction — modern systems achieve high accuracy on well-defined entity types — but normalization and disambiguation. "SOB" in a clinical note could mean "shortness of breath" or "side of bed." "Discharge" could mean leaving the hospital or a wound exudate. Context-dependent disambiguation at scale remains an active research area, though recent LLM-based approaches are showing significant improvements.
Ontology-Based Data Integration
Ontology-based data integration uses formal knowledge representations — ontologies — as the semantic bridge between disparate data sources. Rather than maintaining point-to-point mappings between every pair of terminologies, an ontology provides a shared conceptual framework that multiple terminologies can map to.
The Unified Medical Language System (UMLS), maintained by the NLM, is the most comprehensive biomedical ontology integration framework. UMLS integrates over 200 source vocabularies — including SNOMED CT, ICD-10, LOINC, RxNorm, CPT, and many others — into a shared concept space (the Metathesaurus) where equivalent concepts across terminologies are linked. The UMLS Semantic Network provides a high-level categorization of concept types and their relationships.
For organizations building semantic integration pipelines, UMLS serves as the Rosetta Stone. A concept expressed in any source terminology can be mapped to its UMLS Concept Unique Identifier (CUI), and from there to any other source terminology that contains an equivalent concept. This is not perfect — many concepts have no equivalent in certain terminologies, and the mappings require ongoing maintenance — but it provides a principled approach to cross-terminology integration that scales better than point-to-point mapping.
Common Data Models: OMOP, i2b2, and PCORnet
Common data models (CDMs) take a different approach to the semantic interoperability problem. Rather than translating between terminologies in real-time, CDMs define a target schema and vocabulary to which all source data is mapped during an ETL (Extract, Transform, Load) process. Once data is in the CDM, all analytics, research queries, and decision support operate against a single, normalized representation.
The OMOP Common Data Model, maintained by the OHDSI (Observational Health Data Sciences and Informatics) community, is the most widely adopted CDM for observational research. OMOP maps dozens of source vocabularies — SNOMED, RxNorm, ICD-10, LOINC, Read codes, and others — to a standardized vocabulary, enabling OMOP to act as a Rosetta Stone for interpreting observational data across different sources and global regions. The OMOP2OBO project has produced mappings for 92,367 conditions, 8,611 drug ingredients, and 10,673 measurement results, covering 68-99% of concepts used in clinical practice across 24 hospitals.
i2b2 (Informatics for Integrating Biology and the Bedside) and the PCORnet Common Data Model serve similar purposes for clinical and patient-centered outcomes research respectively. The tradeoff with all CDM approaches is that the ETL process — mapping source data to the CDM vocabulary — is where semantic decisions are made, and those decisions require deep domain expertise in both the source terminology and the target model.
Building a Semantic Layer for Healthcare
Given the approaches above, what does a practical semantic interoperability architecture look like? Organizations that successfully bridge the semantic gap share a common architectural pattern: a dedicated semantic layer that sits between source systems and consuming applications.
Architectural Components
1. Terminology Server: The foundation. A FHIR-compliant terminology server (Ontoserver, Snowstorm, or a managed cloud service) that hosts ValueSets, CodeSystems, and ConceptMaps for all terminologies used across the organization. This server provides $translate, $lookup, $validate-code, and $expand operations as REST APIs consumed by every other component.
2. NLP Pipeline: Processes unstructured clinical text to extract coded concepts. Takes free-text clinical notes, discharge summaries, and pathology reports as input and produces coded entities (SNOMED CT, ICD-10, LOINC, RxNorm) as output. The NLP pipeline feeds into the terminology server for normalization and into the CDM for storage.
3. Ontology Mapper / CDM ETL: Transforms data from source system representations to a normalized common data model. Uses UMLS crosswalks, custom ConceptMaps, and organization-specific mapping rules to convert between terminologies. For research use cases, maps to OMOP CDM. For clinical use cases, normalizes to a preferred internal terminology (typically SNOMED CT for clinical concepts, LOINC for observations, RxNorm for medications).
4. Data Quality Engine: Validates incoming data against expected value sets, detects duplicates across systems, reconciles conflicting information (e.g., allergy lists from multiple EHRs), and flags semantic mismatches for human review. This component is essential because automated mapping is imperfect — a data quality engine ensures that low-confidence mappings are reviewed rather than silently propagated.
Implementation Considerations
Building a semantic layer is a multi-year investment. Organizations should start with the highest-impact use case — typically medication reconciliation or problem list alignment — and expand incrementally. Key decisions include:
- Synchronous vs. asynchronous: Should terminology translation happen in real-time (at the point of data exchange) or in batch (during ETL to a data lake or data warehouse)? Real-time is harder but essential for clinical decision support. Batch is sufficient for research and analytics.
- Centralized vs. federated: Should all data be mapped to a central CDM, or should mapping happen at query time across federated data sources? OMOP favors the centralized approach. FHIR-based queries with terminology expansion favor the federated approach.
- Terminology governance: Who owns the ConceptMaps? Who reviews and approves new mappings? How are mapping errors detected and corrected? Terminology governance is an organizational challenge as much as a technical one, and it requires dedicated clinical informaticists, not just engineers.
The Role of AI and LLMs in Semantic Understanding
Can large language models bridge the semantic interoperability gap? The honest answer is: partially, and with significant caveats.
LLMs have demonstrated strong capabilities in clinical concept extraction. A 2024 study in JMIR on entity extraction pipelines using large language models found that LLMs performed consistently well on binary, well-defined medical history questions, often reaching 100% accuracy. The Truveta research team has published work on LLM-Augmenter approaches that combine large language models with medical knowledge bases for improved clinical information extraction. And emerging agentic systems like CLINES (Clinical LLM-based Information Extraction and Structuring Agent) demonstrate that LLM-powered pipelines can extract, structure, and normalize clinical information from unstructured text at scale.
However, the accuracy performance varies significantly when dealing with questions that involve semantic ambiguities or definitional uncertainties. The same LLM that correctly identifies "metformin 500mg BID" as a diabetic medication may hallucinate a mapping between a rare SNOMED CT concept and an ICD-10 code that does not exist in any official crosswalk. For clinical applications where incorrect mappings can harm patients, LLM outputs must be validated against authoritative terminology services — LLMs should augment, not replace, the semantic infrastructure described above.
Where LLMs add the most value in semantic interoperability is in three specific areas:
- Free-text normalization: Converting clinician-entered free text ("pt c/o SOB x 2 days, worse w/ exertion") into coded concepts (SNOMED CT 267036007 "Dyspnea"). LLMs handle clinical abbreviations, misspellings, and context-dependent disambiguation better than rule-based NLP systems.
- Mapping assistance: Suggesting terminology mappings for concepts that are not covered by existing ConceptMaps. A human informaticist must validate the suggestion, but LLMs can reduce the effort required to maintain comprehensive mapping resources.
- Data quality review: Identifying semantic inconsistencies in patient records — for example, flagging when a medication list includes a drug contraindicated by a condition on the problem list, even when the condition and medication are coded in different terminologies.
The ONC's HTI-5 proposed rule explicitly advances a foundation for AI-enabled interoperability solutions, signaling that the regulatory framework is evolving to accommodate AI-augmented semantic interoperability. But AI is a tool within the semantic layer, not a replacement for it.
Why This Matters for Healthcare AI
Every healthcare AI application — clinical decision support, predictive analytics, population health management, precision medicine — depends on the quality of its input data. The AI community has a shorthand for this: garbage in, garbage out. But in healthcare, the problem is more insidious: it is not that the data is garbage, it is that the data is semantically inconsistent. The glucose values are real. The diagnosis codes are valid. The medication lists are accurate within each source system. But when you combine them across systems without semantic normalization, the AI model trains on noise that looks like signal.
Consider a machine learning model predicting hospital readmission risk. The model uses diagnosis codes, lab values, and medication history from multiple source systems. If diabetes is coded as E11.9 in one system, 44054006 in another, and "DM2" in free text in a third, the model treats these as three different features unless a semantic normalization step unifies them first. The model may learn that E11.9 is predictive while 44054006 is not — an artifact of data representation, not clinical reality. Multiply this across hundreds of diagnosis codes, thousands of lab tests, and tens of thousands of medications, and the model's performance degrades in ways that are difficult to diagnose because the individual data points all look correct.
This is why healthcare data lake architectures that skip the semantic normalization step produce AI models that work well on single-institution data but fail when deployed across a health system with multiple EHRs, lab systems, and pharmacy platforms. The semantic layer is not optional infrastructure — it is a prerequisite for trustworthy healthcare AI.
Organizations investing in AI capabilities without first investing in semantic infrastructure are building on sand. The data will flow, the models will train, the dashboards will render — but the clinical meaning will be inconsistent, and the downstream decisions will reflect that inconsistency. Solving semantic interoperability is not just a data engineering problem. It is the foundation on which every healthcare AI promise depends.
The Path Forward
Semantic interoperability is not a problem that will be solved by a single standard, a single technology, or a single regulatory mandate. It is a problem that requires sustained investment in terminology infrastructure, clinical informatics expertise, and organizational governance. But the organizations that make this investment will have a decisive advantage: their data will not just be exchangeable — it will be understandable, usable, and trustworthy.
The practical steps are clear even if the execution is hard:
- Deploy a terminology server as shared infrastructure. Make $translate, $lookup, and $validate-code available as internal APIs that every application can consume.
- Invest in clinical NLP to convert unstructured text into coded concepts. The volume of free-text clinical data is too large to leave unmapped.
- Adopt a common data model (OMOP for research, a FHIR-based normalized store for clinical operations) that forces semantic decisions to be made explicitly during ETL rather than silently deferred.
- Establish terminology governance with dedicated clinical informaticists who own ConceptMap quality, review automated mappings, and resolve semantic conflicts.
- Use AI as an accelerator within the semantic layer — for free-text normalization, mapping assistance, and data quality review — but never as a standalone replacement for terminology infrastructure.
The systems are connected. The next decade's challenge is making them understand each other.
Frequently Asked Questions
What is semantic interoperability in healthcare, and how is it different from FHIR interoperability?
Semantic interoperability is the ability of two healthcare systems to not only exchange data but to understand the meaning of that data in the same way. FHIR provides structural interoperability — it defines resources and fields so systems know where to find a diagnosis code or lab result — but it does not guarantee that the receiving system interprets the code the same way the sending system intended. Semantic interoperability requires shared terminologies (SNOMED CT, LOINC, RxNorm), consistent terminology binding, and bidirectional concept mapping that sits on top of FHIR, not within it. A system can be fully FHIR-compliant and still fail at semantic interoperability if its terminology usage is inconsistent with the receiving system.
Why do different EHR systems code the same diagnosis differently?
Different EHR systems use different primary terminologies based on their design, configuration, and the clinical context. Epic typically uses ICD-10-CM for diagnosis coding because of its integration with billing workflows, while Oracle Health may use SNOMED CT for clinical documentation and map to ICD-10-CM for claims. Older or smaller systems may still carry legacy ICD-9 codes or proprietary code sets. Additionally, clinicians may enter the same condition at different levels of specificity — "Type 2 diabetes" versus "Type 2 diabetes mellitus with diabetic chronic kidney disease" — creating semantic mismatches even within the same terminology system. The NLM's official SNOMED CT to ICD-10-CM map is inherently lossy because the systems have different granularity and purpose.
Can large language models solve the healthcare semantic interoperability problem?
LLMs can significantly accelerate specific aspects of semantic interoperability — particularly free-text normalization (converting clinical notes into coded concepts), mapping assistance (suggesting terminology crosswalks for unmapped concepts), and data quality review (detecting semantic inconsistencies). However, LLMs cannot replace terminology infrastructure. They hallucinate mappings that do not exist in official crosswalks, and their accuracy drops significantly on semantically ambiguous clinical concepts. The recommended approach is to use LLMs as tools within a broader semantic layer that includes a FHIR terminology server, clinical NLP pipeline, ontology mapper, and data quality engine — with human informaticists validating LLM-suggested mappings before they enter production use.
What is the OMOP Common Data Model and how does it address semantic interoperability?
The OMOP (Observational Medical Outcomes Partnership) Common Data Model, maintained by the OHDSI community, addresses semantic interoperability by defining a single target schema and vocabulary to which all source data is mapped during an ETL process. Rather than translating between terminologies in real-time, OMOP forces semantic decisions to be made upfront: source codes from SNOMED CT, ICD-10, LOINC, RxNorm, and other terminologies are mapped to OMOP standard concepts. Once data is in OMOP, all queries operate against normalized data. The OMOP2OBO project has produced validated mappings covering 68-99% of clinical concepts across 24 hospitals, making OMOP the most practical approach for multi-site observational research requiring consistent semantic alignment.
How should a healthcare organization start building a semantic interoperability strategy?
Start with the highest-impact, most tractable use case — typically medication reconciliation or problem list alignment across your EHR systems. Deploy a FHIR-compliant terminology server (Ontoserver, Snowstorm, or a cloud-managed service like Google Cloud Healthcare API) as shared infrastructure. Establish terminology governance with at least one dedicated clinical informaticist who owns ConceptMap quality. Then expand incrementally: add clinical NLP for unstructured text normalization, adopt OMOP CDM for research analytics, and layer AI-assisted mapping as your terminology infrastructure matures. The key principle is to make semantic decisions explicitly and systematically rather than deferring them to downstream applications. Budget for a multi-year investment — semantic interoperability is infrastructure, not a project.