A radiology group in Texas built a RAG pipeline to help their AI agent answer clinical questions about patient history. They embedded PDF discharge summaries and lab reports. The pipeline worked well in demos. In production, it missed 34% of relevant medications because the PDF chunker split a patient's medication list across two chunks, and the retriever only returned one. The medications were right there in the FHIR MedicationRequest resources, structured and complete, but the pipeline never touched them.
This is the fundamental problem with healthcare RAG today: teams embed PDFs and clinical notes while ignoring the structured FHIR data that already contains the clinical semantics they need. A Condition resource with a SNOMED CT code of 73211009 tells you the patient has diabetes mellitus. A chunk of text that mentions "blood sugar has been running high" might or might not mean the same thing. FHIR resources carry clinical meaning in their structure, not just their text.
This guide shows you how to build a RAG pipeline that embeds FHIR resources directly, preserves clinical semantics through the embedding process, and evaluates retrieval quality with metrics that matter for clinical applications.

Why Standard RAG Pipelines Fail on Clinical Data
Standard RAG pipelines were designed for documents: PDFs, web pages, knowledge base articles. They assume text is the primary carrier of meaning. In healthcare, structured data carries as much clinical meaning as narrative text, often more.
Problem 1: Naive Chunking Destroys Clinical Context
A FHIR Bundle containing a patient's problem list might include 15 Condition resources, each with coded diagnoses, onset dates, clinical status, and verification status. A standard text chunker splitting at 512 tokens will cut this list at arbitrary points, separating a condition from its onset date or splitting a medication from its dosage instructions.
In a comprehensive review of RAG in healthcare, researchers found that document-level chunking strategies consistently underperformed resource-aware chunking for clinical question answering, with recall dropping 15-25% when clinical context boundaries were ignored.
Problem 2: General Embeddings Miss Clinical Semantics
When you embed the text "Patient has type 2 diabetes mellitus" using a general-purpose embedding model like text-embedding-3-small, the vector captures the semantic meaning of the English sentence. But it does not capture the clinical relationship between diabetes and its associated conditions (diabetic nephropathy, retinopathy), its medication implications (metformin as first-line), or its SNOMED CT hierarchy (73211009 is a descendant of Endocrine disorder).
Clinical embedding models trained on medical literature and EHR data encode these relationships in vector space, making retrieval more clinically relevant rather than just textually similar.
Problem 3: Retrieval Evaluation Ignores Clinical Relevance
Standard RAG evaluation measures like Recall@K and NDCG tell you whether the retriever found textually relevant chunks. They do not tell you whether the retrieved chunks contain the clinically correct information for the query. A chunk mentioning "elevated glucose" is textually relevant to a diabetes query but clinically incomplete without the HbA1c value, diagnosis date, and current treatment plan.

Step 1: FHIR Resource Extraction and Normalization
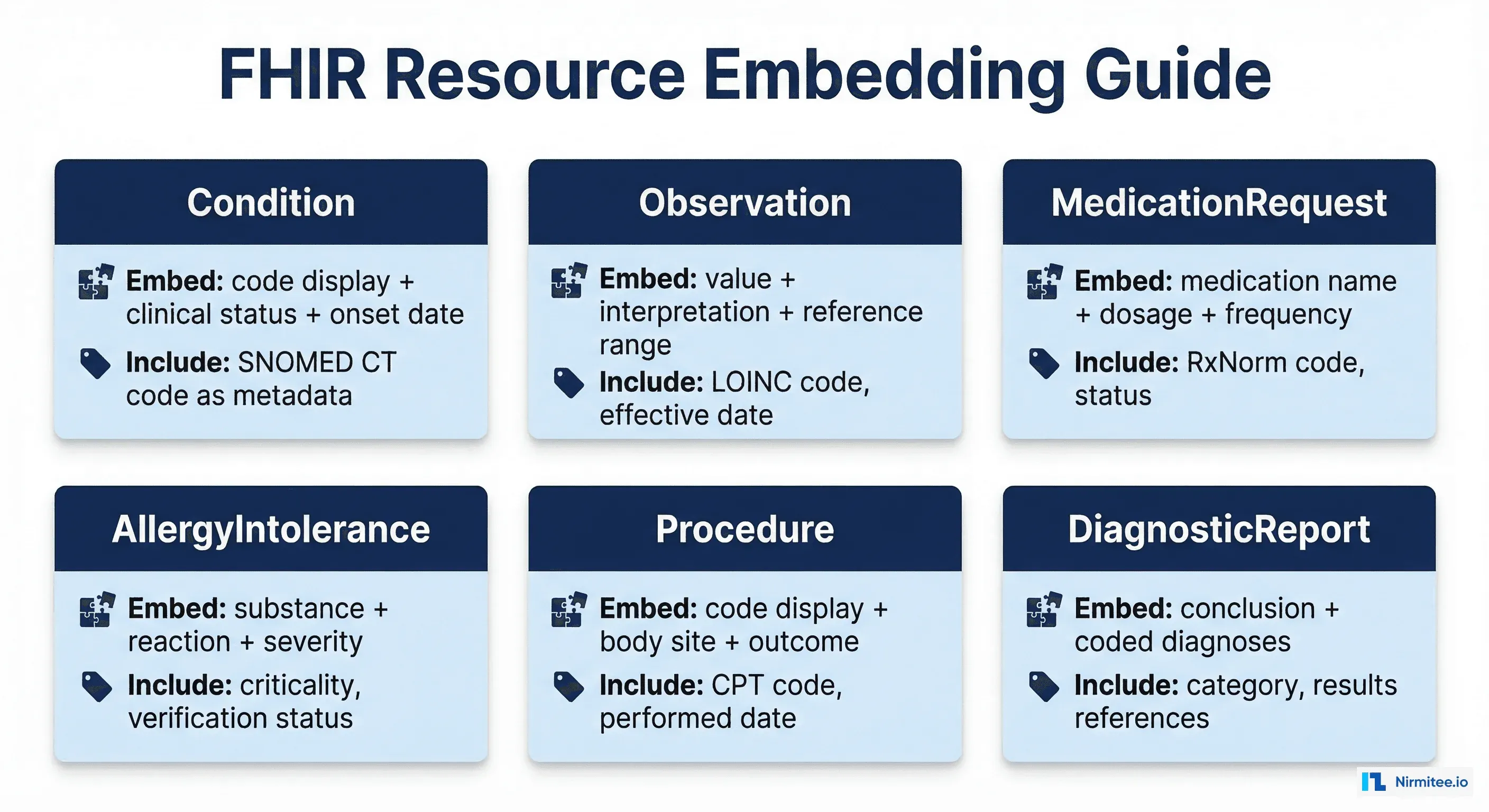
The first step is extracting FHIR resources and converting them into embeddable text that preserves clinical semantics. This is not a simple JSON-to-text serialization. Each resource type needs a specific extraction strategy.
The FHIR-to-Text Pipeline
import json
from typing import List, Dict, Optional
from dataclasses import dataclass
@dataclass
class ClinicalChunk:
text: str
resource_type: str
fhir_resource_id: str
patient_id: str
snomed_codes: List[str]
loinc_codes: List[str]
effective_date: Optional[str]
metadata: Dict
def extract_condition(resource: dict) -> ClinicalChunk:
"""Extract embeddable text from a FHIR Condition resource."""
code = resource.get("code", {})
display = code.get("coding", [{}])[0].get("display", "Unknown condition")
snomed = code.get("coding", [{}])[0].get("code", "")
clinical_status = resource.get("clinicalStatus", {}).get(
"coding", [{}])[0].get("code", "unknown")
verification = resource.get("verificationStatus", {}).get(
"coding", [{}])[0].get("code", "unknown")
onset = resource.get("onsetDateTime", "unknown date")
severity = resource.get("severity", {}).get(
"coding", [{}])[0].get("display", "")
# Structured clinical text — not just the display name
text_parts = [
f"Diagnosis: {display}",
f"Clinical status: {clinical_status}",
f"Verification: {verification}",
f"Onset: {onset}",
]
if severity:
text_parts.append(f"Severity: {severity}")
return ClinicalChunk(
text=" | ".join(text_parts),
resource_type="Condition",
fhir_resource_id=resource["id"],
patient_id=resource.get("subject", {}).get("reference", "").split("/")[-1],
snomed_codes=[snomed] if snomed else [],
loinc_codes=[],
effective_date=onset if onset != "unknown date" else None,
metadata={"category": resource.get("category", [{}])[0].get(
"coding", [{}])[0].get("code", "")}
)
Step 2: Clinical Chunking Strategies
Once you have extracted text from individual FHIR resources, the next decision is how to group them into chunks for embedding. There are three strategies, each with different tradeoffs.
Strategy 1: Resource-Level Chunking (Recommended Starting Point)
One FHIR resource equals one chunk. A single Condition resource becomes one embedding, a single Observation becomes one embedding. This is the simplest approach and preserves resource integrity perfectly.
Pros: Clean boundaries, easy to trace back to source, simple to update when resources change.
Cons: Misses cross-resource relationships. A query about "diabetes management" would need to retrieve the Condition, related MedicationRequest resources, and recent Observation values (HbA1c, glucose) separately.
Strategy 2: Encounter-Level Chunking
Group all resources from a single encounter into one chunk. A clinic visit produces one chunk containing the conditions addressed, observations recorded, medications prescribed, and procedures performed during that visit.
Pros: Captures the clinical context of a single visit, which is often the relevant unit for clinical questions.
Cons: Chunks can be very large for complex encounters (ICU stays, surgical episodes). May exceed embedding model context limits.
Strategy 3: Clinical Concept Chunking (Highest Retrieval Quality)
Group resources by clinical concept using SNOMED CT hierarchies. All resources related to a patient's diabetes (the Condition, HbA1c Observation values, metformin MedicationRequest, diabetic retinopathy screening Procedure) become one chunk.
Pros: Highest retrieval relevance for clinical questions. A single query about diabetes retrieves the complete clinical picture.
Cons: Requires a clinical ontology mapper to group resources by concept. Harder to implement but significantly better retrieval quality.
Step 3: Choosing Your Embedding Model
The embedding model determines how well your vector space captures clinical semantics. General-purpose models work, but clinical models consistently outperform them on healthcare retrieval tasks by 8-15% on clinical concept embedding benchmarks.
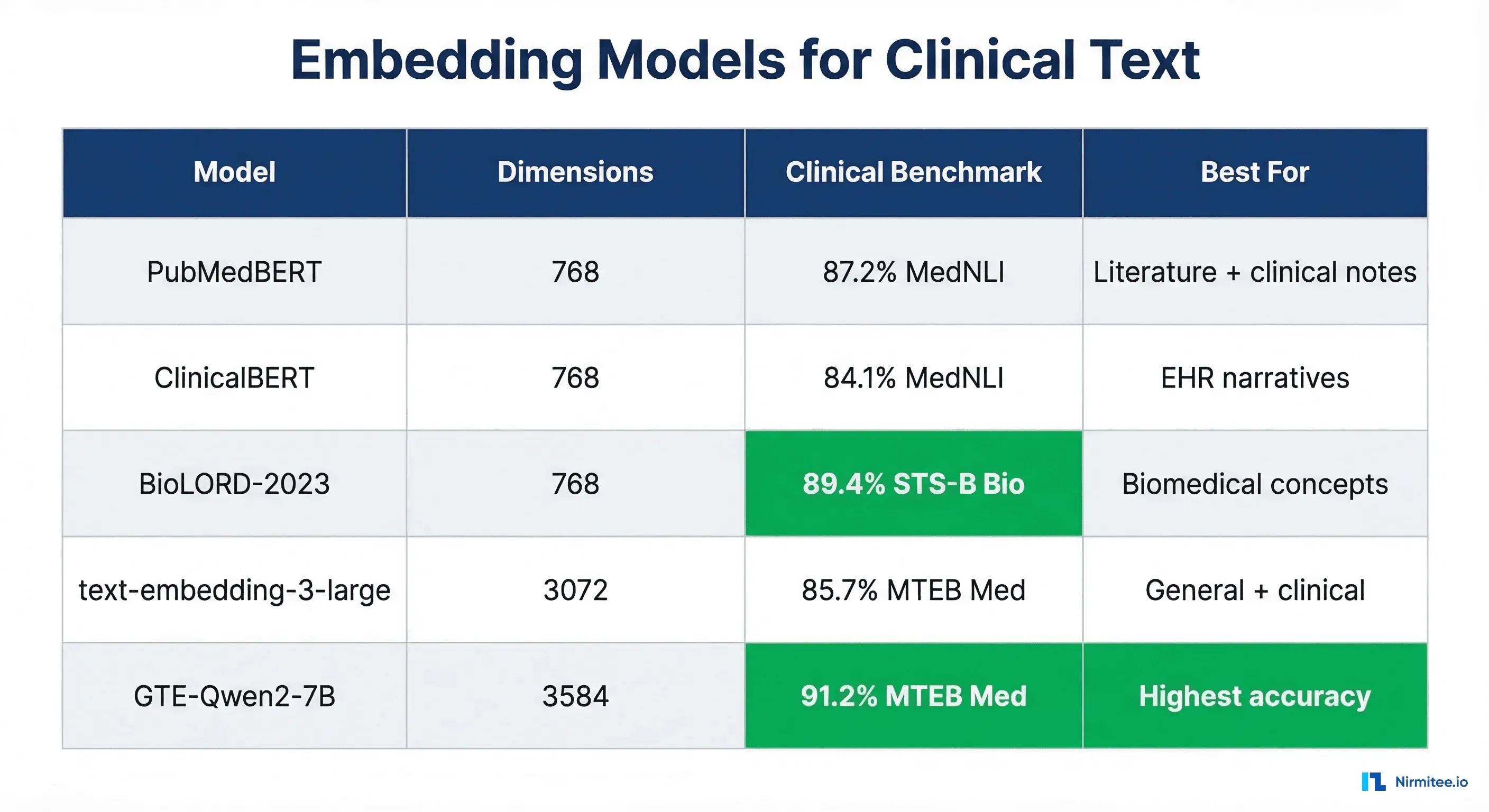
Our Recommendation
For most healthcare RAG pipelines, start with PubMedBERT or BioLORD-2023. They offer the best balance of clinical accuracy, dimension size (768, manageable for pgvector), and inference cost. Move to larger models like GTE-Qwen2 only if you need maximum retrieval accuracy and can afford the compute and storage costs of 3584-dimensional vectors.
Step 4: Vector Store Setup with pgvector
For healthcare RAG, pgvector is the pragmatic choice. You already have PostgreSQL for your FHIR data store and application data. Adding vector search to the same database simplifies HIPAA compliance (one database to audit, one set of access controls) and enables hybrid queries that combine vector similarity with structured FHIR metadata filters.
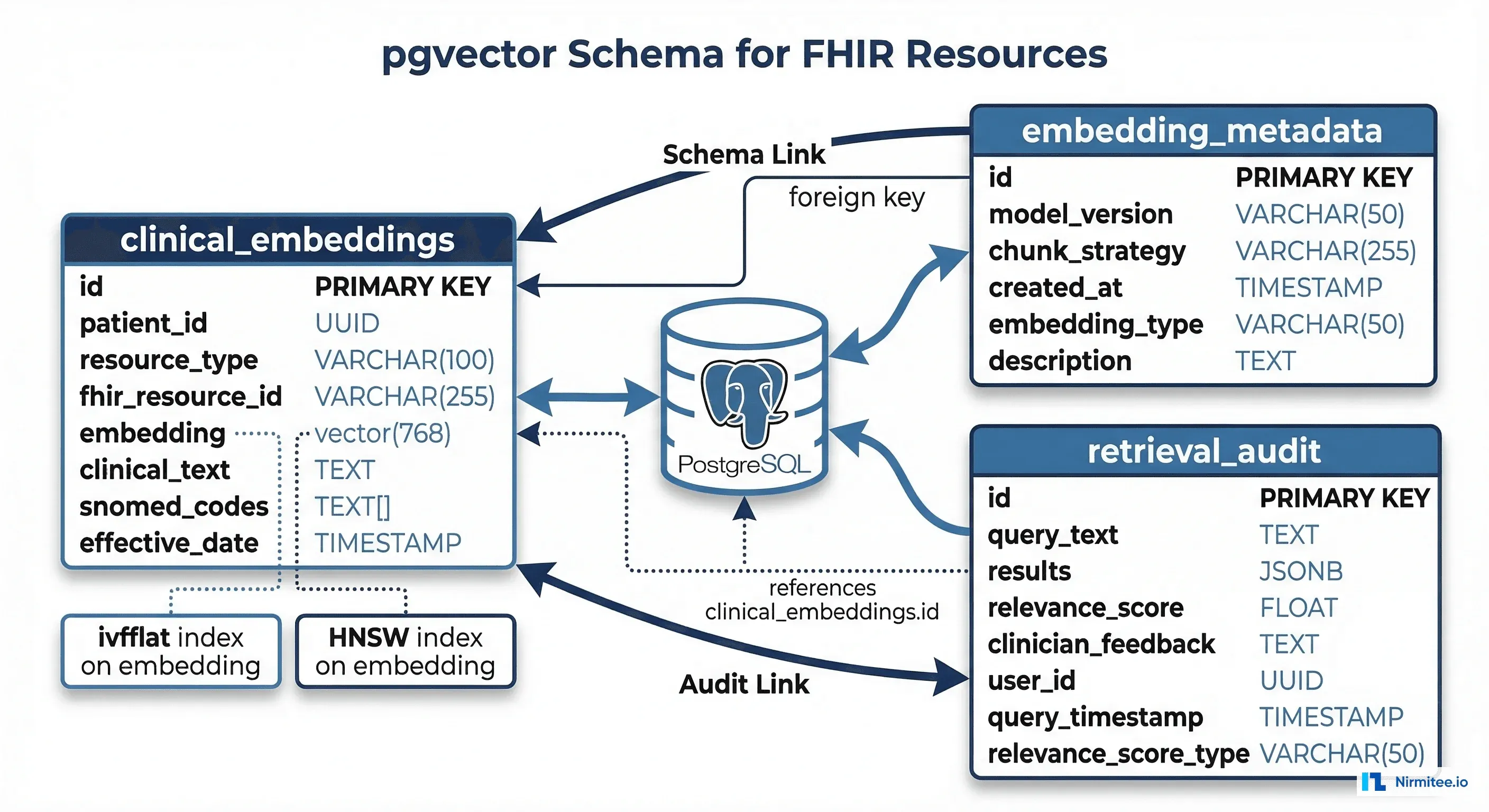
Schema Design
-- Enable pgvector extension
CREATE EXTENSION IF NOT EXISTS vector;
-- Clinical embeddings table
CREATE TABLE clinical_embeddings (
id SERIAL PRIMARY KEY,
patient_id UUID NOT NULL REFERENCES patients(id),
resource_type VARCHAR(50) NOT NULL,
fhir_resource_id VARCHAR(255) NOT NULL,
embedding vector(768) NOT NULL,
clinical_text TEXT NOT NULL,
snomed_codes TEXT[] DEFAULT '{}',
loinc_codes TEXT[] DEFAULT '{}',
effective_date TIMESTAMP,
embedding_model VARCHAR(100) NOT NULL,
chunk_strategy VARCHAR(50) NOT NULL,
created_at TIMESTAMP DEFAULT NOW(),
UNIQUE(fhir_resource_id, embedding_model)
);
-- HNSW index for approximate nearest neighbor search
CREATE INDEX idx_embeddings_hnsw
ON clinical_embeddings
USING hnsw (embedding vector_cosine_ops)
WITH (m = 16, ef_construction = 128);
-- Filtered search indexes
CREATE INDEX idx_embeddings_patient ON clinical_embeddings(patient_id);
CREATE INDEX idx_embeddings_resource ON clinical_embeddings(resource_type);
CREATE INDEX idx_embeddings_snomed ON clinical_embeddings USING gin(snomed_codes);Hybrid Retrieval Query
The power of pgvector in a FHIR context is combining vector similarity with structured metadata filters in a single query:
-- Find relevant clinical context for a medication review agent
SELECT
ce.clinical_text,
ce.resource_type,
ce.fhir_resource_id,
ce.snomed_codes,
1 - (ce.embedding <=> query_embedding) AS similarity
FROM clinical_embeddings ce
WHERE
ce.patient_id = $1
AND ce.resource_type IN ('Condition', 'MedicationRequest', 'AllergyIntolerance')
AND ce.effective_date > NOW() - INTERVAL '2 years'
ORDER BY ce.embedding <=> query_embedding
LIMIT 20;This query retrieves the 20 most semantically similar clinical chunks, filtered to only active conditions, medications, and allergies from the last 2 years. A pure vector store cannot do this without post-retrieval filtering, which wastes retrieval budget on irrelevant results.
Step 5: Clinical Retrieval Evaluation
Standard RAG evaluation is not sufficient for clinical applications. You need metrics that capture clinical relevance, not just textual similarity. Based on systematic reviews of RAG in biomedical applications, we use a three-tier evaluation framework.
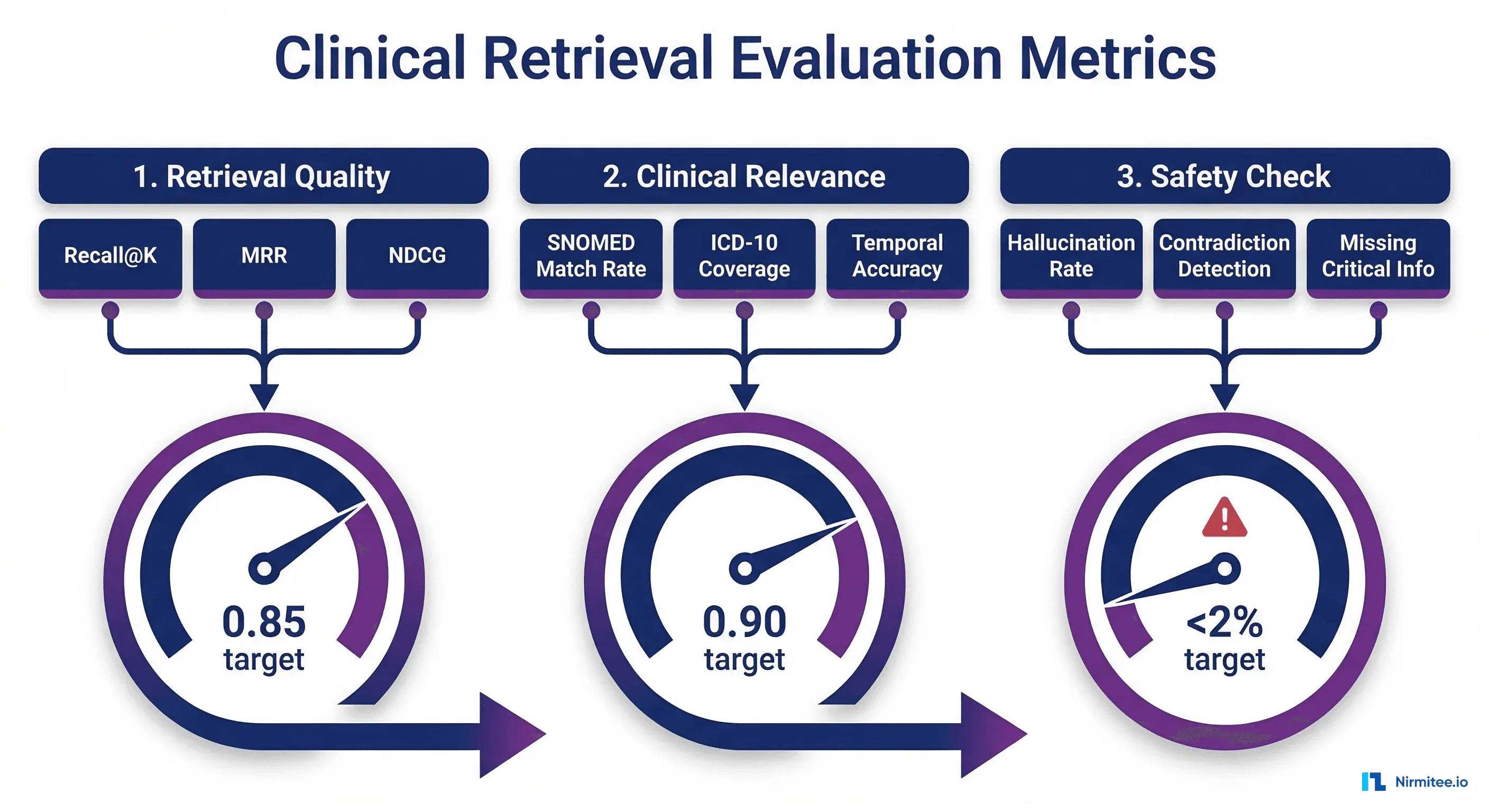
Tier 1: Standard Retrieval Metrics
| Metric | Target | What It Measures |
|---|---|---|
| Recall@10 | >0.85 | Did the top 10 results include the relevant resources? |
| MRR | >0.70 | How high was the first relevant result ranked? |
| NDCG@10 | >0.80 | Were relevant results ranked higher than less relevant ones? |
Tier 2: Clinical Relevance Metrics
| Metric | Target | What It Measures |
|---|---|---|
| SNOMED Match Rate | >0.90 | Did retrieved chunks contain the correct SNOMED codes? |
| Clinical Completeness | >0.85 | Did retrieval capture all clinically necessary information? |
| Temporal Accuracy | >0.95 | Are retrieved results from the clinically relevant time window? |
Tier 3: Safety Metrics
| Metric | Target | What It Measures |
|---|---|---|
| Critical Miss Rate | <2% | How often does retrieval miss allergy or contraindication data? |
| Hallucination Rate | <1% | Does the LLM generate information not in retrieved context? |
| Contradiction Rate | <0.5% | Does the response contradict retrieved FHIR resources? |
Pinecone Alternative: When to Choose Managed Vector Search
pgvector is ideal when you are already running PostgreSQL and want to minimize your HIPAA compliance surface area. Choose Pinecone or Weaviate when you need sub-10ms retrieval at scale (1M+ embeddings), dedicated vector search infrastructure with automatic scaling, or when your team does not want to manage pgvector index tuning. Both offer HIPAA BAAs and SOC 2 compliance.
Production Considerations
Embedding Updates When FHIR Resources Change
FHIR resources are not static. A Condition can change from active to resolved. A MedicationRequest can be cancelled. Your embedding pipeline needs to detect these changes and update vectors accordingly. Use Change Data Capture (CDC) on your FHIR server to trigger re-embedding when resources are created or updated.
PHI in Vector Embeddings
Embeddings derived from PHI are themselves PHI under HIPAA. Your vector store needs the same access controls, encryption at rest, and audit logging as your FHIR server. This is another reason pgvector is attractive: it inherits PostgreSQL's existing security infrastructure.
Model Versioning
When you upgrade your embedding model, you need to re-embed your entire corpus. The embedding_model column in our schema supports running both old and new embeddings simultaneously, with a gradual cutover after validating retrieval quality. Track this in your model registry alongside your inference models.
Getting Started
Begin with resource-level chunking using PubMedBERT and pgvector. This combination ships fast, integrates with your existing PostgreSQL infrastructure, and gives you a solid baseline to iterate from. Add clinical concept chunking and more sophisticated embedding models as your retrieval evaluation metrics tell you where the pipeline is falling short.
The key insight is that FHIR resources already carry clinical semantics in their structure. Your RAG pipeline should embed that structure, not flatten it into unstructured text and hope the embedding model reconstructs the meaning.
At Nirmitee, we build FHIR-native data pipelines for healthcare AI systems. From FHIR agent integration to production RAG pipelines with clinical evaluation, our team has the healthcare data engineering expertise to get your clinical AI from prototype to production.