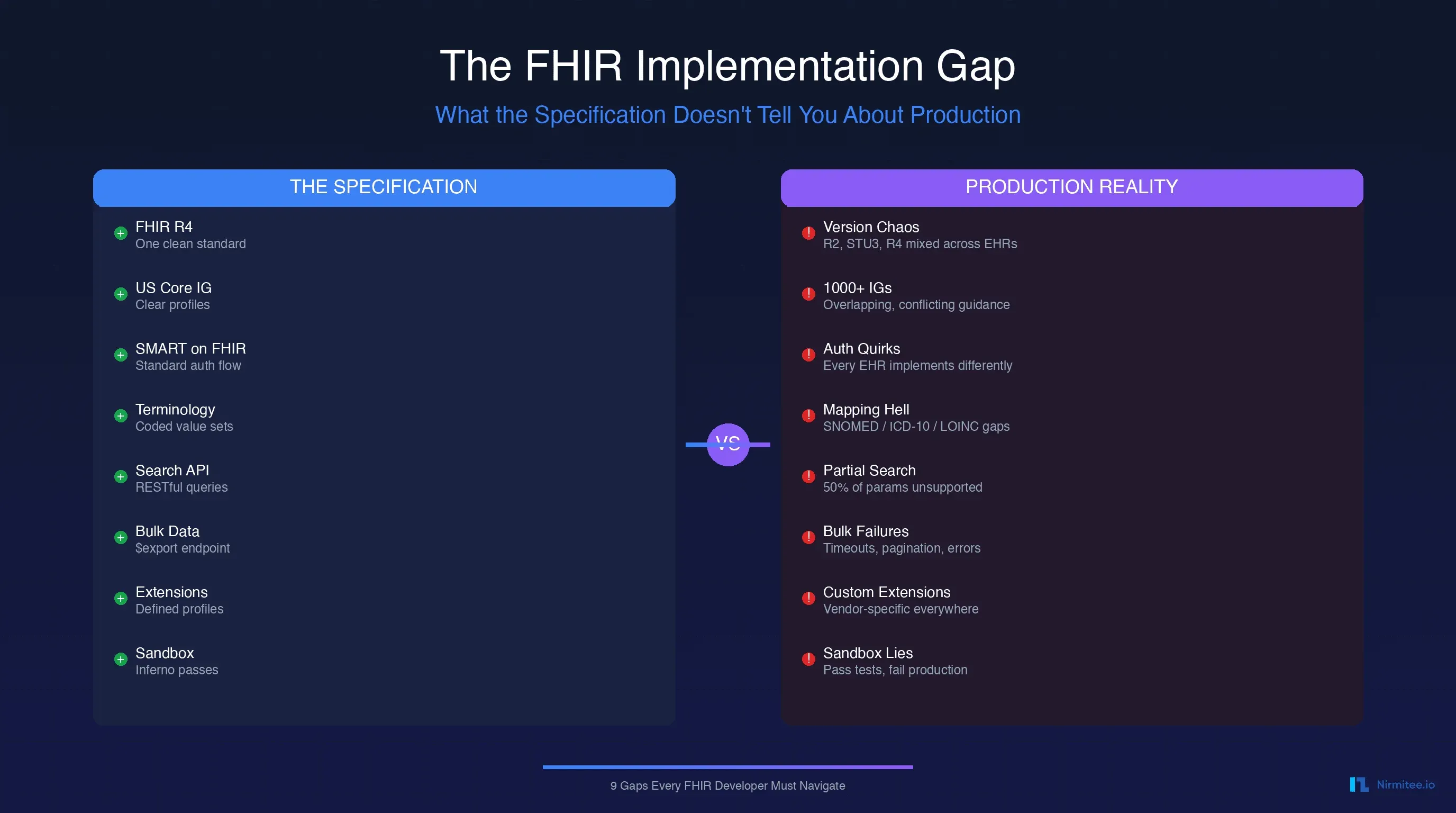
Every healthcare organization has data governance on its roadmap. Few have it in production. The gap between intent and implementation is enormous — and the consequences are real: a researcher pulls a report from a dashboard showing readmission rates dropping by 15%, presents it to the board, and nobody can answer the question "Where did this number come from?"
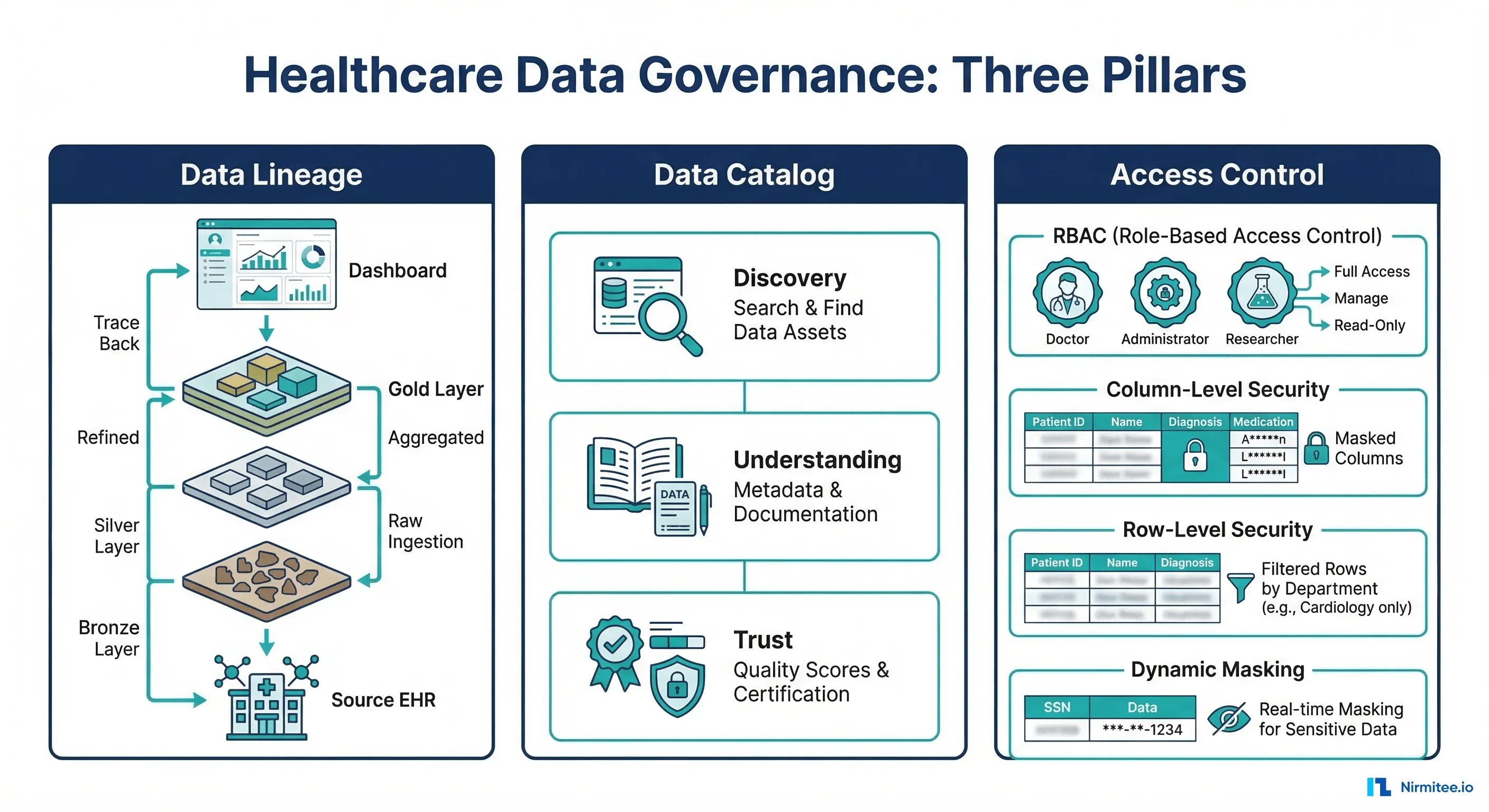
Healthcare data governance is not a single tool or policy. It is the combination of three capabilities: data lineage (tracing data from the dashboard back to source), data catalog (discovering and understanding what data exists), and access control (ensuring the right people see the right data at the right level of detail). Layer on top of that the unique requirements of PHI — HIPAA minimum 6-year retention, dynamic masking for researchers vs. clinicians, column-level security for SSN and DOB fields — and you have a complex but essential infrastructure challenge.
This guide provides production-ready patterns for healthcare data governance, with code examples for Unity Catalog, Apache Ranger, column-level security, and automated data quality monitoring.
Data Lineage: Where Did This Number Come From?
Why Lineage Matters in Healthcare
When a quality officer reports HEDIS measures to CMS, or when a researcher publishes findings from your clinical data warehouse, they need to trust the numbers. Data lineage provides that trust by answering:
- Provenance: What source system did this data originate from? (Epic? Cerner? Lab instrument?)
- Transformation history: What ETL steps were applied? Were codes mapped? Were duplicates removed?
- Freshness: When was this data last updated? Is it current enough for clinical decisions?
- Impact analysis: If we change the source schema, what downstream reports and dashboards break?
Implementing Lineage with OpenLineage
OpenLineage is an open standard for lineage metadata that works with Spark, Airflow, dbt, and other pipeline tools. Here is how to instrument a healthcare ETL pipeline:
# lineage_instrumented_etl.py
# Healthcare ETL with OpenLineage instrumentation
from openlineage.client import OpenLineageClient
from openlineage.client.run import (
RunEvent, RunState, Run, Job, Dataset, InputDataset, OutputDataset
)
from openlineage.client.facet import (
SchemaField, SchemaDatasetFacet,
DataQualityMetricsInputDatasetFacet
)
import uuid
from datetime import datetime
client = OpenLineageClient(url="http://lineage-server:5000")
namespace = "healthcare-data-platform"
def emit_lineage_event(
job_name, run_id, state,
inputs, outputs, quality_metrics=None
):
"""Emit an OpenLineage event for pipeline tracking."""
input_datasets = []
for inp in inputs:
facets = {}
if quality_metrics and inp["name"] in quality_metrics:
facets["dataQuality"] = DataQualityMetricsInputDatasetFacet(
rowCount=quality_metrics[inp["name"]].get("row_count"),
bytes=quality_metrics[inp["name"]].get("bytes"),
)
input_datasets.append(
InputDataset(
namespace=namespace,
name=inp["name"],
facets=facets
)
)
output_datasets = [
OutputDataset(
namespace=namespace,
name=out["name"],
facets={
"schema": SchemaDatasetFacet(
fields=[
SchemaField(name=f["name"], type=f["type"])
for f in out.get("fields", [])
]
)
}
)
for out in outputs
]
event = RunEvent(
eventType=state,
eventTime=datetime.utcnow().isoformat() + "Z",
run=Run(runId=str(run_id)),
job=Job(namespace=namespace, name=job_name),
inputs=input_datasets,
outputs=output_datasets,
)
client.emit(event)
# Example: Lab results ETL with lineage
run_id = uuid.uuid4()
# START event
emit_lineage_event(
job_name="lab-results-bronze-to-silver",
run_id=run_id,
state=RunState.START,
inputs=[
{"name": "bronze.lab.fhir_observations"},
{"name": "reference.loinc_codes"},
],
outputs=[
{"name": "silver.lab.validated_results",
"fields": [
{"name": "patient_id", "type": "STRING"},
{"name": "loinc_code", "type": "STRING"},
{"name": "result_value", "type": "DECIMAL"},
{"name": "result_unit", "type": "STRING"},
{"name": "reference_range_low", "type": "DECIMAL"},
{"name": "reference_range_high", "type": "DECIMAL"},
{"name": "effective_date", "type": "TIMESTAMP"},
]}
]
)
# ... run actual ETL ...
# COMPLETE event with quality metrics
emit_lineage_event(
job_name="lab-results-bronze-to-silver",
run_id=run_id,
state=RunState.COMPLETE,
inputs=[
{"name": "bronze.lab.fhir_observations"},
],
outputs=[
{"name": "silver.lab.validated_results",
"fields": []}
],
quality_metrics={
"bronze.lab.fhir_observations": {
"row_count": 1_247_893,
"bytes": 3_200_000_000,
}
}
)Data Catalog: Discover, Understand, Trust
Choosing a Catalog for Healthcare
| Catalog | Strengths | Healthcare Fit |
|---|---|---|
| Unity Catalog (Databricks) | Deep Delta Lake integration, built-in access control, column masking | Best if already on Databricks; native PHI masking and row-level security |
| Apache Atlas | Open-source, strong lineage, Hadoop ecosystem | Good for on-premise health systems; requires significant ops investment |
| DataHub (LinkedIn) | Open-source, modern UI, broad integrations | Best open-source option; extensible metadata model for PHI classification |
| Collibra | Enterprise governance, business glossary, stewardship workflows | Best for large IDNs (integrated delivery networks) with dedicated governance teams |
| Alation | SQL query recommendations, behavioral analysis | Good for analyst-heavy organizations; strong search and discovery |
Unity Catalog Setup for Healthcare
-- Unity Catalog: Healthcare data governance setup
-- Step 1: Create catalog and schemas
CREATE CATALOG IF NOT EXISTS health_system;
USE CATALOG health_system;
CREATE SCHEMA IF NOT EXISTS clinical_bronze
COMMENT 'Raw clinical data ingested from EHR systems (FHIR NDJSON, HL7v2)';
CREATE SCHEMA IF NOT EXISTS clinical_silver
COMMENT 'Cleaned, validated, LOINC/SNOMED coded clinical data';
CREATE SCHEMA IF NOT EXISTS clinical_gold
COMMENT 'Business-ready data products: aggregated metrics, OMOP CDM tables';
CREATE SCHEMA IF NOT EXISTS research_omop
COMMENT 'OMOP CDM tables for observational research (de-identified)';
-- Step 2: Create tables with column-level tags
CREATE TABLE clinical_gold.patient_demographics (
patient_id STRING COMMENT 'Internal patient identifier',
mrn STRING COMMENT 'Medical Record Number'
TAGS ('phi_level' = 'direct_identifier'),
first_name STRING COMMENT 'Patient first name'
TAGS ('phi_level' = 'direct_identifier'),
last_name STRING COMMENT 'Patient last name'
TAGS ('phi_level' = 'direct_identifier'),
ssn STRING COMMENT 'Social Security Number'
TAGS ('phi_level' = 'direct_identifier'),
date_of_birth DATE COMMENT 'Date of birth'
TAGS ('phi_level' = 'quasi_identifier'),
zip_code STRING COMMENT '5-digit ZIP code'
TAGS ('phi_level' = 'quasi_identifier'),
gender STRING COMMENT 'Administrative gender',
race STRING COMMENT 'Race category',
ethnicity STRING COMMENT 'Ethnicity category',
primary_language STRING COMMENT 'Preferred language',
insurance_type STRING COMMENT 'Primary insurance category'
)
COMMENT 'Patient demographics with PHI classification tags'
TBLPROPERTIES (
'data_product.domain' = 'patient_access',
'data_product.owner' = 'patient-data-team',
'data_product.freshness_sla' = 'daily',
'compliance.hipaa' = 'true',
'retention.years' = '10'
);
-- Step 3: Create column masking functions
CREATE FUNCTION mask_ssn(ssn STRING)
RETURNS STRING
RETURN CONCAT('***-**-', RIGHT(ssn, 4));
CREATE FUNCTION mask_name(name STRING)
RETURNS STRING
RETURN 'REDACTED';
CREATE FUNCTION mask_dob_year_only(dob DATE)
RETURNS STRING
RETURN CAST(YEAR(dob) AS STRING);
-- Step 4: Apply column masks by role
-- Researchers see masked PHI
ALTER TABLE clinical_gold.patient_demographics
ALTER COLUMN ssn SET MASK mask_ssn
USING COLUMNS (ssn)
WHERE IS_MEMBER('researcher_group');
ALTER TABLE clinical_gold.patient_demographics
ALTER COLUMN first_name SET MASK mask_name
USING COLUMNS (first_name)
WHERE IS_MEMBER('researcher_group');
ALTER TABLE clinical_gold.patient_demographics
ALTER COLUMN last_name SET MASK mask_name
USING COLUMNS (last_name)
WHERE IS_MEMBER('researcher_group');
ALTER TABLE clinical_gold.patient_demographics
ALTER COLUMN date_of_birth SET MASK mask_dob_year_only
USING COLUMNS (date_of_birth)
WHERE IS_MEMBER('researcher_group');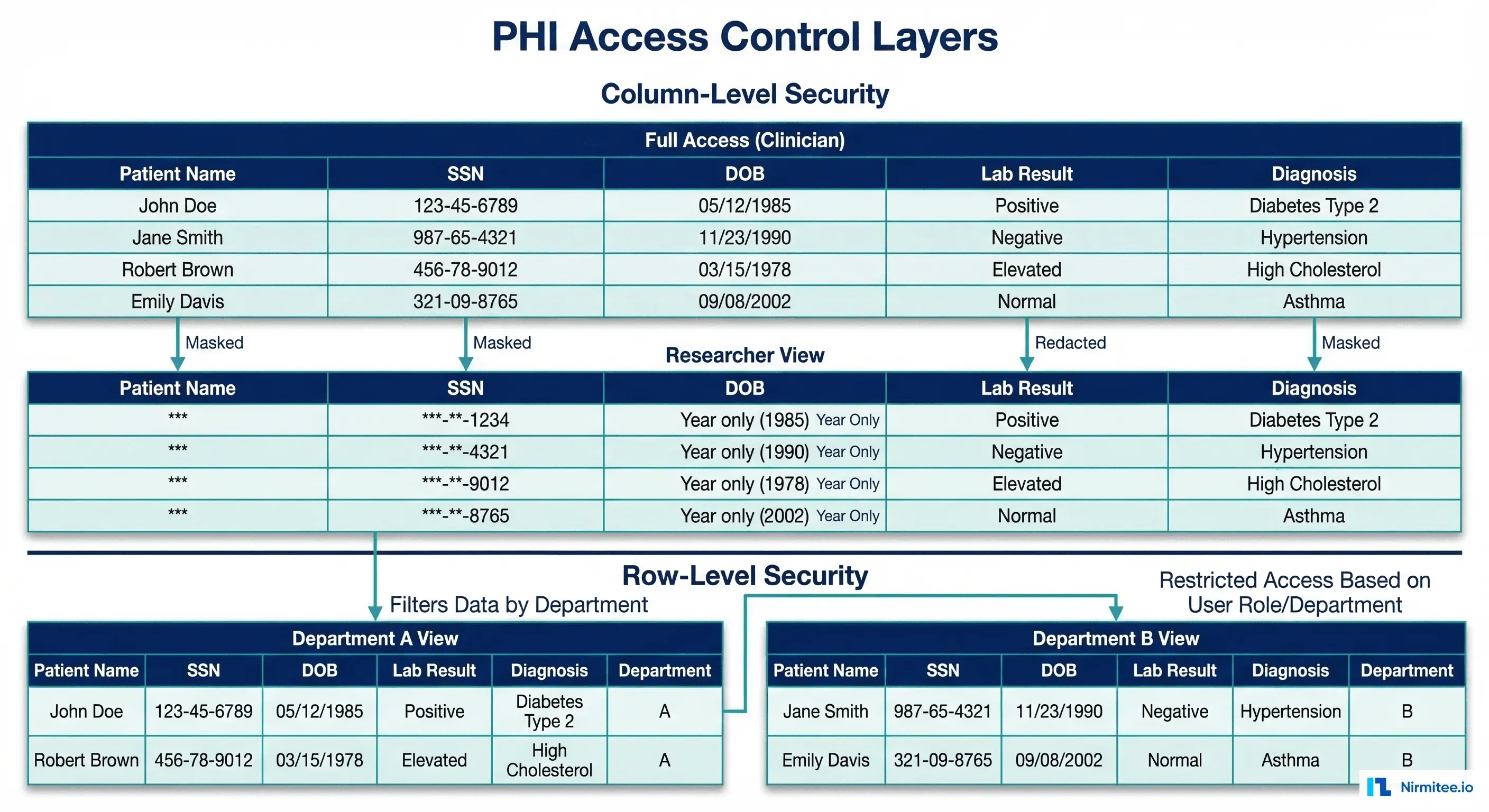
Access Control for PHI at Scale
The Three Layers of Healthcare Access Control
Healthcare data access requires three simultaneous controls that most general-purpose systems do not provide out of the box:
- Role-Based Access Control (RBAC): Clinicians, researchers, billing analysts, and administrators each have different base permissions
- Column-Level Security: PHI fields (SSN, name, DOB, MRN) are masked or hidden based on the user's role and purpose
- Row-Level Security: Users see only patients in their department, facility, or care team — not the entire population
Row-Level Security Implementation
-- Row-level security: Department-scoped access
-- Users see only patients from their department/facility
-- Step 1: Create a mapping table of user-to-facility access
CREATE TABLE governance.user_facility_access (
user_email STRING,
facility_id STRING,
department STRING,
access_level STRING -- 'full', 'aggregate', 'de-identified'
);
-- Step 2: Create row filter function
CREATE FUNCTION filter_by_facility(facility_id STRING)
RETURNS BOOLEAN
RETURN (
EXISTS (
SELECT 1 FROM governance.user_facility_access ufa
WHERE ufa.user_email = current_user()
AND ufa.facility_id = filter_by_facility.facility_id
)
OR IS_MEMBER('enterprise_admin_group')
);
-- Step 3: Apply row filter to clinical tables
ALTER TABLE clinical_gold.lab_results
SET ROW FILTER filter_by_facility ON (facility_id);
ALTER TABLE clinical_gold.patient_encounters
SET ROW FILTER filter_by_facility ON (facility_id);
-- Now: Dr. Smith at Hospital A queries lab_results
-- She sees only Hospital A patients automatically
-- Enterprise analytics team (admin group) sees all facilities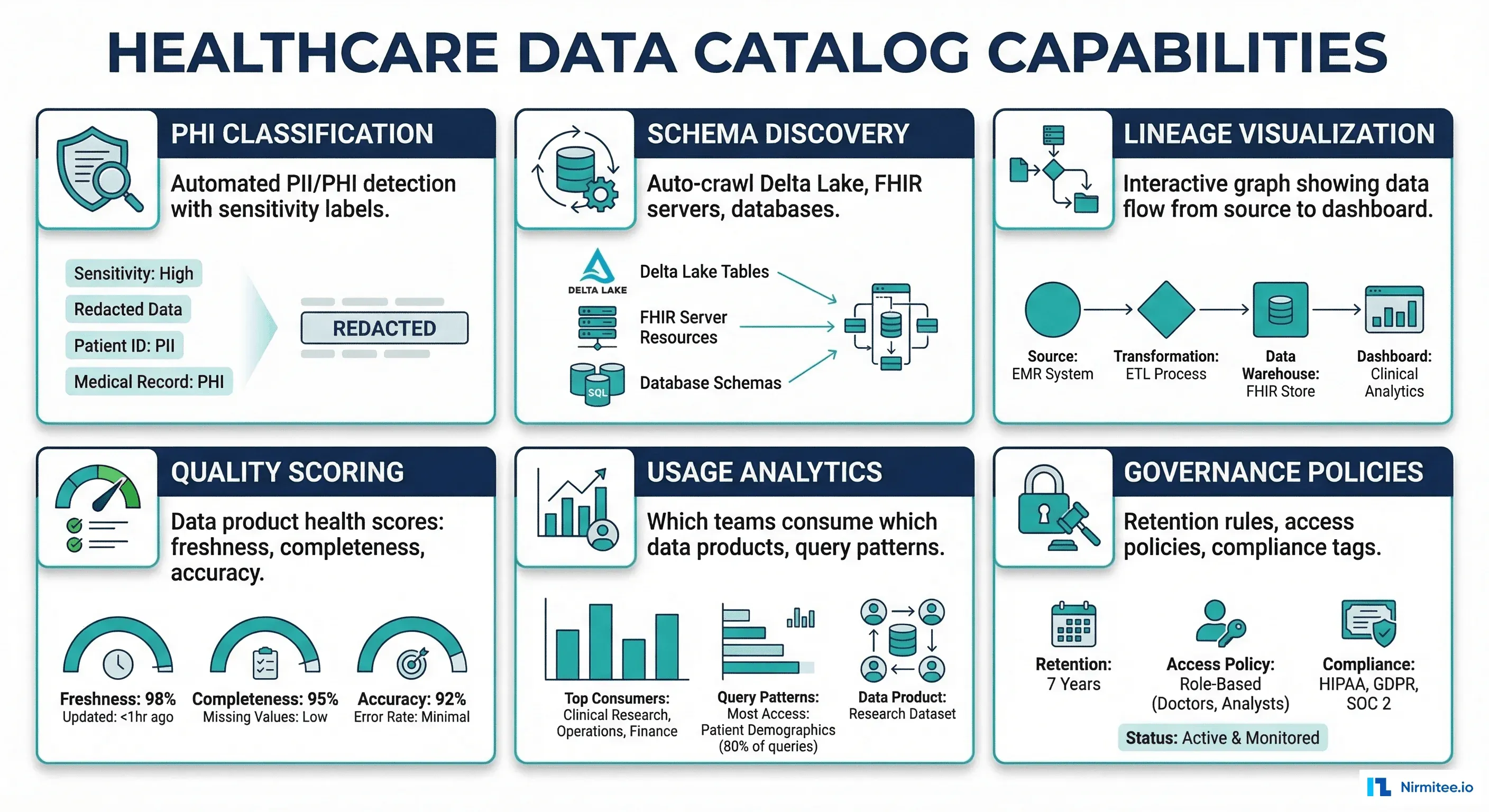
Dynamic Data Masking Policies
# dynamic_masking_policies.py
# Define masking policies for different consumer roles
MASKING_POLICIES = {
"treating_clinician": {
"description": "Full access to patients in their care team",
"patient_name": "VISIBLE",
"ssn": "VISIBLE",
"date_of_birth": "VISIBLE",
"mrn": "VISIBLE",
"diagnoses": "VISIBLE",
"lab_results": "VISIBLE",
"row_filter": "patient IN user.care_team",
"audit": "standard (logged, not alerted)"
},
"researcher": {
"description": "De-identified access for IRB-approved research",
"patient_name": "REDACTED",
"ssn": "LAST_4_ONLY", # ***-**-1234
"date_of_birth": "YEAR_ONLY", # 1985
"mrn": "HASHED", # SHA-256 pseudonym
"diagnoses": "VISIBLE",
"lab_results": "VISIBLE",
"row_filter": "patient.consent_for_research = TRUE",
"audit": "enhanced (logged + quarterly review)"
},
"billing_analyst": {
"description": "Financial data with minimal clinical detail",
"patient_name": "REDACTED",
"ssn": "REDACTED",
"date_of_birth": "REDACTED",
"mrn": "VISIBLE", # needed for charge reconciliation
"diagnoses": "ICD_CODE_ONLY", # code visible, not narrative
"lab_results": "HIDDEN",
"row_filter": "encounter.has_open_claim = TRUE",
"audit": "standard"
},
"quality_analyst": {
"description": "Aggregate metrics only, no individual patient data",
"patient_name": "REDACTED",
"ssn": "REDACTED",
"date_of_birth": "AGE_BUCKET", # 18-34, 35-49, 50-64, 65+
"mrn": "REDACTED",
"diagnoses": "VISIBLE",
"lab_results": "AGGREGATE_ONLY", # mean, median, percentiles
"row_filter": "none (aggregate queries only)",
"minimum_cell_size": 11, # suppress cells with < 11 patients
"audit": "standard"
},
"break_the_glass": {
"description": "Emergency full access with mandatory justification",
"patient_name": "VISIBLE",
"ssn": "VISIBLE",
"date_of_birth": "VISIBLE",
"mrn": "VISIBLE",
"diagnoses": "VISIBLE",
"lab_results": "VISIBLE",
"row_filter": "none",
"requires": "written justification + supervisor notification",
"audit": "immediate alert to privacy officer",
"auto_expire": "24 hours"
}
}Data Quality Contracts
Automated Quality Monitoring
Every data product must have machine-readable quality contracts that are continuously monitored:
# data_quality_monitor.py
# Automated data quality monitoring for healthcare data products
from great_expectations.core import ExpectationSuite
from great_expectations.dataset import SparkDFDataset
from datetime import datetime, timedelta
import json
def build_lab_results_quality_suite():
"""Define quality expectations for lab results data product."""
suite = ExpectationSuite(
expectation_suite_name="lab_results_quality"
)
# Completeness checks
suite.add_expectation({
"expectation_type": "expect_column_values_to_not_be_null",
"kwargs": {"column": "loinc_code", "mostly": 0.995},
"meta": {"severity": "critical",
"description": "99.5% of results must have LOINC code"}
})
suite.add_expectation({
"expectation_type": "expect_column_values_to_not_be_null",
"kwargs": {"column": "result_value", "mostly": 0.99},
"meta": {"severity": "critical",
"description": "99% of results must have a value"}
})
suite.add_expectation({
"expectation_type": "expect_column_values_to_not_be_null",
"kwargs": {"column": "result_unit", "mostly": 0.99},
"meta": {"severity": "warning",
"description": "99% should have UCUM units"}
})
# Freshness check
suite.add_expectation({
"expectation_type": "expect_column_max_to_be_between",
"kwargs": {
"column": "effective_datetime",
"min_value": (datetime.utcnow() - timedelta(minutes=30)).isoformat(),
"max_value": datetime.utcnow().isoformat()
},
"meta": {"severity": "critical",
"description": "Most recent result within 30 minutes"}
})
# Clinical plausibility checks
suite.add_expectation({
"expectation_type": "expect_column_values_to_be_between",
"kwargs": {
"column": "result_value",
"min_value": 0,
"max_value": 100000,
"mostly": 0.999
},
"meta": {"severity": "warning",
"description": "Result values within instrument range"}
})
# Volume anomaly detection
suite.add_expectation({
"expectation_type": "expect_table_row_count_to_be_between",
"kwargs": {
"min_value": 5000, # minimum daily volume
"max_value": 50000 # maximum daily volume
},
"meta": {"severity": "critical",
"description": "Daily volume within expected range"}
})
return suite
def check_quality_and_alert(dataset, suite):
"""Run quality checks and alert on failures."""
results = dataset.validate(suite)
failures = [
r for r in results.results
if not r.success
]
if failures:
critical = [
f for f in failures
if f.expectation_config.meta.get("severity") == "critical"
]
if critical:
send_alert(
channel="pagerduty",
severity="high",
message=f"CRITICAL: {len(critical)} data quality "
f"checks failed for lab results",
details=json.dumps([
{
"check": f.expectation_config.expectation_type,
"description": f.expectation_config.meta.get(
"description", ""
),
}
for f in critical
])
)
return results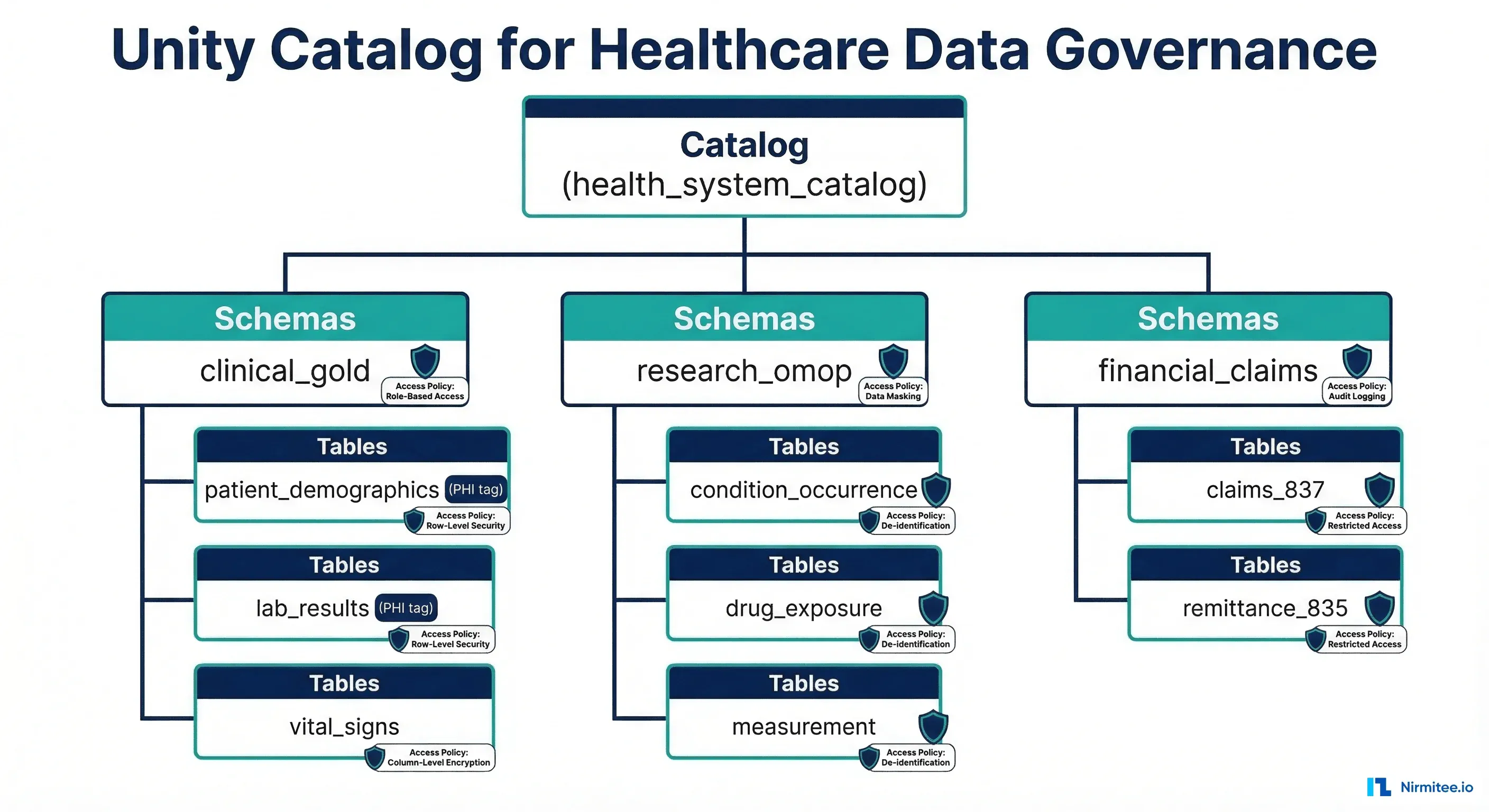
Retention Policies and Automated Purging
Healthcare Retention Requirements
| Data Type | Minimum Retention | Source | Notes |
|---|---|---|---|
| Medical records (general) | 6 years | HIPAA 45 CFR 164.530(j) | From date of creation or last effective date |
| Medical records (state) | 7-10 years (varies) | State medical record laws | California: 7 years; New York: 6 years; Florida: 7 years |
| Minor patient records | Until age 21 + state retention | State laws | Could extend to 25+ years for newborns |
| Medicare/Medicaid records | 10 years | 42 CFR 482.24 | Includes claims, cost reports, eligibility records |
| Clinical trial data | Per protocol (15+ years typical) | 21 CFR Part 11, ICH GCP | Sponsor determines retention; often indefinite |
| HIPAA policies and procedures | 6 years | 45 CFR 164.530(j) | From date of creation or last effective date |
| Audit logs (PHI access) | 6 years | HIPAA Security Rule | All system activity logs involving PHI |
| Employee training records | 6 years from training date | HIPAA | Including HIPAA training certifications |
Automated Retention and Purging
# retention_manager.py
# Automated retention policy enforcement for healthcare data
from dataclasses import dataclass
from datetime import datetime, timedelta
from typing import Optional
import logging
logger = logging.getLogger("retention_manager")
@dataclass
class RetentionPolicy:
data_type: str
retention_years: int
source_regulation: str
has_legal_hold: bool = False
minor_patient_extension: bool = False
POLICIES = {
"medical_records": RetentionPolicy(
data_type="medical_records",
retention_years=10, # using max of HIPAA + most states
source_regulation="HIPAA + State Law (max)",
minor_patient_extension=True
),
"claims_data": RetentionPolicy(
data_type="claims_data",
retention_years=10,
source_regulation="42 CFR 482.24 (Medicare)"
),
"audit_logs": RetentionPolicy(
data_type="audit_logs",
retention_years=6,
source_regulation="HIPAA Security Rule"
),
"research_data": RetentionPolicy(
data_type="research_data",
retention_years=25, # conservative default
source_regulation="21 CFR Part 11 / Protocol"
),
"analytics_derived": RetentionPolicy(
data_type="analytics_derived",
retention_years=3, # non-PHI derived data
source_regulation="Internal policy"
),
}
def calculate_purge_date(
record_date: datetime,
policy: RetentionPolicy,
patient_dob: Optional[datetime] = None
) -> datetime:
"""Calculate when a record can be purged."""
base_purge = record_date + timedelta(
days=policy.retention_years * 365
)
# Minor extension: retain until age 21 + retention period
if policy.minor_patient_extension and patient_dob:
age_21 = patient_dob + timedelta(days=21 * 365)
minor_purge = age_21 + timedelta(
days=policy.retention_years * 365
)
base_purge = max(base_purge, minor_purge)
return base_purge
def run_retention_scan(dry_run: bool = True):
"""Scan all data products and identify records eligible for purging."""
now = datetime.utcnow()
for policy_name, policy in POLICIES.items():
if policy.has_legal_hold:
logger.info(
f"Skipping {policy_name}: legal hold active"
)
continue
# Query for records past retention date
logger.info(
f"Scanning {policy_name} "
f"(retain {policy.retention_years} years)..."
)
# In production: query Delta Lake / database
# eligible_count = spark.sql(f"""
# SELECT COUNT(*) FROM {policy_name}
# WHERE purge_eligible_date < current_timestamp()
# AND legal_hold = FALSE
# """).first()[0]
if dry_run:
logger.info(f" DRY RUN: would purge eligible records")
else:
logger.info(f" PURGING eligible records...")
# Execute deletion with audit trailPutting It All Together: Governance Architecture
# governance-architecture.yaml
# Complete healthcare data governance stack
governance_stack:
layer_1_catalog:
primary: "Unity Catalog" # or DataHub for non-Databricks
features:
- automated_crawling: "Delta Lake, FHIR server, databases"
- phi_classification: "ML-based PII/PHI detection on ingest"
- lineage: "OpenLineage integration for end-to-end tracking"
- search: "natural language search across all data assets"
- quality_scores: "real-time health scores per data product"
layer_2_access_control:
primary: "Unity Catalog ACLs + Apache Ranger"
capabilities:
rbac: "role-based by job function (clinician, researcher, analyst)"
column_masking: "dynamic PHI masking per role"
row_filtering: "department/facility scoped access"
break_the_glass: "emergency access with justification + alert"
audit: "all PHI access logged, 6-year retention, quarterly review"
layer_3_quality:
primary: "Great Expectations + custom clinical validators"
monitoring:
freshness: "continuous SLA monitoring per data product"
completeness: "critical field null rates checked hourly"
plausibility: "clinical range validation on ingest"
volume: "anomaly detection on daily record counts"
alerting:
critical: "PagerDuty immediate alert"
warning: "Slack notification to domain team"
info: "dashboard update, weekly digest"
layer_4_retention:
primary: "Custom retention manager + Delta Lake time travel"
policies:
medical_records: "10 years (HIPAA + state max)"
claims: "10 years (Medicare requirement)"
audit_logs: "6 years (HIPAA Security Rule)"
research: "per protocol (25 years default)"
automation:
scan_frequency: "weekly"
legal_hold: "supported, overrides purge schedule"
purge_method: "soft delete -> 90 day grace -> hard delete"
layer_5_compliance:
hipaa:
risk_assessment: "annual, documented in governance catalog"
baa_tracking: "all data sharing agreements tracked"
breach_notification: "automated detection + 60-day reporting"
state_laws:
tracking: "per-state requirements mapped to retention policies"
updates: "quarterly review of regulatory changes"From architecture to production, our Healthcare Software Product Development team builds healthcare platforms that perform at scale. We also offer specialized Healthcare Interoperability Solutions services. Talk to our team to get started.
Frequently Asked Questions
Which data catalog should we use for healthcare?
If you are on Databricks, Unity Catalog is the clear choice — it integrates natively with Delta Lake and provides column masking, row filtering, and lineage out of the box. For multi-cloud or non-Databricks environments, DataHub (open-source) or Collibra (enterprise) are strong options.
The key healthcare requirement is automated PHI classification — whatever catalog you choose must tag sensitive columns automatically and enforce access policies based on those tags.
How do we handle break-the-glass access for emergencies?
Break-the-glass (BTG) access must be available for genuine clinical emergencies where a provider needs to see a patient's full record outside normal access controls. Implementation: allow override with mandatory justification text, immediate notification to the privacy officer, automatic expiration (typically 24 hours), and post-hoc review.
The key is making BTG access frictionless for real emergencies while creating enough accountability to prevent abuse. Most EHRs have BTG built in; extend this pattern to your data mesh analytics layer.
How granular should column-level security be?
Follow the HIPAA Safe Harbor method as your baseline: the 18 identifier types (name, SSN, DOB, address, etc.) must be masked or removed for non-treatment uses. Beyond that, apply the minimum necessary standard — each role should see only the data elements required for their job function.
Billing analysts need MRN and diagnosis codes, but not lab results. Researchers need diagnosis and lab data, but not names. Quality analysts need aggregate metrics, not individual records.
What about de-identification for research?
HIPAA provides two de-identification methods: Safe Harbor (remove/generalize 18 specific identifiers) and Expert Determination (a qualified statistician certifies re-identification risk is very small). For most data governance implementations, apply Safe Harbor through dynamic masking policies.
Expert Determination is used for datasets that need more granular data (e.g., keeping exact dates for time-series analysis) and require engagement with a qualified privacy expert.
How do we audit data access at scale?
Centralize audit logs from all systems (EHR, data lake, analytics tools, applications) into a tamper-proof log store (e.g., AWS CloudTrail, Azure Monitor, or a dedicated SIEM). Set up automated alerts for: access to sensitive records outside the care team, bulk data exports, BTG events, and access patterns that deviate from baseline. HIPAA requires 6 years of audit log retention. Use OpenTelemetry to correlate data access traces across your entire pipeline.