Why Snowflake for Healthcare Data
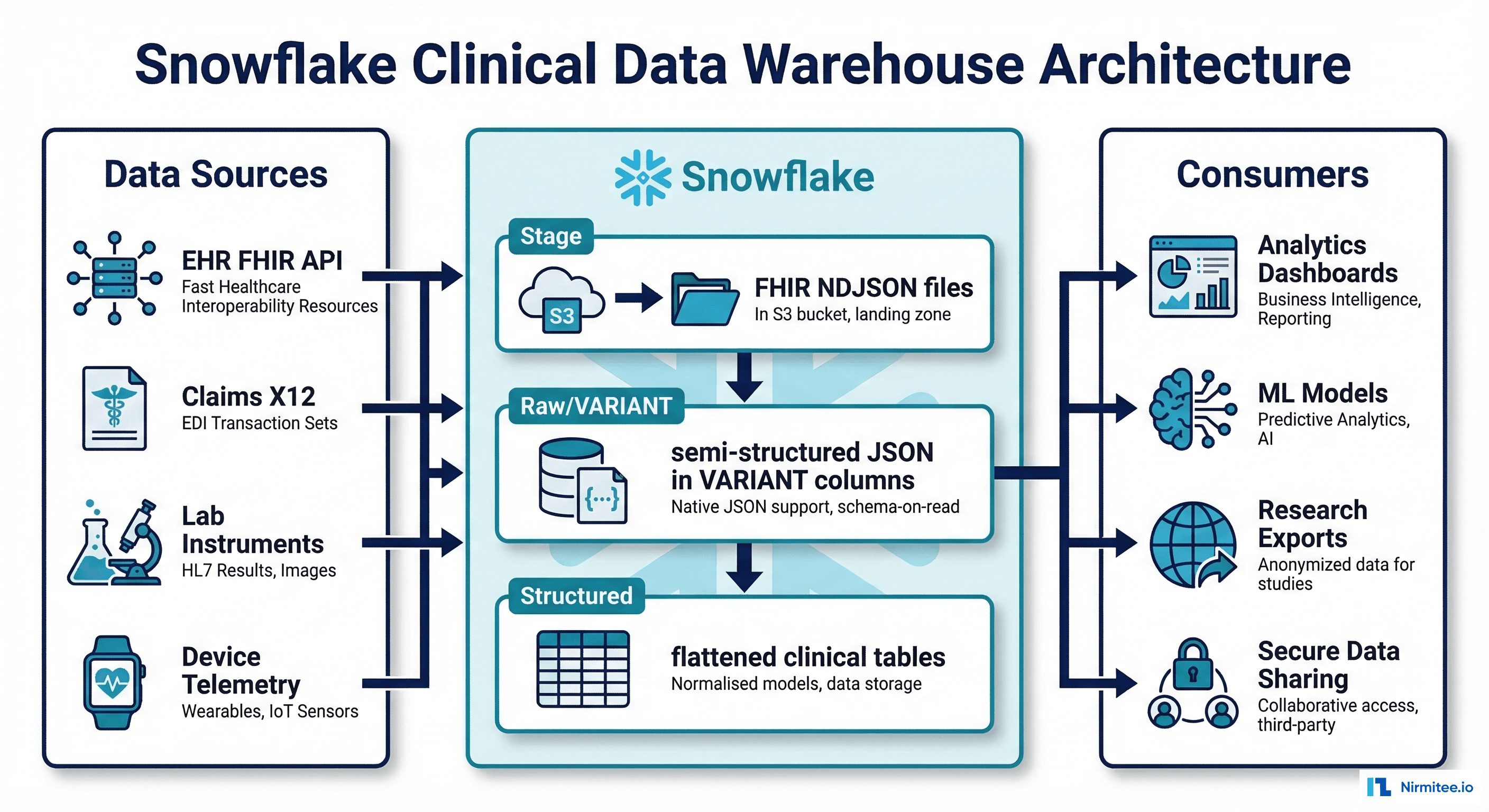
Healthcare organizations generate vast amounts of semi-structured clinical data — FHIR JSON resources, HL7v2 messages, claims EDI files — that traditional relational databases struggle to handle efficiently. Snowflake's VARIANT data type and LATERAL FLATTEN function make it uniquely suited for loading, querying, and analyzing FHIR JSON at scale without forcing a rigid schema upfront.
Beyond JSON handling, Snowflake offers three capabilities that healthcare data teams increasingly depend on: Secure Data Sharing (share analytics with research partners without copying data), Dynamic Data Masking (PHI fields automatically masked based on the querying user's role), and Snowpark (run Python/Java/Scala ML pipelines directly on Snowflake compute, keeping data in place). Together, these features address the core challenges of building a clinical data warehouse that is both analytically powerful and HIPAA-compliant.
This guide covers the full implementation: loading FHIR NDJSON, flattening nested resources, setting up masking policies, configuring secure shares, and building ML features with Snowpark.

Loading FHIR NDJSON into Snowflake
Stage and COPY INTO with VARIANT
FHIR Bulk Data Export produces NDJSON files — one JSON object per line, one file per resource type. Snowflake loads these directly into VARIANT columns without any schema transformation:
-- Step 1: Create file format for FHIR NDJSON
CREATE OR REPLACE FILE FORMAT fhir_ndjson_format
TYPE = 'JSON'
STRIP_OUTER_ARRAY = FALSE
STRIP_NULL_VALUES = FALSE
COMPRESSION = 'AUTO';
-- Step 2: Create external stage pointing to S3/Azure/GCS
CREATE OR REPLACE STAGE fhir_bulk_export_stage
URL = 's3://ehr-fhir-exports/bulk-export/'
CREDENTIALS = (AWS_KEY_ID = '...' AWS_SECRET_KEY = '...')
FILE_FORMAT = fhir_ndjson_format;
-- Step 3: Create raw tables with VARIANT columns
CREATE OR REPLACE TABLE raw_fhir.patients (
raw_data VARIANT,
source_file STRING DEFAULT METADATA$FILENAME,
load_timestamp TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP()
);
CREATE OR REPLACE TABLE raw_fhir.conditions (
raw_data VARIANT,
source_file STRING DEFAULT METADATA$FILENAME,
load_timestamp TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP()
);
CREATE OR REPLACE TABLE raw_fhir.observations (
raw_data VARIANT,
source_file STRING DEFAULT METADATA$FILENAME,
load_timestamp TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP()
);
CREATE OR REPLACE TABLE raw_fhir.medication_requests (
raw_data VARIANT,
source_file STRING DEFAULT METADATA$FILENAME,
load_timestamp TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP()
);
-- Step 4: Load FHIR resources
COPY INTO raw_fhir.patients
FROM @fhir_bulk_export_stage/Patient/
FILE_FORMAT = fhir_ndjson_format
PATTERN = '.*Patient.*\.ndjson'
ON_ERROR = 'CONTINUE';
COPY INTO raw_fhir.conditions
FROM @fhir_bulk_export_stage/Condition/
FILE_FORMAT = fhir_ndjson_format
PATTERN = '.*Condition.*\.ndjson'
ON_ERROR = 'CONTINUE';
COPY INTO raw_fhir.observations
FROM @fhir_bulk_export_stage/Observation/
FILE_FORMAT = fhir_ndjson_format
PATTERN = '.*Observation.*\.ndjson'
ON_ERROR = 'CONTINUE';
-- Verify load
SELECT COUNT(*) AS patient_count FROM raw_fhir.patients;
SELECT COUNT(*) AS condition_count FROM raw_fhir.conditions;
SELECT COUNT(*) AS observation_count FROM raw_fhir.observations;Flattening FHIR Resources with LATERAL FLATTEN
Patient Demographics
-- Flatten FHIR Patient resources into relational format
CREATE OR REPLACE VIEW clinical.patient_demographics AS
SELECT
raw_data:id::STRING AS patient_id,
raw_data:identifier[0].value::STRING AS mrn,
raw_data:name[0].family::STRING AS last_name,
raw_data:name[0].given[0]::STRING AS first_name,
raw_data:birthDate::DATE AS date_of_birth,
raw_data:gender::STRING AS gender,
raw_data:telecom[0].value::STRING AS phone,
raw_data:address[0].line[0]::STRING AS address_line,
raw_data:address[0].city::STRING AS city,
raw_data:address[0].state::STRING AS state,
raw_data:address[0].postalCode::STRING AS zip_code,
-- US Core Race extension
raw_data:extension[0].extension[0].valueCoding.display::STRING AS race,
-- US Core Ethnicity extension
raw_data:extension[1].extension[0].valueCoding.display::STRING AS ethnicity,
raw_data:meta.lastUpdated::TIMESTAMP_NTZ AS last_updated
FROM raw_fhir.patients;Lab Results with Component Flattening
-- Flatten Observation resources (including multi-component like Blood Pressure)
CREATE OR REPLACE VIEW clinical.lab_results AS
-- Simple observations (single value)
SELECT
raw_data:id::STRING AS observation_id,
raw_data:subject.reference::STRING AS patient_ref,
raw_data:code.coding[0].system::STRING AS code_system,
raw_data:code.coding[0].code::STRING AS loinc_code,
raw_data:code.coding[0].display::STRING AS test_name,
raw_data:valueQuantity.value::FLOAT AS result_value,
raw_data:valueQuantity.unit::STRING AS result_unit,
raw_data:referenceRange[0].low.value::FLOAT AS ref_range_low,
raw_data:referenceRange[0].high.value::FLOAT AS ref_range_high,
raw_data:effectiveDateTime::TIMESTAMP_NTZ AS effective_date,
raw_data:status::STRING AS status,
'single' AS observation_type
FROM raw_fhir.observations
WHERE raw_data:valueQuantity IS NOT NULL
UNION ALL
-- Multi-component observations (e.g., Blood Pressure)
SELECT
raw_data:id::STRING AS observation_id,
raw_data:subject.reference::STRING AS patient_ref,
comp.value:code.coding[0].system::STRING AS code_system,
comp.value:code.coding[0].code::STRING AS loinc_code,
comp.value:code.coding[0].display::STRING AS test_name,
comp.value:valueQuantity.value::FLOAT AS result_value,
comp.value:valueQuantity.unit::STRING AS result_unit,
NULL AS ref_range_low,
NULL AS ref_range_high,
raw_data:effectiveDateTime::TIMESTAMP_NTZ AS effective_date,
raw_data:status::STRING AS status,
'component' AS observation_type
FROM raw_fhir.observations,
LATERAL FLATTEN(input => raw_data:component) comp
WHERE raw_data:component IS NOT NULL;Conditions with Multiple Codings
-- Flatten Condition resources with all coding systems
CREATE OR REPLACE VIEW clinical.conditions AS
SELECT
raw_data:id::STRING AS condition_id,
raw_data:subject.reference::STRING AS patient_ref,
coding.value:system::STRING AS code_system,
coding.value:code::STRING AS diagnosis_code,
coding.value:display::STRING AS diagnosis_display,
raw_data:clinicalStatus.coding[0].code::STRING AS clinical_status,
raw_data:verificationStatus.coding[0].code::STRING AS verification_status,
raw_data:onsetDateTime::DATE AS onset_date,
raw_data:abatementDateTime::DATE AS abatement_date,
raw_data:recordedDate::DATE AS recorded_date,
raw_data:category[0].coding[0].code::STRING AS category
FROM raw_fhir.conditions,
LATERAL FLATTEN(input => raw_data:code.coding) coding;Dynamic Data Masking for PHI
Masking Policy Setup
Snowflake's dynamic data masking applies column-level masking rules based on the querying user's role — the underlying data remains unchanged, and different users see different views of the same table:
-- Create masking policies for PHI fields
-- SSN masking: show last 4 to researchers, full to clinicians
CREATE OR REPLACE MASKING POLICY phi_ssn_mask AS
(val STRING) RETURNS STRING ->
CASE
WHEN IS_ROLE_IN_SESSION('TREATING_CLINICIAN') THEN val
WHEN IS_ROLE_IN_SESSION('CARE_TEAM_MEMBER') THEN val
WHEN IS_ROLE_IN_SESSION('RESEARCHER') THEN
CONCAT('***-**-', RIGHT(val, 4))
ELSE '***-**-****'
END;
-- Name masking: visible to clinicians, redacted for others
CREATE OR REPLACE MASKING POLICY phi_name_mask AS
(val STRING) RETURNS STRING ->
CASE
WHEN IS_ROLE_IN_SESSION('TREATING_CLINICIAN') THEN val
WHEN IS_ROLE_IN_SESSION('CARE_TEAM_MEMBER') THEN val
WHEN IS_ROLE_IN_SESSION('RESEARCHER') THEN
CONCAT('Patient-', MD5(val)::STRING(8))
ELSE 'REDACTED'
END;
-- DOB masking: full date for clinicians, year only for researchers
CREATE OR REPLACE MASKING POLICY phi_dob_mask AS
(val DATE) RETURNS DATE ->
CASE
WHEN IS_ROLE_IN_SESSION('TREATING_CLINICIAN') THEN val
WHEN IS_ROLE_IN_SESSION('CARE_TEAM_MEMBER') THEN val
WHEN IS_ROLE_IN_SESSION('RESEARCHER') THEN
DATE_FROM_PARTS(YEAR(val), 1, 1) -- year only
ELSE NULL
END;
-- Apply masking policies to patient table
ALTER TABLE clinical.patient_demographics_table
MODIFY COLUMN last_name
SET MASKING POLICY phi_name_mask;
ALTER TABLE clinical.patient_demographics_table
MODIFY COLUMN first_name
SET MASKING POLICY phi_name_mask;
ALTER TABLE clinical.patient_demographics_table
MODIFY COLUMN date_of_birth
SET MASKING POLICY phi_dob_mask;
-- Now: same query, different results per role
-- Clinician sees: John Smith, 1985-03-22
-- Researcher sees: Patient-a7x9f2e1, 1985-01-01
-- External sees: REDACTED, NULLSecure Data Sharing for Research
Zero-Copy Sharing Setup
Snowflake's Secure Data Sharing allows you to share clinical analytics with research partners, payer organizations, or public health agencies without copying or moving data. The consumer gets a read-only view of live data in your account:
-- Step 1: Create a share
CREATE OR REPLACE SHARE clinical_research_share
COMMENT = 'De-identified clinical data for approved research partners';
-- Step 2: Create a secure view with de-identification
CREATE OR REPLACE SECURE VIEW shared.research_conditions AS
SELECT
MD5(patient_ref) AS patient_pseudonym, -- pseudonymized ID
diagnosis_code,
diagnosis_display,
clinical_status,
YEAR(onset_date) AS onset_year, -- year only, not full date
category
FROM clinical.conditions
WHERE verification_status = 'confirmed'
AND clinical_status IN ('active', 'resolved');
CREATE OR REPLACE SECURE VIEW shared.research_lab_results AS
SELECT
MD5(patient_ref) AS patient_pseudonym,
loinc_code,
test_name,
result_value,
result_unit,
DATE_TRUNC('MONTH', effective_date) AS effective_month,
status
FROM clinical.lab_results
WHERE status = 'final';
-- Step 3: Grant to share
GRANT USAGE ON DATABASE analytics_db TO SHARE clinical_research_share;
GRANT USAGE ON SCHEMA shared TO SHARE clinical_research_share;
GRANT SELECT ON shared.research_conditions TO SHARE clinical_research_share;
GRANT SELECT ON shared.research_lab_results TO SHARE clinical_research_share;
-- Step 4: Add consumer accounts
ALTER SHARE clinical_research_share
ADD ACCOUNTS = 'RESEARCH_PARTNER_ACCOUNT';
-- Consumer side: create database from share
-- CREATE DATABASE research_data FROM SHARE provider_account.clinical_research_share;Secure Sharing vs. Traditional Data Exchange
| Aspect | Traditional (File Export) | Snowflake Secure Sharing |
|---|---|---|
| Data Movement | Copy to SFTP/S3, consumer loads into their system | No movement — consumer queries data in provider account |
| Freshness | Batch exports (daily/weekly), always stale | Real-time — consumer sees latest data immediately |
| Security | Data sits in multiple locations, harder to audit | Single copy, provider controls access, full audit trail |
| Cost | Storage duplication, ETL maintenance | No additional storage cost; consumer pays only for compute |
| Revocation | Cannot recall exported data | Revoke access instantly by removing consumer account |
| HIPAA Compliance | Requires BAA with file transfer platform | Covered under Snowflake BAA; de-identification in secure views |
Streams and Tasks: Near Real-Time Updates
Continuous Ingestion with Snowpipe
-- Snowpipe: Auto-ingest FHIR NDJSON as files land in S3
CREATE OR REPLACE PIPE fhir_patient_pipe
AUTO_INGEST = TRUE
AS
COPY INTO raw_fhir.patients
FROM @fhir_bulk_export_stage/Patient/
FILE_FORMAT = fhir_ndjson_format;
-- Stream: Capture changes to raw table
CREATE OR REPLACE STREAM patient_changes_stream
ON TABLE raw_fhir.patients
APPEND_ONLY = TRUE;
-- Task: Transform new records every 5 minutes
CREATE OR REPLACE TASK transform_patients
WAREHOUSE = 'ETL_WH'
SCHEDULE = '5 MINUTE'
WHEN SYSTEM$STREAM_HAS_DATA('patient_changes_stream')
AS
MERGE INTO clinical.patient_demographics_table tgt
USING (
SELECT
raw_data:id::STRING AS patient_id,
raw_data:name[0].family::STRING AS last_name,
raw_data:name[0].given[0]::STRING AS first_name,
raw_data:birthDate::DATE AS date_of_birth,
raw_data:gender::STRING AS gender,
raw_data:address[0].city::STRING AS city,
raw_data:address[0].state::STRING AS state,
raw_data:address[0].postalCode::STRING AS zip_code,
raw_data:meta.lastUpdated::TIMESTAMP_NTZ AS last_updated
FROM patient_changes_stream
) src
ON tgt.patient_id = src.patient_id
WHEN MATCHED AND src.last_updated > tgt.last_updated THEN
UPDATE SET
tgt.last_name = src.last_name,
tgt.first_name = src.first_name,
tgt.date_of_birth = src.date_of_birth,
tgt.gender = src.gender,
tgt.city = src.city,
tgt.state = src.state,
tgt.zip_code = src.zip_code,
tgt.last_updated = src.last_updated
WHEN NOT MATCHED THEN
INSERT (patient_id, last_name, first_name, date_of_birth,
gender, city, state, zip_code, last_updated)
VALUES (src.patient_id, src.last_name, src.first_name,
src.date_of_birth, src.gender, src.city, src.state,
src.zip_code, src.last_updated);
-- Resume task
ALTER TASK transform_patients RESUME;Snowpark for Clinical ML
Feature Engineering on FHIR Data
# snowpark_clinical_features.py
# Build ML features from FHIR data without moving data out of Snowflake
from snowflake.snowpark import Session
from snowflake.snowpark.functions import (
col, count, avg, max as sf_max, min as sf_min,
datediff, lit, when, sum as sf_sum
)
from snowflake.snowpark.types import FloatType
session = Session.builder.configs({
"account": "healthcare_org",
"user": "ml_engineer",
"password": "...",
"database": "ANALYTICS_DB",
"schema": "ML_FEATURES",
"warehouse": "ML_WH"
}).create()
# Load clinical tables (already flattened from FHIR)
patients = session.table("CLINICAL.PATIENT_DEMOGRAPHICS_TABLE")
conditions = session.table("CLINICAL.CONDITIONS")
lab_results = session.table("CLINICAL.LAB_RESULTS")
encounters = session.table("CLINICAL.ENCOUNTERS")
# Feature 1: Comorbidity count (number of active conditions)
comorbidity_features = conditions \
.filter(col("CLINICAL_STATUS") == "active") \
.group_by("PATIENT_REF") \
.agg(
count("*").alias("active_condition_count"),
count(when(col("CATEGORY") == "encounter-diagnosis", 1))
.alias("encounter_diagnosis_count")
)
# Feature 2: Lab result trends (last 90 days)
lab_features = lab_results \
.filter(col("EFFECTIVE_DATE") >= "2026-01-01") \
.group_by("PATIENT_REF") \
.agg(
count("*").alias("lab_count_90d"),
avg(when(col("LOINC_CODE") == "2339-0", col("RESULT_VALUE")))
.alias("avg_glucose"),
avg(when(col("LOINC_CODE") == "2160-0", col("RESULT_VALUE")))
.alias("avg_creatinine"),
sf_max(when(col("LOINC_CODE") == "4548-4", col("RESULT_VALUE")))
.alias("latest_hba1c")
)
# Feature 3: Utilization patterns
utilization_features = encounters \
.filter(col("ENCOUNTER_START") >= "2025-03-16") \
.group_by("PATIENT_REF") \
.agg(
count("*").alias("encounters_12m"),
count(when(col("ENCOUNTER_TYPE") == "emergency", 1))
.alias("ed_visits_12m"),
count(when(col("ENCOUNTER_TYPE") == "inpatient", 1))
.alias("admissions_12m"),
avg(datediff("day", col("ENCOUNTER_START"), col("ENCOUNTER_END")))
.alias("avg_los_days")
)
# Combine features
ml_features = patients \
.join(comorbidity_features, patients["PATIENT_ID"] == comorbidity_features["PATIENT_REF"], "left") \
.join(lab_features, patients["PATIENT_ID"] == lab_features["PATIENT_REF"], "left") \
.join(utilization_features, patients["PATIENT_ID"] == utilization_features["PATIENT_REF"], "left") \
.select(
col("PATIENT_ID"),
col("GENDER"),
datediff("year", col("DATE_OF_BIRTH"), lit("2026-03-16"))
.alias("AGE"),
col("ACTIVE_CONDITION_COUNT"),
col("LAB_COUNT_90D"),
col("AVG_GLUCOSE"),
col("AVG_CREATININE"),
col("LATEST_HBA1C"),
col("ENCOUNTERS_12M"),
col("ED_VISITS_12M"),
col("ADMISSIONS_12M"),
col("AVG_LOS_DAYS")
)
# Save features table (data never leaves Snowflake)
ml_features.write.mode("overwrite").save_as_table("ML_FEATURES.READMISSION_FEATURES")
print(f"Feature table created: {ml_features.count()} patients")Snowflake vs. Alternatives for Healthcare
| Capability | Snowflake | Databricks | BigQuery | Redshift |
|---|---|---|---|---|
| Semi-structured (FHIR JSON) | Native VARIANT + FLATTEN | Native JSON support | Native JSON support | Limited (SUPER type) |
| Secure Data Sharing | Zero-copy, cross-account | Delta Sharing (open protocol) | Analytics Hub | Data sharing via Redshift |
| Dynamic Masking | Built-in, policy-based | Unity Catalog column masking | Column-level security | Limited |
| HIPAA BAA | Available (Business Critical+) | Available | Available | Available |
| ML/Python | Snowpark (in-warehouse) | Native (MLflow, notebooks) | BigQuery ML | Redshift ML |
| Streaming | Snowpipe + Streams/Tasks | Structured Streaming | Streaming inserts | Streaming ingestion |
| Best For | SQL-heavy analytics, data sharing | ML-heavy, lakehouse architecture | Google Cloud ecosystem | AWS-native workloads |
For healthcare organizations where the primary use case is SQL analytics on clinical data with secure sharing to research partners, Snowflake is the strongest choice. For organizations that need heavy ML workloads with Delta Lake and OMOP CDM, Databricks may be a better fit. Many health systems use both — Snowflake for the consumption/sharing layer and Databricks for the transformation/ML layer.
Cost Optimization for Healthcare Workloads
-- Snowflake cost optimization for healthcare data warehouse
-- 1. Use warehouse auto-suspend and auto-resume
ALTER WAREHOUSE ANALYTICS_WH SET
AUTO_SUSPEND = 120, -- suspend after 2 min idle
AUTO_RESUME = TRUE,
MIN_CLUSTER_COUNT = 1,
MAX_CLUSTER_COUNT = 4, -- scale up for peak hours
SCALING_POLICY = 'ECONOMY';
-- 2. Separate warehouses by workload
CREATE WAREHOUSE ETL_WH WITH
WAREHOUSE_SIZE = 'MEDIUM'
AUTO_SUSPEND = 60; -- ETL jobs: medium, suspend quickly
CREATE WAREHOUSE DASHBOARD_WH WITH
WAREHOUSE_SIZE = 'SMALL'
AUTO_SUSPEND = 300; -- dashboards: small, keep warm
CREATE WAREHOUSE ML_WH WITH
WAREHOUSE_SIZE = 'XLARGE'
AUTO_SUSPEND = 60; -- ML: large but suspend fast
-- 3. Cluster keys for common clinical queries
-- Lab results: most queries filter by patient + date
ALTER TABLE clinical.lab_results_table
CLUSTER BY (patient_ref, effective_date);
-- Conditions: queries filter by diagnosis code
ALTER TABLE clinical.conditions_table
CLUSTER BY (diagnosis_code, onset_date);
-- 4. Materialized views for common aggregations
CREATE OR REPLACE MATERIALIZED VIEW clinical.daily_lab_summary AS
SELECT
DATE_TRUNC('DAY', effective_date) AS result_date,
loinc_code,
test_name,
COUNT(*) AS result_count,
AVG(result_value) AS avg_value,
MIN(result_value) AS min_value,
MAX(result_value) AS max_value
FROM clinical.lab_results
WHERE status = 'final'
GROUP BY 1, 2, 3;Frequently Asked Questions
Is Snowflake HIPAA-compliant?
Yes — Snowflake offers HIPAA compliance on its Business Critical edition and above. This includes a signed Business Associate Agreement (BAA), encryption at rest (AES-256) and in transit (TLS 1.2+), private connectivity options (AWS PrivateLink, Azure Private Link), and HITRUST CSF certification. You must use Business Critical or higher for PHI workloads — the Standard and Enterprise editions do not support BAA.
Should we store raw FHIR JSON or flatten everything?
Both. Store raw FHIR NDJSON in VARIANT columns as your immutable bronze layer. Create flattened views and tables as your silver/gold layers for analytics. This gives you the best of both worlds: the raw JSON preserves all FHIR extensions and metadata that you might need later, while the flattened tables give analysts fast, SQL-friendly access to common fields. The flattening views cost nothing until queried — they are just metadata.
How does Snowflake handle FHIR references and joins?
FHIR resources reference each other via the reference field (e.g., "subject": {"reference": "Patient/12345"}). In Snowflake, extract the reference string, split on '/' to get the resource type and ID, and join across tables. For frequently used joins, create materialized views or pre-join tables in your gold layer. Snowflake's query optimizer handles these joins efficiently at scale.
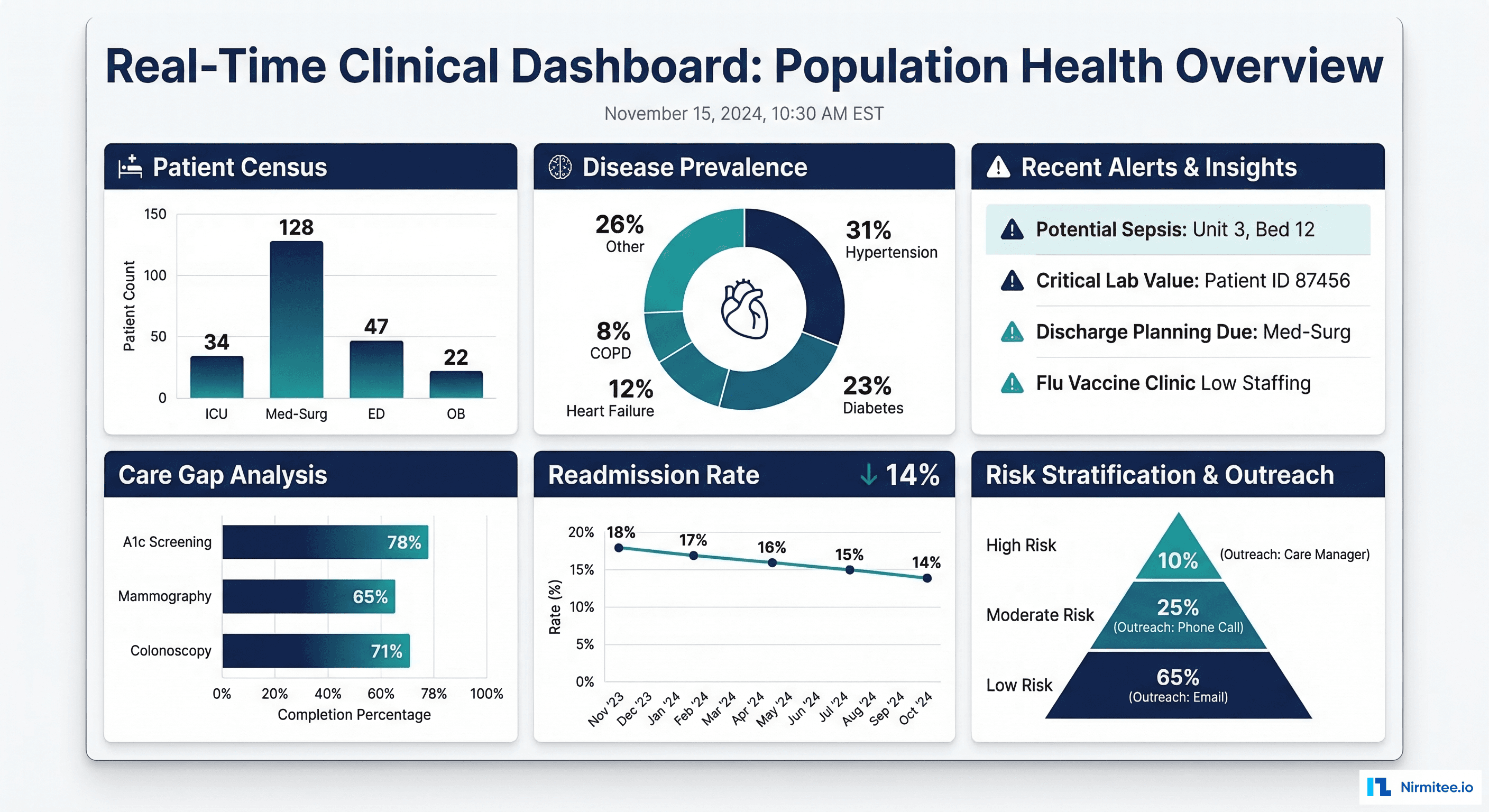
What about real-time clinical dashboards on Snowflake?
Snowflake is not a real-time database — it is designed for analytical workloads with latency measured in seconds, not milliseconds. Using Snowpipe, Streams, and Tasks, you can achieve near-real-time updates with 5-15-minute latency, which is sufficient for most clinical dashboards (patient census, lab result trends, population health metrics). For true sub-second latency (ED tracking boards, monitor alarms), use a dedicated streaming platform and reserve Snowflake for the analytical layer.
How do we handle FHIR extensions in Snowflake?
FHIR extensions (like US Core Race and Ethnicity) are nested arrays in the JSON. Access them with Snowflake's path notation: raw_data:extension[0].extension[0].valueCoding.display. For complex extensions, use LATERAL FLATTEN to unnest the extension array and filter by URL. Since extensions vary by implementation, document your specific extension paths in a metadata table that analysts can reference.