Medical imaging accounts for 90% of all healthcare data by volume. Every year, more than 3.6 billion diagnostic imaging procedures are performed worldwide, generating petabytes of pixel data in DICOM format. Yet when a referring physician wants to see those images alongside a patient's medications, lab results, and clinical notes, they hit a wall. The imaging data lives in a PACS silo. The clinical data lives in the EHR. The two systems speak fundamentally different languages.
This is the last frontier of healthcare interoperability. HL7v2 connected labs. C-CDA connected documents. FHIR R4 is unifying clinical data through RESTful APIs. But medical imaging -- the richest, most data-intensive domain in healthcare -- has largely remained a walled garden. The DICOM standard, which has governed medical imaging since 1993, was designed for a world of dedicated PACS workstations and radiology reading rooms, not for the API-driven, cloud-native, AI-augmented healthcare infrastructure being built today.
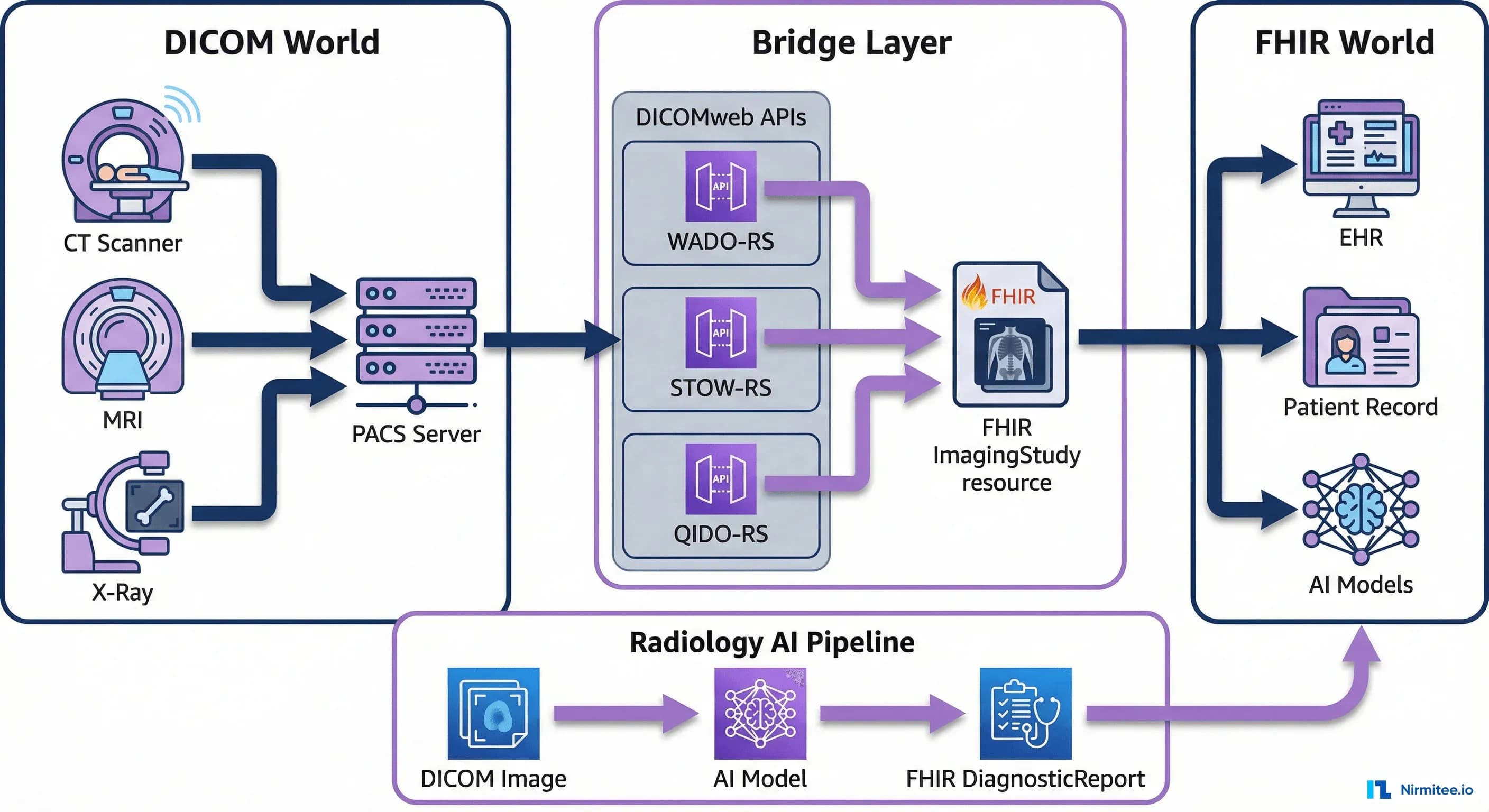
That is changing. The convergence of DICOM and FHIR is rewriting how imaging data flows through health systems, how radiology AI gets deployed, and how enterprise imaging architectures are designed. This guide covers the complete technical landscape: from the ImagingStudy resource that bridges both worlds, to the DICOMweb APIs that make pixels accessible over HTTP, to the cloud PACS architectures that are replacing on-premises infrastructure. Whether you are building an imaging AI pipeline, migrating off a legacy PACS, or designing an interoperable healthcare platform, this is the technical reference you need.
DICOM and FHIR -- Two Standards, One Patient
DICOM (Digital Imaging and Communications in Medicine) and FHIR (Fast Healthcare Interoperability Resources) were designed decades apart for fundamentally different problems. Understanding what each does -- and where they necessarily overlap -- is the foundation for any integration architecture.
What DICOM Does
DICOM is a complete imaging ecosystem standard. It defines:
- File format: The .dcm file encodes pixel data alongside metadata (patient demographics, study information, acquisition parameters, device settings) in a binary header with tag-length-value encoding
- Network protocol: DICOM DIMSE services (C-STORE, C-FIND, C-MOVE, C-GET, C-ECHO) operate over TCP connections between imaging devices, PACS, and workstations
- Information model: A four-level hierarchy -- Patient, Study, Series, Instance -- organizes every imaging exam
- Structured reporting: DICOM SR (Structured Reports) encode findings using coded entries from RadLex, SNOMED CT, and other terminologies
- Worklist management: Modality Worklist (MWL) pushes scheduled procedure information from the RIS to imaging devices
DICOM was built when imaging lived exclusively in radiology departments. A CT scanner sends images to a PACS via C-STORE. A radiologist reviews them on a dedicated workstation. The report goes back to the RIS, which sends it to the EHR as an HL7v2 ORU message. This workflow has served radiology for 30 years, but it was never designed for cross-departmental access, API-driven integrations, or cloud-native architectures.
What FHIR Does
FHIR R4 is a RESTful API standard for clinical data exchange. It defines resources for clinical concepts (Patient, Condition, Observation, MedicationRequest), a JSON/XML serialization format, a REST API surface (CRUD operations, search parameters, batch/transaction bundles), and an authorization framework (SMART on FHIR). For imaging specifically, FHIR provides the ImagingStudy resource to represent imaging exam metadata and the ImagingSelection resource (FHIR R5) for referencing specific frames or regions of interest.
FHIR was not designed to transmit pixel data. A chest CT generating 500 MB of DICOM instances is not going inside a FHIR Bundle. FHIR's role in imaging is metadata and orchestration: indexing what studies exist for a patient, linking them to clinical encounters and diagnostic reports, and providing the URLs where the actual pixel data can be retrieved via DICOMweb.
Where They Overlap -- and Why That Matters
Both standards carry patient demographics, study-level metadata (accession number, study date, referring physician, procedure codes), and references to clinical context. This overlap creates both opportunity and risk. The opportunity is a natural join point: a FHIR ImagingStudy resource can reference the same study that lives in DICOM, creating a unified view. The risk is data inconsistency: if the patient's name is updated in the EHR (FHIR) but not in the PACS (DICOM), downstream systems will disagree.
The IHE (Integrating the Healthcare Enterprise) profiles address this with defined workflows. The IHE RAD profiles (specifically XDS-I.b for cross-enterprise imaging sharing and MHD for mobile access) define how DICOM and FHIR metadata stay synchronized. The IHE Mobile access to Health Documents (MHD) profile maps document sharing to FHIR transactions. In practice, most production systems treat DICOM as the source of truth for imaging metadata and use FHIR as a federated index.
The ImagingStudy Resource -- FHIR's Bridge to DICOM
The ImagingStudy resource is FHIR's primary mechanism for representing imaging exams. It does not contain pixel data. Instead, it holds the metadata needed to locate, identify, and contextualize an imaging study within a patient's clinical record, while providing the endpoint references needed to retrieve the actual images via DICOMweb.
Resource Structure
An ImagingStudy has three levels that mirror the DICOM information model:
- Study level: Patient reference, encounter reference, study instance UID, accession number, modality list, number of series, number of instances, procedure codes, reason codes, and the endpoint where the study can be retrieved
- Series level: Series instance UID, modality (CT, MR, US, etc.), body site, laterality, number of instances, and series-specific endpoints
- Instance level: SOP instance UID, SOP class (specifying the type of DICOM object -- CT Image Storage, MR Image Storage, etc.), and instance number
Here is a complete FHIR R4 ImagingStudy resource for a chest CT scan, with DICOMweb endpoint references:
{
"resourceType": "ImagingStudy",
"id": "chest-ct-example",
"status": "available",
"subject": {
"reference": "Patient/pat-john-smith",
"display": "John Smith"
},
"encounter": {
"reference": "Encounter/enc-20260315-er"
},
"started": "2026-03-15T10:30:00Z",
"basedOn": [
{
"reference": "ServiceRequest/sr-chest-ct-001"
}
],
"referrer": {
"reference": "Practitioner/prac-dr-chen",
"display": "Dr. Wei Chen"
},
"endpoint": [
{
"reference": "Endpoint/wado-rs-server"
}
],
"numberOfSeries": 3,
"numberOfInstances": 487,
"procedureCode": [
{
"coding": [
{
"system": "http://loinc.org",
"code": "36643-5",
"display": "CT Chest WO Contrast"
}
]
}
],
"reasonCode": [
{
"coding": [
{
"system": "http://snomed.info/sct",
"code": "49727002",
"display": "Cough"
}
]
}
],
"description": "CT Chest without contrast",
"series": [
{
"uid": "2.16.124.113543.6003.1154777499.30246.19789.3503430046",
"number": 1,
"modality": {
"system": "http://dicom.nema.org/resources/ontology/DCM",
"code": "CT"
},
"description": "Axial 1.25mm",
"numberOfInstances": 412,
"bodySite": {
"system": "http://snomed.info/sct",
"code": "51185008",
"display": "Thorax"
},
"instance": [
{
"uid": "2.16.124.113543.6003.189642796.63084.16748.2599092903",
"sopClass": {
"system": "urn:ietf:rfc:3986",
"code": "urn:oid:1.2.840.10008.5.1.4.1.1.2"
},
"number": 1
}
]
},
{
"uid": "2.16.124.113543.6003.1154777499.30246.19789.3503430047",
"number": 2,
"modality": {
"system": "http://dicom.nema.org/resources/ontology/DCM",
"code": "CT"
},
"description": "Coronal 3mm MPR",
"numberOfInstances": 45,
"bodySite": {
"system": "http://snomed.info/sct",
"code": "51185008",
"display": "Thorax"
}
},
{
"uid": "2.16.124.113543.6003.1154777499.30246.19789.3503430048",
"number": 3,
"modality": {
"system": "http://dicom.nema.org/resources/ontology/DCM",
"code": "CT"
},
"description": "Sagittal 3mm MPR",
"numberOfInstances": 30,
"bodySite": {
"system": "http://snomed.info/sct",
"code": "51185008",
"display": "Thorax"
}
}
]
}The Endpoint Resource -- Connecting FHIR to DICOMweb
The Endpoint resource referenced in ImagingStudy tells the consuming system where and how to retrieve the actual images. For DICOMweb, it looks like this:
{
"resourceType": "Endpoint",
"id": "wado-rs-server",
"status": "active",
"connectionType": {
"system": "http://terminology.hl7.org/CodeSystem/endpoint-connection-type",
"code": "dicom-wado-rs"
},
"name": "Enterprise PACS DICOMweb",
"payloadType": [
{
"coding": [
{
"system": "http://dicom.nema.org/resources/ontology/DCM",
"code": "113014",
"display": "Study"
}
]
}
],
"address": "https://pacs.hospital.org/dicomweb"
}This design is intentional. FHIR provides the clinical index -- "Patient John Smith had a chest CT on March 15 for cough, with 487 instances across 3 series." DICOMweb provides the pixel pipeline -- "retrieve the actual DICOM instances from this URL." Neither standard tries to do the other's job.
DICOMweb APIs Explained
DICOMweb is the RESTful evolution of DICOM's legacy network services. Instead of DIMSE commands over TCP connections requiring AE title configuration and dedicated ports, DICOMweb provides three HTTP-based services that work with standard web infrastructure: load balancers, CDNs, API gateways, OAuth tokens, and CORS policies. Published as DICOM PS3.18, DICOMweb has become the standard API layer for modern PACS, VNA, and cloud imaging services.
WADO-RS: Web Access to DICOM Objects (Retrieve)
WADO-RS retrieves DICOM objects -- studies, series, or individual instances -- over HTTP. It supports content negotiation via the Accept header, so consumers can request DICOM binary, bulk data, or rendered images (JPEG, PNG) from the same endpoint.
Retrieve an entire study (all series, all instances):
GET /dicomweb/studies/2.16.124.113543.6003.1154777499.30246.19789.3503430045
Accept: multipart/related; type="application/dicom"
Authorization: Bearer eyJhbGciOiJSUzI1NiIs...Retrieve a single instance as rendered JPEG (for viewer thumbnails):
GET /dicomweb/studies/2.16.124.113543.6003.1154777499.30246.19789.3503430045/series/2.16.124.113543.6003.1154777499.30246.19789.3503430046/instances/2.16.124.113543.6003.189642796.63084.16748.2599092903/rendered
Accept: image/jpeg
Authorization: Bearer eyJhbGciOiJSUzI1NiIs...Retrieve instance metadata only (no pixel data):
GET /dicomweb/studies/2.16.124.113543.6003.1154777499.30246.19789.3503430045/metadata
Accept: application/dicom+json
Authorization: Bearer eyJhbGciOiJSUzI1NiIs...STOW-RS: Store Over the Web (Upload)
STOW-RS enables HTTP-based DICOM storage. Modalities and gateways POST DICOM instances to a DICOMweb server instead of using the legacy C-STORE DIMSE service. This is critical for cloud architectures where imaging devices need to send data to a cloud PACS without VPN tunnels to legacy DIMSE listeners.
POST /dicomweb/studies
Content-Type: multipart/related; type="application/dicom"; boundary=boundary123
Authorization: Bearer eyJhbGciOiJSUzI1NiIs...
--boundary123
Content-Type: application/dicom
[DICOM binary instance 1]
--boundary123
Content-Type: application/dicom
[DICOM binary instance 2]
--boundary123--The server returns a DICOM JSON response with the stored instance UIDs and any warnings or failures:
{
"00081190": {
"vr": "UR",
"Value": [
"https://pacs.hospital.org/dicomweb/studies/2.16.124.113543.6003.1154777499.30246.19789.3503430045"
]
},
"00081199": {
"vr": "SQ",
"Value": [
{
"00081150": {
"vr": "UI",
"Value": ["1.2.840.10008.5.1.4.1.1.2"]
},
"00081155": {
"vr": "UI",
"Value": ["2.16.124.113543.6003.189642796.63084.16748.2599092903"]
}
}
]
}
}QIDO-RS: Query Based on ID for DICOM Objects (Search)
QIDO-RS is the search API. It replaces C-FIND with HTTP GET requests that support query parameters for filtering and pagination. QIDO-RS returns DICOM JSON metadata -- not pixel data -- making it lightweight enough for study lists, worklists, and search interfaces.
Search for all CT studies for a patient in the last 30 days:
GET /dicomweb/studies?PatientID=PAT-12345&ModalitiesInStudy=CT&StudyDate=20260301-20260401&limit=25&offset=0
Accept: application/dicom+json
Authorization: Bearer eyJhbGciOiJSUzI1NiIs...Search for series within a specific study:
GET /dicomweb/studies/2.16.124.113543.6003.1154777499.30246.19789.3503430045/series?Modality=CT
Accept: application/dicom+jsonSearch with includefield for selective metadata retrieval:
GET /dicomweb/studies?PatientID=PAT-12345&includefield=00081030,00080060,00201206
Accept: application/dicom+jsonThe includefield parameter controls which DICOM tags are returned, reducing payload size for study list UIs that only need study description, modality, and number of instances.
Architecture Patterns for DICOM-FHIR Integration
There is no single right way to integrate DICOM and FHIR. The optimal architecture depends on your existing infrastructure, cloud strategy, and the use cases you need to support. Three dominant patterns have emerged in production deployments.
Pattern 1: FHIR as Index, DICOMweb for Pixels
This is the most common pattern and the one recommended by IHE profiles. The FHIR server acts as a clinical index that knows what imaging studies exist for each patient. The PACS or VNA serves the actual pixel data via DICOMweb. Consuming applications query FHIR for study metadata, then follow the endpoint URLs to DICOMweb for image retrieval.
Data flow:
- Modality completes a scan and sends DICOM instances to the PACS via STOW-RS or C-STORE
- A DICOM-to-FHIR bridge service (or PACS notification handler) creates or updates an
ImagingStudyresource on the FHIR server with the study metadata and DICOMweb endpoint reference - When a clinician opens the patient chart in the EHR, the EHR queries the FHIR server:
GET /ImagingStudy?patient=Patient/pat-john-smith&_sort=-date - The EHR displays the study list. When the clinician clicks to view images, the EHR launches an embedded viewer (OHIF, Cornerstone) pointing at the DICOMweb endpoint from the ImagingStudy's Endpoint reference
Advantages: Minimal disruption to existing PACS infrastructure. The PACS continues to be the system of record for imaging. FHIR adds a clinical access layer without duplicating pixel data. This pattern works well with any FHIR server -- HAPI, Google Cloud Healthcare API, AWS HealthLake.
Disadvantages: Requires a synchronization mechanism between PACS and FHIR. If a study is deleted or corrected in the PACS, the FHIR ImagingStudy must be updated. Network topology must allow consuming applications to reach both the FHIR server and the DICOMweb endpoint.
Pattern 2: Full FHIR with Embedded ImagingStudy References
In this pattern, the FHIR server is the primary clinical data platform. ImagingStudy resources are first-class citizens alongside Conditions, Observations, and DiagnosticReports. The FHIR server may proxy or cache imaging metadata, and DiagnosticReport resources for radiology directly reference the ImagingStudy:
{
"resourceType": "DiagnosticReport",
"id": "rad-report-chest-ct-001",
"status": "final",
"category": [
{
"coding": [
{
"system": "http://terminology.hl7.org/CodeSystem/v2-0074",
"code": "RAD",
"display": "Radiology"
}
]
}
],
"code": {
"coding": [
{
"system": "http://loinc.org",
"code": "36643-5",
"display": "CT Chest WO Contrast"
}
]
},
"subject": {
"reference": "Patient/pat-john-smith"
},
"effectiveDateTime": "2026-03-15T11:45:00Z",
"issued": "2026-03-15T14:30:00Z",
"performer": [
{
"reference": "Practitioner/prac-dr-radiologist",
"display": "Dr. Sarah Martinez, MD"
}
],
"imagingStudy": [
{
"reference": "ImagingStudy/chest-ct-example"
}
],
"conclusion": "No acute cardiopulmonary process. Stable 4mm right upper lobe nodule, recommend follow-up CT in 12 months per Fleischner criteria.",
"conclusionCode": [
{
"coding": [
{
"system": "http://snomed.info/sct",
"code": "168731009",
"display": "Chest CT normal"
}
]
}
]
}Advantages: Single API surface for all clinical data. Applications query one FHIR server and get the complete picture -- imaging, labs, medications, notes. Simplifies authorization (one SMART on FHIR token covers everything). This pattern is favored by organizations building new FHIR-native platforms rather than retrofitting existing systems.
Disadvantages: Higher complexity in the FHIR server layer. Requires robust synchronization and careful handling of large imaging metadata sets.
Pattern 3: Cloud-Native Medical Imaging
Cloud providers now offer managed services that handle both DICOM storage and FHIR indexing natively, eliminating the need for a separate synchronization layer.
- Google Cloud Healthcare API: A single API that supports both FHIR R4 and DICOMweb natively. When a DICOM instance is stored via STOW-RS, the service can automatically create or update a corresponding FHIR ImagingStudy resource. Both are accessible through the same project and authentication mechanism.
- AWS HealthImaging: Purpose-built for medical imaging at scale. Stores imaging data in a cloud-optimized format with sub-second retrieval. Integrates with AWS HealthLake (FHIR) for clinical data correlation.
- Azure Health Data Services: DICOM service alongside a FHIR service in the same workspace, with built-in synchronization pipelines.
Advantages: No infrastructure to manage. Built-in synchronization. Elastic scaling for imaging workloads that can spike dramatically (a single MRI study can be 2-3 GB). HIPAA BAA coverage included.
Disadvantages: Cloud vendor lock-in for the imaging layer. Egress costs for large imaging datasets can be significant (retrieving 1 TB of imaging data from cloud storage incurs real networking costs). Latency requirements for real-time radiology reading may not be met for all geographic regions.
Radiology AI Pipeline -- DICOM to FHIR DiagnosticReport
The most transformative application of DICOM-FHIR integration is the radiology AI pipeline. AI models that detect pulmonary nodules, classify chest pathologies, or measure cardiac function all consume DICOM pixel data and produce clinical findings. The challenge is getting those findings back into the clinical workflow in a structured, actionable format. This is where FHIR closes the loop.
The End-to-End Pipeline
A production radiology AI pipeline has five stages:
- Trigger: A new imaging study arrives in the PACS. A DICOM router (or DICOMweb subscription) detects the study and checks routing rules (e.g., all chest CTs go to the lung nodule AI).
- Retrieve: The AI orchestrator retrieves the DICOM instances via WADO-RS:
GET /dicomweb/studies/{studyUID}. For efficiency, it may retrieve only the axial series or a specific reconstruction kernel. - Inference: The DICOM instances are preprocessed (windowing, resampling to isotropic voxels, normalization) and fed to the AI model. The model outputs structured findings: nodule locations (x, y, z coordinates), sizes, Lung-RADS classifications, confidence scores.
- Store results: The AI findings are encoded as a DICOM Structured Report (SR) or DICOM Secondary Capture with annotations, stored back to the PACS via STOW-RS. Simultaneously, a FHIR
DiagnosticReportis created with coded findings. - Alert: If the findings meet alert criteria (e.g., Lung-RADS 4B nodule), a CDS Hooks card is fired to the ordering clinician's EHR session, or a FHIR
Communicationresource triggers a notification workflow.
FHIR DiagnosticReport with AI Findings
Here is what the AI's output looks like as a FHIR DiagnosticReport with structured observations:
{
"resourceType": "DiagnosticReport",
"id": "ai-lung-nodule-report-001",
"status": "preliminary",
"category": [
{
"coding": [
{
"system": "http://terminology.hl7.org/CodeSystem/v2-0074",
"code": "RAD"
}
]
}
],
"code": {
"coding": [
{
"system": "http://loinc.org",
"code": "82688-3",
"display": "Lung nodule assessment"
}
]
},
"subject": {
"reference": "Patient/pat-john-smith"
},
"effectiveDateTime": "2026-03-15T10:35:00Z",
"performer": [
{
"reference": "Device/ai-lung-cad-v3",
"display": "LungCAD AI v3.2.1"
}
],
"imagingStudy": [
{
"reference": "ImagingStudy/chest-ct-example"
}
],
"result": [
{
"reference": "Observation/ai-nodule-finding-001",
"display": "RUL nodule, 4mm, Lung-RADS 2"
}
],
"conclusion": "AI-detected: 1 pulmonary nodule in right upper lobe (4mm solid, Lung-RADS 2). Recommend follow-up CT in 12 months per Fleischner criteria. Confidence: 0.94.",
"extension": [
{
"url": "http://example.org/fhir/StructureDefinition/ai-model-metadata",
"extension": [
{
"url": "model-name",
"valueString": "LungCAD"
},
{
"url": "model-version",
"valueString": "3.2.1"
},
{
"url": "confidence-score",
"valueDecimal": 0.94
},
{
"url": "fda-clearance",
"valueString": "K203456"
}
]
}
]
}CDS Hooks Integration for Real-Time AI Alerts
CDS Hooks provides a standardized mechanism for AI systems to push alerts into the clinician's workflow at the point of care. When integrated with a radiology AI pipeline, the flow works as follows:
The EHR fires a order-select or custom imaging-study-complete hook. The CDS service checks if AI findings exist for the study. If critical findings are present, it returns a card:
{
"cards": [
{
"uuid": "card-lung-nodule-alert-001",
"summary": "AI Alert: Lung nodule detected on chest CT",
"detail": "LungCAD AI detected a 4mm solid nodule in the right upper lobe (Lung-RADS 2, confidence 0.94). Fleischner criteria recommend follow-up CT in 12 months.",
"indicator": "warning",
"source": {
"label": "LungCAD AI v3.2.1",
"url": "https://lungcad.example.com",
"icon": "https://lungcad.example.com/icon.png"
},
"suggestions": [
{
"label": "Order follow-up CT Chest in 12 months",
"uuid": "suggestion-followup-ct",
"actions": [
{
"type": "create",
"description": "Create follow-up CT order",
"resource": {
"resourceType": "ServiceRequest",
"status": "draft",
"intent": "proposal",
"code": {
"coding": [
{
"system": "http://loinc.org",
"code": "36643-5",
"display": "CT Chest WO Contrast"
}
]
},
"occurrenceDateTime": "2027-03-15"
}
}
]
}
],
"links": [
{
"label": "View AI findings in OHIF Viewer",
"url": "https://viewer.hospital.org/viewer/2.16.124.113543.6003.1154777499.30246.19789.3503430045",
"type": "absolute"
}
]
}
]
}This integration pattern connects directly to clinical decision support systems and the broader healthcare AI agent pipeline. The AI model processes unstructured pixel data (DICOM), produces structured clinical findings (FHIR), and delivers actionable recommendations at the point of care (CDS Hooks).
Enterprise Imaging Architecture
Enterprise imaging extends radiology's imaging infrastructure to every department that captures visual clinical data: cardiology (echocardiograms, cath lab), pathology (whole slide imaging), dermatology (clinical photographs), ophthalmology (OCT, fundoscopy), wound care, and point-of-care ultrasound. The goal is a single imaging platform that serves all departments, replacing the departmental silos that historically managed their own images independently.
Vendor Neutral Archive (VNA)
The VNA is the storage backbone of enterprise imaging. Unlike a department-specific PACS that is tightly coupled to its viewer and workflow engine, a VNA stores imaging data in a vendor-neutral format and serves it via DICOMweb (and increasingly, FHIR) to any consuming application.
Key VNA capabilities for DICOM-FHIR integration:
- Multi-format ingestion: DICOM from radiology and cardiology, JPEG/PNG from dermatology and wound care, TIFF from pathology whole slide imaging, video from endoscopy and surgical capture
- DICOMweb API layer: All content accessible via WADO-RS/QIDO-RS/STOW-RS regardless of source format. Non-DICOM content is wrapped in DICOM Secondary Capture or Visible Light Photographic Image objects
- FHIR resource generation: Automatic creation of ImagingStudy resources when new content arrives, pushed to the enterprise FHIR server for clinical indexing
- Lifecycle management: Retention policies, legal hold, migration between storage tiers (hot/warm/cold) based on study age and access patterns
Universal Viewer Strategy
Enterprise imaging requires a viewer that works across departments. The OHIF Viewer (Open Health Imaging Foundation) has become the de facto open-source standard for web-based DICOM viewing. It connects to any DICOMweb-compatible backend, supports measurement tools, MPR reconstruction, and AI overlay display, and can be embedded in EHR portals via SMART on FHIR launch.
A typical universal viewer deployment serves multiple constituencies from a single platform:
- Radiologists: Full diagnostic reading with hanging protocols, prior comparison, voice dictation integration
- Cardiologists: Echo measurements, ejection fraction calculation, strain analysis
- Surgeons: 3D volume rendering, surgical planning overlays
- Referring physicians: Simplified view with key images, report overlay, and AI findings highlights
- Patients: Consumer-grade viewer via patient portal with SMART on FHIR patient-facing launch
PACS Migration Path
Most health systems are not building enterprise imaging from scratch. They are migrating from legacy departmental PACS to a consolidated architecture. The migration typically follows this sequence:
- Deploy VNA: Stand up the vendor-neutral archive alongside existing PACS. Configure DICOMweb endpoints.
- Route new studies: Configure modality routing to send copies to both the existing PACS and the VNA. This parallel-run phase validates data integrity.
- Historical migration: Migrate historical studies from the legacy PACS to the VNA. This is the longest phase -- large health systems have petabytes of imaging history. Typical migration rates: 50,000-100,000 studies per day with dedicated migration tools.
- Deploy universal viewer: Roll out OHIF or commercial viewer connected to the VNA's DICOMweb API. Validate reading workflows with radiologists.
- FHIR integration: Connect the VNA to the enterprise FHIR server. ImagingStudy resources are created for new and migrated studies. EHR embedding goes live.
- Decommission legacy PACS: Once all historical data is migrated and validated, the legacy PACS is retired.
Cloud PACS: AWS, Azure, and GCP Medical Imaging Services
The three major cloud providers have each built purpose-specific medical imaging services. For organizations evaluating cloud PACS architectures, the differences are significant in pricing model, DICOMweb compliance, FHIR integration depth, and AI/ML ecosystem connectivity.
| Capability | AWS HealthImaging | Azure DICOM Service | Google Cloud Healthcare API |
|---|---|---|---|
| DICOMweb compliance | WADO-RS, STOW-RS, QIDO-RS (full) | WADO-RS, STOW-RS, QIDO-RS (full) | WADO-RS, STOW-RS, QIDO-RS (full) |
| FHIR integration | Separate service (HealthLake FHIR R4). Manual or Lambda-based sync | Same workspace (Azure Health Data Services). Built-in DICOM-to-FHIR sync pipeline | Same API (Healthcare API). Native FHIR R4 + DICOMweb in single project. Auto ImagingStudy creation |
| Storage format | Cloud-optimized HTJ2K (High Throughput JPEG 2000). Stores decoded pixel data for sub-second retrieval | Standard DICOM storage with configurable transcoding | Standard DICOM storage with on-the-fly transcoding |
| AI/ML integration | SageMaker pipelines, Lambda triggers on new studies | Azure ML, Azure AI Services, MONAI integration | Vertex AI, AutoML Vision, Cloud AI Platform notebooks |
| Pricing model | Per-study ingestion + storage per GB/month + retrieval per frame | Per-transaction + storage per GB/month | Per-operation + storage per GB/month |
| De-identification | Built-in configurable de-id at copy time | Built-in with DICOM tag-level control | Built-in with tag-level control + keep-list/remove-list profiles |
| Viewer integration | OHIF via DICOMweb, Amazon HealthScribe for reports | OHIF via DICOMweb, Medical Imaging Server for DICOM (OSS) | OHIF via DICOMweb, Cloud Healthcare DICOM viewer (preview) |
| Certification | HIPAA, HITRUST, SOC 2 | HIPAA, HITRUST, SOC 2 | HIPAA, HITRUST, SOC 2 |
Key Differentiators
Google Cloud Healthcare API has the tightest DICOM-FHIR integration. Both protocols share the same API surface, authentication, and IAM policies. When a DICOM study is stored, a FHIR ImagingStudy resource can be automatically generated -- no custom integration code required. For organizations that want DICOM and FHIR as a single managed service, Google leads.
AWS HealthImaging differentiates on retrieval performance. Its HTJ2K cloud-optimized format enables sub-second first-image display for studies of any size, which matters for teleradiology workflows where latency kills productivity. The tradeoff is that HealthLake FHIR and HealthImaging are separate services requiring custom synchronization via Lambda or EventBridge.
Azure Health Data Services offers the most cohesive healthcare platform play. DICOM, FHIR, and MedTech (device data ingestion) services live in a single workspace with built-in data pipelines. The DICOM-to-FHIR sync pipeline handles ImagingStudy creation automatically. For Microsoft-stack health systems already on Azure AD and Teams, the integration story is strongest.
All three support OHIF Viewer connectivity via DICOMweb, and all three are suitable backends for FHIR-first AI infrastructure.
Implementation Guide -- From Zero to Integrated Imaging
This section walks through a concrete implementation: deploying the OHIF Viewer connected to an Orthanc DICOMweb server, creating FHIR ImagingStudy resources, and linking everything to patient records. This is a production-viable architecture suitable for development environments and smaller deployments, and the same patterns apply at scale with commercial PACS or cloud services.
Step 1: Deploy Orthanc as a DICOMweb Server
Orthanc is an open-source, lightweight DICOM server with a DICOMweb plugin. Deploy it with Docker:
# docker-compose.yml
version: "3.8"
services:
orthanc:
image: orthancteam/orthanc:24.3.3
ports:
- "8042:8042" # Orthanc REST API + DICOMweb
- "4242:4242" # DICOM DIMSE port
environment:
ORTHANC__DICOM_WEB__ENABLE: "true"
ORTHANC__DICOM_WEB__ROOT: "/dicomweb/"
ORTHANC__DICOM_WEB__STUDIES_METADATA: "MainDicomTags"
ORTHANC__REGISTERED_USERS: '{"admin": "orthanc-secret"}'
ORTHANC__AUTHENTICATION_ENABLED: "true"
volumes:
- orthanc-data:/var/lib/orthanc/db
volumes:
orthanc-data:# Start Orthanc
docker compose up -d
# Verify DICOMweb is active
curl -u admin:orthanc-secret http://localhost:8042/dicom-web/studiesStep 2: Upload Sample DICOM Data
Use STOW-RS to upload DICOM instances to Orthanc:
# Upload a DICOM file via STOW-RS
curl -u admin:orthanc-secret \
-X POST \
-H "Content-Type: application/dicom" \
--data-binary @chest_ct_001.dcm \
http://localhost:8042/dicom-web/studies
# Verify the study was stored
curl -u admin:orthanc-secret \
-H "Accept: application/dicom+json" \
http://localhost:8042/dicom-web/studies | python3 -m json.toolStep 3: Deploy OHIF Viewer with DICOMweb Configuration
The OHIF Viewer connects to any DICOMweb-compliant server. Configure it to point at Orthanc:
// ohif-config.js — OHIF v3 configuration
window.config = {
routerBasename: '/',
showStudyList: true,
dataSources: [
{
namespace: '@ohif/extension-default.dataSourcesModule.dicomweb',
sourceName: 'orthanc',
configuration: {
friendlyName: 'Orthanc DICOMweb',
name: 'orthanc',
wadoUriRoot: 'http://localhost:8042/wado',
qidoRoot: 'http://localhost:8042/dicom-web',
wadoRoot: 'http://localhost:8042/dicom-web',
qidoSupportsIncludeField: false,
imageRendering: 'wadors',
thumbnailRendering: 'wadors',
enableStudyLazyLoad: true,
supportsFuzzyMatching: false,
supportsWildcard: true,
bulkDataURI: {
enabled: true
}
}
}
],
defaultDataSourceName: 'orthanc'
};# Deploy OHIF via Docker
docker run -d \
--name ohif-viewer \
-p 3000:80 \
-v $(pwd)/ohif-config.js:/usr/share/nginx/html/app-config.js \
ohif/app:v3.8Access the viewer at http://localhost:3000. It will query Orthanc's QIDO-RS endpoint for the study list and use WADO-RS for image retrieval.
Step 4: Create FHIR ImagingStudy Resources
Write a bridge script that listens for new studies in Orthanc (via its Change API or Lua callbacks) and creates corresponding FHIR ImagingStudy resources. Here is a Python implementation:
import requests
import json
ORTHANC_URL = "http://localhost:8042"
ORTHANC_AUTH = ("admin", "orthanc-secret")
FHIR_URL = "http://localhost:8080/fhir" # HAPI FHIR server
def create_imaging_study(orthanc_study_id: str) -> dict:
"""Fetch DICOM metadata from Orthanc and create a FHIR ImagingStudy."""
# Get study metadata from Orthanc REST API
study = requests.get(
f"{ORTHANC_URL}/studies/{orthanc_study_id}",
auth=ORTHANC_AUTH
).json()
main_tags = study["MainDicomTags"]
study_uid = main_tags["StudyInstanceUID"]
patient_id = study["PatientMainDicomTags"]["PatientID"]
# Build series array
series_list = []
for series_id in study["Series"]:
series_meta = requests.get(
f"{ORTHANC_URL}/series/{series_id}",
auth=ORTHANC_AUTH
).json()
s_tags = series_meta["MainDicomTags"]
series_list.append({
"uid": s_tags["SeriesInstanceUID"],
"number": int(s_tags.get("SeriesNumber", 0)),
"modality": {
"system": "http://dicom.nema.org/resources/ontology/DCM",
"code": s_tags.get("Modality", "OT")
},
"description": s_tags.get("SeriesDescription", ""),
"numberOfInstances": len(series_meta["Instances"])
})
# Create FHIR ImagingStudy
imaging_study = {
"resourceType": "ImagingStudy",
"status": "available",
"subject": {
"reference": f"Patient/{patient_id}"
},
"started": main_tags.get("StudyDate", ""),
"numberOfSeries": len(study["Series"]),
"numberOfInstances": sum(s["numberOfInstances"] for s in series_list),
"description": main_tags.get("StudyDescription", ""),
"endpoint": [
{"reference": "Endpoint/orthanc-dicomweb"}
],
"identifier": [
{
"system": "urn:dicom:uid",
"value": f"urn:oid:{study_uid}"
}
],
"series": series_list
}
# POST to FHIR server
response = requests.post(
f"{FHIR_URL}/ImagingStudy",
json=imaging_study,
headers={"Content-Type": "application/fhir+json"}
)
response.raise_for_status()
return response.json()
# Example: sync all Orthanc studies to FHIR
studies = requests.get(f"{ORTHANC_URL}/studies", auth=ORTHANC_AUTH).json()
for study_id in studies:
result = create_imaging_study(study_id)
print(f"Created ImagingStudy: {result['id']}")Step 5: Link to Patient Records via FHIR
With ImagingStudy resources on the FHIR server, you can now query imaging in the context of a patient's complete clinical record:
# Get all imaging studies for a patient
curl -H "Accept: application/fhir+json" \
"http://localhost:8080/fhir/ImagingStudy?patient=Patient/pat-john-smith&_sort=-date&_count=10"
# Get imaging studies with their associated diagnostic reports
curl -H "Accept: application/fhir+json" \
"http://localhost:8080/fhir/ImagingStudy?patient=Patient/pat-john-smith&_revinclude=DiagnosticReport:imaging-study"
# Search for specific modality studies
curl -H "Accept: application/fhir+json" \
"http://localhost:8080/fhir/ImagingStudy?patient=Patient/pat-john-smith&modality=CT,MR"Security and Compliance Considerations
Medical imaging introduces unique security challenges beyond standard FHIR data protection. The volume of data (a single study can be gigabytes), the sensitivity of the content (images can identify patients even without metadata), and the cross-system access patterns require specific architectural safeguards.
Authentication and Authorization
DICOMweb endpoints must be protected with the same rigor as FHIR endpoints. The recommended approach is SMART on FHIR for DICOMweb: use the same OAuth 2.0 tokens that authorize FHIR API access to also authorize DICOMweb requests. The viewer obtains a SMART token during launch and includes it as a Bearer token in all WADO-RS/QIDO-RS requests. This provides:
- Patient-scoped access: The token's
patientscope ensures the viewer can only retrieve imaging for the authorized patient - Single sign-on: Clinicians authenticate once in the EHR; the viewer inherits the session
- Audit trail: Every image retrieval is logged with the authenticated user identity
De-identification for Research and AI
DICOM files contain PHI in both metadata tags and burned-in pixel annotations. A compliant de-identification pipeline must handle both:
- Tag-level de-identification: Remove or modify DICOM tags containing PHI (Patient Name, Patient ID, Study Date, Referring Physician, etc.) per DICOM PS3.15 Appendix E profiles (Basic, Retain Dates, Retain Device Identity)
- Pixel-level de-identification: Detect and redact burned-in annotations on ultrasound images, screen captures, and secondary captures that may contain patient names, MRN, or dates. Tools like RSNA CTP (Clinical Trial Processor) and cloud-native de-id services handle this
- FHIR resource de-identification: Strip patient references, dates, and identifiers from ImagingStudy and DiagnosticReport resources when exporting for research
Frequently Asked Questions
Can FHIR replace DICOM for medical imaging?
No. FHIR is not designed to store or transmit medical image pixel data. FHIR provides the clinical metadata layer -- indexing what imaging studies exist, linking them to patient records and clinical encounters, and referencing the endpoints where images can be retrieved. DICOM remains the standard for image acquisition, storage, and pixel-level operations. The two standards are complementary: DICOM handles the images, FHIR handles the clinical context, and DICOMweb provides the HTTP bridge between them.
What is the difference between DICOMweb and legacy DICOM networking?
Legacy DICOM networking (DIMSE) uses persistent TCP connections with application-level commands (C-STORE, C-FIND, C-MOVE). It requires configuring AE (Application Entity) titles on both ends, opening specific ports, and often VPN tunnels for remote access. DICOMweb replaces this with standard HTTP REST calls: STOW-RS (POST), QIDO-RS (GET with query parameters), and WADO-RS (GET for retrieval). DICOMweb works with standard web infrastructure -- load balancers, API gateways, OAuth tokens, CDNs -- making it dramatically easier to integrate imaging into web applications and cloud architectures.
How do I handle large imaging studies in a FHIR-based system?
The FHIR ImagingStudy resource stores only metadata, not pixel data, so resource size is not a concern. For the actual image retrieval via DICOMweb, use progressive loading: retrieve the series list first (QIDO-RS), then load instances on demand as the user navigates through the study. Modern viewers like OHIF implement streaming protocols (HTJ2K progressive decoding) that display images before the full dataset is downloaded. For AI pipelines processing large studies, retrieve only the relevant series rather than the entire study.
Which cloud provider should I choose for medical imaging?
Choose Google Cloud Healthcare API if you want the tightest native DICOM-FHIR integration with automatic ImagingStudy resource creation. Choose AWS HealthImaging if retrieval performance is your top priority -- its HTJ2K format enables sub-second first-image display for any study size. Choose Azure Health Data Services if you are a Microsoft-stack organization that wants DICOM, FHIR, and device data in a single managed workspace with built-in sync pipelines. All three are HIPAA-compliant and support DICOMweb and OHIF Viewer connectivity.
How do I integrate radiology AI with existing PACS?
Use a DICOM router (such as Orthanc with Lua scripting, or commercial solutions like Nuance AI Marketplace or Blackford) that intercepts new studies and routes copies to your AI inference service. The AI service retrieves images via DICOMweb (WADO-RS), runs inference, and stores results back as DICOM Structured Reports (STOW-RS) and FHIR DiagnosticReports. CDS Hooks can deliver real-time alerts to the ordering clinician's EHR session. This approach works without modifying the existing PACS -- the router sits between the modalities and the PACS as a transparent intermediary.
What is ImagingSelection in FHIR R5, and should I wait for it?
ImagingSelection is a new FHIR R5 resource that references specific frames, regions of interest, or segments within an imaging study -- not just the study-level metadata that ImagingStudy provides. It is useful for AI annotations (referencing the exact frame where a nodule was detected), surgical planning (referencing a 3D region), and clinical documentation (linking a report finding to a specific image). If you are building today, use FHIR R4 ImagingStudy with extension elements for frame-level references. If your timeline extends to 2027 and beyond, evaluate FHIR R5 adoption for the ImagingSelection capability.
The Road Ahead: DICOM-FHIR Convergence
The convergence of DICOM and FHIR is not a theoretical exercise. It is happening now, driven by three forces: the ONC mandate for API-based interoperability that now touches imaging, the explosion of radiology AI that needs structured output channels, and the migration to cloud infrastructure that demands HTTP-based protocols.
The IHE has published IHE RAD IMR (Interactive Multimedia Report), a profile that uses FHIR DiagnosticReport with embedded image references, enabling radiology reports that link directly to the key images and measurements that support each finding. DICOM Working Group 20 continues to refine the DICOMweb specification, with recent additions for bulk data retrieval and subscription-based notification of new studies. The HL7 Imaging Integration work group is defining how ImagingStudy and ImagingSelection should evolve in FHIR R6.
For healthcare developers building today, the playbook is clear:
- Use DICOMweb as your imaging API layer. Whether your backend is Orthanc, a commercial PACS, or a cloud service, expose images via WADO-RS/QIDO-RS/STOW-RS.
- Use FHIR ImagingStudy as your clinical index. Link imaging studies to patients, encounters, and diagnostic reports through standard FHIR references.
- Deploy a DICOMweb-native viewer. OHIF is production-ready, open source, and connects to any DICOMweb backend.
- Build AI pipelines on DICOM in, FHIR out. Consume DICOM pixel data, produce FHIR DiagnosticReports with coded findings and CDS Hooks alerts.
- Secure everything with SMART on FHIR. Use the same OAuth 2.0 infrastructure for FHIR API and DICOMweb access.
Medical imaging has been the last interoperability frontier for a reason -- the data is massive, the workflows are specialized, and the standards predate the web. But the tools to bridge DICOM and FHIR are now mature, the cloud services are production-ready, and the regulatory pressure is real. The organizations that integrate imaging into their interoperability strategy today will have a decisive advantage in deploying radiology AI, enabling enterprise imaging, and delivering the unified patient record that clinicians have needed for decades.
Building a DICOM-FHIR integration or deploying radiology AI? Nirmitee's healthcare interoperability team has deep expertise in DICOMweb, FHIR R4, and enterprise imaging architecture. Let's talk about your imaging modernization roadmap.