Up to 80% of healthcare data is unstructured. Clinical notes, scanned PDFs, faxed documents, discharge summaries, pathology reports, referral letters — all containing critical clinical information locked in free text. Your AI agent cannot reason about a patient's history if that history exists only in a scanned PDF from 2019 or a faxed prior authorization form.
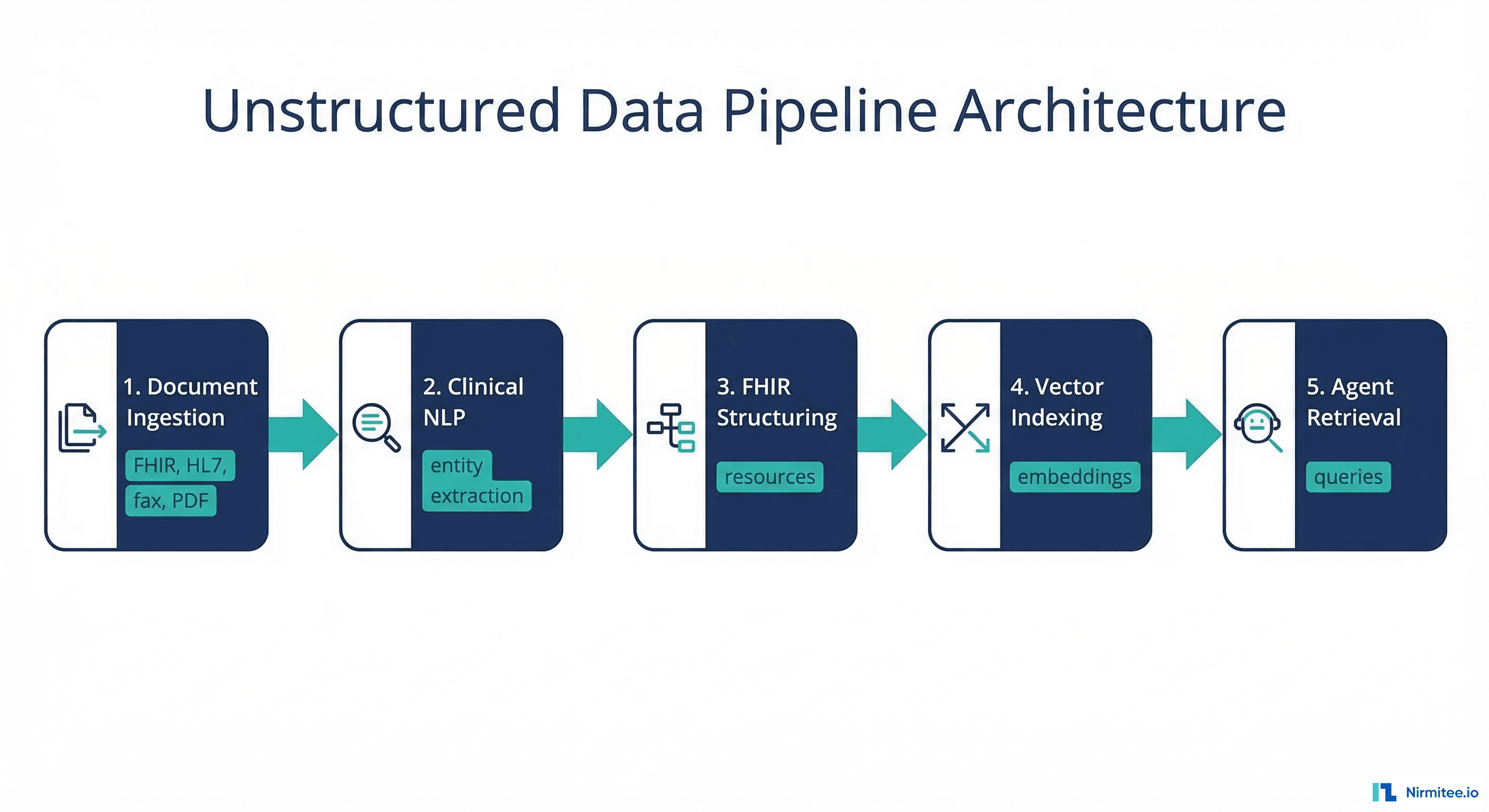
This guide is the practical HOW. We cover the complete pipeline: document ingestion (FHIR DocumentReference, HL7 MDM messages, fax OCR, scanned PDF OCR), clinical NLP (medical entity extraction, negation detection, temporal reasoning), structuring extracted data into FHIR resources, vector indexing for retrieval-augmented generation (RAG), and agent retrieval patterns. Every stage includes working Python code.
For the foundational context on why unstructured data is the critical blocker for healthcare AI, see our companion guide on Prerequisites Before Building an AI Agent for Healthcare.
Stage 1: Document Ingestion
Before any NLP runs, you need to get the documents into your pipeline. Healthcare documents arrive through four primary channels:
FHIR DocumentReference
Modern EHRs expose clinical documents through the FHIR DocumentReference resource. The document content may be inline (base64-encoded) or referenced via a URL to a Binary resource:
# fhir_document_ingestion.py
import base64
import httpx
from typing import Optional
class FHIRDocumentIngester:
"""Ingest clinical documents from FHIR DocumentReference resources."""
def __init__(self, fhir_base: str, token: str):
self.client = httpx.Client(
base_url=fhir_base,
headers={
"Authorization": f"Bearer {token}",
"Accept": "application/fhir+json"
},
timeout=30.0
)
def get_patient_documents(self, patient_id: str,
category: str = None) -> list[dict]:
"""Fetch all documents for a patient."""
params = {"patient": patient_id, "_sort": "-date"}
if category:
params["category"] = category
resp = self.client.get("/DocumentReference", params=params)
resp.raise_for_status()
bundle = resp.json()
return [e["resource"] for e in bundle.get("entry", [])]
def extract_text(self, doc_ref: dict) -> Optional[str]:
"""Extract text content from a DocumentReference."""
for content in doc_ref.get("content", []):
attachment = content.get("attachment", {})
content_type = attachment.get("contentType", "")
# Inline base64 content
if "data" in attachment:
raw = base64.b64decode(attachment["data"])
if "text" in content_type:
return raw.decode("utf-8")
elif "pdf" in content_type:
return self._extract_pdf_text(raw)
# Referenced Binary resource
elif "url" in attachment:
binary_resp = self.client.get(attachment["url"])
if binary_resp.status_code == 200:
if "text" in content_type:
return binary_resp.text
elif "pdf" in content_type:
return self._extract_pdf_text(binary_resp.content)
return None
def _extract_pdf_text(self, pdf_bytes: bytes) -> str:
"""Extract text from PDF using PyMuPDF."""
import fitz # PyMuPDF
doc = fitz.open(stream=pdf_bytes, filetype="pdf")
text = ""
for page in doc:
text += page.get_text()
return textHL7 v2 MDM Messages
Many hospitals still send clinical documents through HL7 v2 MDM (Medical Document Management) messages. The document text lives in the OBX-5 segment. If your integration handles HL7 v2, see our guide on Turning HL7 v2 Streams into FHIR APIs Using Mirth Connect for the conversion pipeline.
Fax and Scanned Document OCR
Fax remains a primary communication channel in US healthcare — see our analysis of Why Fax Remains Mandatory in Healthcare. Scanned faxes and PDFs require OCR before NLP processing:
# ocr_pipeline.py
import io
from PIL import Image
def ocr_document(image_bytes: bytes, engine: str = "tesseract") -> str:
"""OCR a scanned document image."""
if engine == "tesseract":
import pytesseract
image = Image.open(io.BytesIO(image_bytes))
# Pre-process for better OCR accuracy
image = image.convert("L") # Grayscale
text = pytesseract.image_to_string(
image,
config="--psm 6 --oem 3" # Assume uniform block of text, LSTM engine
)
return text
elif engine == "google_vision":
from google.cloud import vision
client = vision.ImageAnnotatorClient()
image = vision.Image(content=image_bytes)
response = client.document_text_detection(image=image)
return response.full_text_annotation.text
elif engine == "aws_textract":
import boto3
client = boto3.client("textract")
response = client.detect_document_text(
Document={"Bytes": image_bytes}
)
blocks = [b["Text"] for b in response["Blocks"] if b["BlockType"] == "LINE"]
return "\n".join(blocks)
raise ValueError(f"Unknown OCR engine: {engine}")For production OCR, Google Cloud Vision and AWS Textract significantly outperform Tesseract on handwritten notes and low-quality fax scans. Budget $1.50 per 1000 pages for cloud OCR.
Stage 2: Clinical NLP
Once you have text, clinical NLP extracts structured medical concepts. This is fundamentally different from general NLP — clinical language is dense, abbreviated, full of negation, and uses domain-specific terminology.
Medical Named Entity Recognition (NER)
Medical NER identifies mentions of clinical concepts in text: conditions, medications, procedures, lab values, anatomical sites, and dosages. Here is a complete pipeline using spaCy and medspaCy:
# clinical_nlp.py
import spacy
import medspacy
from medspacy.ner import TargetRule
from medspacy.context import ConTextRule
from dataclasses import dataclass, field
from typing import Optional
@dataclass
class ClinicalEntity:
text: str
label: str # CONDITION, MEDICATION, PROCEDURE, LAB_VALUE
start: int
end: int
is_negated: bool = False
is_historical: bool = False
is_family_history: bool = False
temporal_info: Optional[str] = None
confidence: float = 0.0
standard_code: Optional[str] = None
standard_system: Optional[str] = None
class ClinicalNLPPipeline:
"""Full clinical NLP pipeline for medical entity extraction."""
def __init__(self):
self.nlp = medspacy.load(enable=["medspacy_pyrush",
"medspacy_context"])
self._add_target_rules()
self._add_context_rules()
def _add_target_rules(self):
"""Add clinical concept patterns for NER."""
target_rules = [
# Conditions
TargetRule("diabetes", "CONDITION", pattern=[{"LOWER": {"IN": ["diabetes", "dm", "dm2", "dm1"]}}]),
TargetRule("hypertension", "CONDITION", pattern=[{"LOWER": {"IN": ["hypertension", "htn", "high blood pressure"]}}]),
TargetRule("heart failure", "CONDITION", pattern=[{"LOWER": "heart"}, {"LOWER": "failure"}]),
TargetRule("COPD", "CONDITION", pattern=[{"LOWER": {"IN": ["copd", "chronic obstructive pulmonary disease"]}}]),
# Medications
TargetRule("metformin", "MEDICATION", pattern=[{"LOWER": "metformin"}]),
TargetRule("lisinopril", "MEDICATION", pattern=[{"LOWER": "lisinopril"}]),
TargetRule("insulin", "MEDICATION", pattern=[{"LOWER": "insulin"}]),
# Procedures
TargetRule("colonoscopy", "PROCEDURE", pattern=[{"LOWER": "colonoscopy"}]),
TargetRule("MRI", "PROCEDURE", pattern=[{"LOWER": {"IN": ["mri", "magnetic resonance imaging"]}}]),
]
self.nlp.get_pipe("medspacy_target_matcher").add(target_rules)
def _add_context_rules(self):
"""Add context rules for negation, historical, family history."""
context_rules = [
ConTextRule("no evidence of", "NEGATED_EXISTENCE", direction="forward"),
ConTextRule("denies", "NEGATED_EXISTENCE", direction="forward"),
ConTextRule("no history of", "NEGATED_EXISTENCE", direction="forward"),
ConTextRule("ruled out", "NEGATED_EXISTENCE", direction="backward"),
ConTextRule("history of", "HISTORICAL", direction="forward"),
ConTextRule("previously", "HISTORICAL", direction="forward"),
ConTextRule("family history of", "FAMILY", direction="forward"),
ConTextRule("mother had", "FAMILY", direction="forward"),
ConTextRule("father had", "FAMILY", direction="forward"),
]
self.nlp.get_pipe("medspacy_context").add(context_rules)
def process(self, text: str) -> list[ClinicalEntity]:
"""Process clinical text and return extracted entities."""
doc = self.nlp(text)
entities = []
for ent in doc.ents:
entity = ClinicalEntity(
text=ent.text,
label=ent.label_,
start=ent.start_char,
end=ent.end_char,
is_negated=ent._.is_negated,
is_historical=ent._.is_historical,
is_family_history=ent._.is_family,
confidence=0.85 # Base confidence for rule-based NER
)
entities.append(entity)
return entities
# Example usage
if __name__ == "__main__":
nlp_pipeline = ClinicalNLPPipeline()
clinical_note = """
Patient is a 62-year-old male with a history of type 2 diabetes
and hypertension, currently on metformin 1000mg BID and lisinopril 20mg daily.
Patient denies chest pain. No history of heart failure.
Family history of COPD (mother). Colonoscopy scheduled for next month.
Last HbA1c was 7.2% (2026-01-15).
"""
entities = nlp_pipeline.process(clinical_note)
for e in entities:
neg = " [NEGATED]" if e.is_negated else ""
hist = " [HISTORICAL]" if e.is_historical else ""
fam = " [FAMILY]" if e.is_family_history else ""
print(f" {e.label}: {e.text}{neg}{hist}{fam}")Negation Detection: The Make-or-Break Feature
Negation detection is the single most important feature in clinical NLP. Without it, "patient denies chest pain" becomes a positive finding of chest pain. "No history of diabetes" becomes a diabetes diagnosis. Getting negation wrong does not just reduce accuracy — it creates clinically dangerous false positives.
The medspaCy ConText algorithm handles common negation patterns, but production systems need additional patterns for clinical abbreviations: "r/o" (rule out), "neg" (negative), "w/o" (without), and assertion-modifying phrases like "possible," "suspected," and "unlikely."
Stage 3: Structuring Into FHIR Resources
After NLP extracts clinical entities, convert them into proper FHIR resources. This is where the extracted intelligence becomes computable:
# fhir_structuring.py
from datetime import datetime
import uuid
class FHIRStructurer:
"""Convert extracted clinical entities into FHIR resources."""
def entity_to_fhir(self, entity, patient_id: str,
encounter_id: str = None) -> dict | None:
"""Convert a ClinicalEntity to a FHIR resource."""
if entity.is_negated:
return None # Don't create resources for negated findings
if entity.label == "CONDITION":
return self._to_condition(entity, patient_id, encounter_id)
elif entity.label == "MEDICATION":
return self._to_medication_statement(entity, patient_id)
elif entity.label == "PROCEDURE":
return self._to_procedure(entity, patient_id, encounter_id)
elif entity.label == "LAB_VALUE":
return self._to_observation(entity, patient_id)
return None
def _to_condition(self, entity, patient_id, encounter_id) -> dict:
resource = {
"resourceType": "Condition",
"id": str(uuid.uuid4()),
"clinicalStatus": {
"coding": [{
"system": "http://terminology.hl7.org/CodeSystem/condition-clinical",
"code": "active" if not entity.is_historical else "resolved"
}]
},
"verificationStatus": {
"coding": [{
"system": "http://terminology.hl7.org/CodeSystem/condition-ver-status",
"code": "confirmed" if entity.confidence > 0.9 else "provisional"
}]
},
"code": {"text": entity.text},
"subject": {"reference": f"Patient/{patient_id}"},
"meta": {
"tag": [{
"system": "https://your-org.com/nlp-extraction",
"code": "nlp-derived",
"display": f"NLP confidence: {entity.confidence:.2f}"
}]
}
}
if entity.standard_code:
resource["code"]["coding"] = [{
"system": entity.standard_system,
"code": entity.standard_code
}]
if encounter_id:
resource["encounter"] = {"reference": f"Encounter/{encounter_id}"}
return resource
def _to_medication_statement(self, entity, patient_id) -> dict:
return {
"resourceType": "MedicationStatement",
"id": str(uuid.uuid4()),
"status": "active",
"medicationCodeableConcept": {"text": entity.text},
"subject": {"reference": f"Patient/{patient_id}"},
"meta": {
"tag": [{
"system": "https://your-org.com/nlp-extraction",
"code": "nlp-derived",
"display": f"NLP confidence: {entity.confidence:.2f}"
}]
}
}
def _to_procedure(self, entity, patient_id, encounter_id) -> dict:
resource = {
"resourceType": "Procedure",
"id": str(uuid.uuid4()),
"status": "completed" if entity.is_historical else "preparation",
"code": {"text": entity.text},
"subject": {"reference": f"Patient/{patient_id}"},
"meta": {
"tag": [{
"system": "https://your-org.com/nlp-extraction",
"code": "nlp-derived"
}]
}
}
if encounter_id:
resource["encounter"] = {"reference": f"Encounter/{encounter_id}"}
return resource
def _to_observation(self, entity, patient_id) -> dict:
return {
"resourceType": "Observation",
"id": str(uuid.uuid4()),
"status": "final",
"code": {"text": entity.text},
"subject": {"reference": f"Patient/{patient_id}"},
"meta": {
"tag": [{
"system": "https://your-org.com/nlp-extraction",
"code": "nlp-derived"
}]
}
}Critical detail: every NLP-derived FHIR resource is tagged with a meta.tag marking it as NLP-derived with a confidence score. This lets downstream consumers distinguish between clinician-entered structured data and machine-extracted data. Never mix NLP-derived resources with clinician-verified data without clear provenance. This ties into the broader data quality considerations covered in Medallion Architecture for Healthcare Data.
Stage 4: Vector Indexing for RAG
For AI agents using retrieval-augmented generation, clinical text needs to be chunked, embedded, and indexed in a vector database:
# vector_indexing.py
from dataclasses import dataclass
from typing import Optional
import hashlib
@dataclass
class ClinicalChunk:
text: str
patient_id: str
document_id: str
document_type: str # progress_note, discharge_summary, lab_report
document_date: str
chunk_index: int
embedding: list[float] = None
metadata: dict = None
class ClinicalChunker:
"""Chunk clinical documents for vector indexing."""
def __init__(self, chunk_size: int = 512, overlap: int = 64):
self.chunk_size = chunk_size
self.overlap = overlap
def chunk_document(self, text: str, patient_id: str,
document_id: str, document_type: str,
document_date: str) -> list[ClinicalChunk]:
"""Chunk clinical text with section-aware splitting."""
# Split by clinical sections first
sections = self._split_by_sections(text)
chunks = []
chunk_idx = 0
for section_name, section_text in sections:
# Sub-chunk sections that exceed chunk_size
words = section_text.split()
for i in range(0, len(words), self.chunk_size - self.overlap):
chunk_words = words[i:i + self.chunk_size]
chunk_text = " ".join(chunk_words)
if len(chunk_text.strip()) < 20:
continue
chunks.append(ClinicalChunk(
text=chunk_text,
patient_id=patient_id,
document_id=document_id,
document_type=document_type,
document_date=document_date,
chunk_index=chunk_idx,
metadata={
"section": section_name,
"chunk_hash": hashlib.md5(
chunk_text.encode()
).hexdigest()
}
))
chunk_idx += 1
return chunks
def _split_by_sections(self, text: str) -> list[tuple[str, str]]:
"""Split clinical text by common section headers."""
import re
section_pattern = re.compile(
r'^(CHIEF COMPLAINT|HISTORY OF PRESENT ILLNESS|'
r'PAST MEDICAL HISTORY|MEDICATIONS|ALLERGIES|'
r'REVIEW OF SYSTEMS|PHYSICAL EXAMINATION|'
r'ASSESSMENT|PLAN|DISCHARGE INSTRUCTIONS|'
r'HPI|PMH|ROS|PE|A/P):?\s*$',
re.MULTILINE | re.IGNORECASE
)
sections = []
matches = list(section_pattern.finditer(text))
if not matches:
return [("FULL_TEXT", text)]
for i, match in enumerate(matches):
section_name = match.group(1).upper()
start = match.end()
end = matches[i + 1].start() if i + 1 < len(matches) else len(text)
section_text = text[start:end].strip()
if section_text:
sections.append((section_name, section_text))
return sections
class VectorIndexer:
"""Index clinical chunks in a vector database."""
def __init__(self, collection_name: str = "clinical_documents"):
# Using chromadb for simplicity; swap for Pinecone/Weaviate in production
import chromadb
self.client = chromadb.PersistentClient(path="./chroma_db")
self.collection = self.client.get_or_create_collection(
name=collection_name,
metadata={"hnsw:space": "cosine"}
)
def index_chunks(self, chunks: list[ClinicalChunk]):
"""Index chunks with embeddings and metadata."""
ids = [f"{c.document_id}_{c.chunk_index}" for c in chunks]
documents = [c.text for c in chunks]
metadatas = [{
"patient_id": c.patient_id,
"document_type": c.document_type,
"document_date": c.document_date,
"section": c.metadata.get("section", "unknown"),
} for c in chunks]
self.collection.add(
ids=ids, documents=documents, metadatas=metadatas
)
def query(self, query_text: str, patient_id: str = None,
document_type: str = None, n_results: int = 5) -> list[dict]:
"""Query the vector index for relevant clinical chunks."""
where_filter = {}
if patient_id:
where_filter["patient_id"] = patient_id
if document_type:
where_filter["document_type"] = document_type
results = self.collection.query(
query_texts=[query_text],
n_results=n_results,
where=where_filter if where_filter else None
)
return [{
"text": doc,
"metadata": meta,
"distance": dist
} for doc, meta, dist in zip(
results["documents"][0],
results["metadatas"][0],
results["distances"][0]
)]Stage 5: Agent Retrieval
The final stage connects the vector index to your AI agent. The agent queries clinical data by concept, filters by date and document type, and uses the retrieved context to answer clinical questions:
# agent_retrieval.py
class ClinicalRetriever:
"""Retrieval interface for AI agents to query clinical documents."""
def __init__(self, vector_indexer: VectorIndexer,
fhir_client=None):
self.vector_index = vector_indexer
self.fhir_client = fhir_client
def get_clinical_context(self, patient_id: str,
query: str,
max_chunks: int = 10) -> dict:
"""Get comprehensive clinical context for a patient query."""
# Retrieve relevant unstructured data
unstructured = self.vector_index.query(
query_text=query,
patient_id=patient_id,
n_results=max_chunks
)
# Retrieve structured FHIR data if client available
structured = {}
if self.fhir_client:
structured = {
"conditions": self.fhir_client.get_conditions(patient_id),
"medications": self.fhir_client.get_medications(patient_id),
"recent_labs": self.fhir_client.get_observations(
patient_id, category="laboratory"
)
}
return {
"patient_id": patient_id,
"query": query,
"unstructured_context": unstructured,
"structured_data": structured,
"retrieval_metadata": {
"chunks_retrieved": len(unstructured),
"has_structured_data": bool(structured)
}
}
def summarize_for_prompt(self, context: dict,
max_tokens: int = 2000) -> str:
"""Format retrieved context for an LLM prompt."""
parts = []
parts.append(f"=== Clinical Context for Patient {context['patient_id']} ===")
# Add structured data summary
if context.get("structured_data"):
sd = context["structured_data"]
if sd.get("conditions"):
conditions = [c.get("code", {}).get("text", "Unknown")
for c in sd["conditions"][:10]]
parts.append(f"Active Conditions: {', '.join(conditions)}")
if sd.get("medications"):
meds = [m.get("medicationCodeableConcept", {}).get("text", "Unknown")
for m in sd["medications"][:10]]
parts.append(f"Active Medications: {', '.join(meds)}")
# Add relevant unstructured chunks
parts.append("\n=== Relevant Clinical Notes ===")
for chunk in context.get("unstructured_context", []):
doc_type = chunk["metadata"].get("document_type", "note")
doc_date = chunk["metadata"].get("document_date", "unknown")
parts.append(f"[{doc_type} | {doc_date}]")
parts.append(chunk["text"])
parts.append("---")
return "\n".join(parts)This retrieval layer gives your AI agent both structured FHIR data and semantically relevant unstructured clinical text. The agent can reason across both: "The patient has active diabetes (structured) and the progress note from January mentions poor medication adherence (unstructured)." For patterns on building AI agents that use this data effectively, see our guide on Observability for Agentic AI in Healthcare.
NLP Tool Comparison
| Tool | Type | Best For | Cost | PHI Handling |
|---|---|---|---|---|
| spaCy + medspaCy | Open Source | Custom clinical NLP pipelines | Free | On-premise |
| Amazon Comprehend Medical | Cloud API | Quick start, medication extraction | $0.01/unit | BAA available |
| Google Healthcare NLP | Cloud API | High-volume processing | Pay per use | BAA available |
| Hugging Face Clinical Models | Open Source | Custom fine-tuning, research | Free | On-premise |
| John Snow Labs Spark NLP | Commercial | Enterprise clinical NLP | License | On-premise |
For HIPAA-compliant production deployments, the choice is between on-premise open-source tools (spaCy/medspaCy, Hugging Face) and cloud APIs with BAAs (AWS, Google). Cloud APIs are faster to deploy but create PHI processing dependencies. On-premise gives you full control but requires ML infrastructure. Many teams use a hybrid: cloud OCR for document ingestion, on-premise NLP for entity extraction.
Frequently Asked Questions
How accurate is clinical NLP compared to manual chart review?
State-of-the-art clinical NER achieves 85-95% F1 scores for common entity types (medications, conditions, procedures). Negation detection typically achieves 90-95% accuracy with ConText-based approaches. However, accuracy varies significantly by document type: structured progress notes perform better than handwritten scanned documents. Always validate NLP output against a gold-standard annotated dataset before production use.
How do I handle PHI in the NLP pipeline?
All text processing happens within your HIPAA-compliant environment. Never send PHI to a cloud NLP service without a BAA in place. For vector databases, ensure they are deployed on HIPAA-eligible infrastructure (not a developer laptop). De-identification should happen early in the pipeline if downstream consumers do not need PHI.
What embedding model should I use for clinical text?
General-purpose embeddings (OpenAI text-embedding-3, Cohere embed) work surprisingly well for clinical text. Clinical-specific models like PubMedBERT and ClinicalBERT provide marginal accuracy improvements for medical concept retrieval. In practice, the chunk size and section-aware splitting have more impact on retrieval quality than the embedding model choice.
How do I handle multi-language clinical documents?
In the US, clinical documents are predominantly English, but patient-facing documents (consent forms, discharge instructions) may be in Spanish, Mandarin, or other languages. Use language detection first (fastText langdetect), then route to language-specific NLP pipelines. AWS Comprehend Medical and Google Healthcare NLP support limited multilingual capabilities.
What is the latency budget for real-time NLP processing?
For real-time use cases (clinical decision support during a visit), target under 2 seconds end-to-end: OCR (500ms), NLP (800ms), FHIR structuring (200ms), vector indexing (500ms). For batch processing (nightly population health analytics), latency is less critical — focus on throughput (documents per minute).
Conclusion
The unstructured data pipeline is what separates a proof-of-concept AI agent from a production-grade one. Without it, your agent is blind to 80% of the clinical picture. With it, your agent can reason across discharge summaries, progress notes, scanned referral letters, and faxed prior authorization forms — the same information a clinician uses to make decisions.
Start with Stage 1 (document ingestion) and Stage 2 (basic NER). Get extraction accuracy above 90% before moving to FHIR structuring and vector indexing. The pipeline is only as good as its NLP layer — invest in annotation, validation, and continuous improvement.
For the architectural foundations that support this pipeline, see our guides on Domain-Driven Design in Healthcare and Building an AI Clinical Scribe.