Every health system in the US is sitting on a goldmine. Decades of clinical notes, lab results, imaging reports, medication histories, and longitudinal patient journeys — the exact data that could train the next generation of clinical AI models. The problem? HIPAA makes sharing it terrifying, and for good reason. A single breach of Protected Health Information (PHI) carries penalties of up to $2.1 million per violation category, per year. The 2024 Change Healthcare breach alone affected 100 million patients and is projected to cost UnitedHealth Group over $2.5 billion.
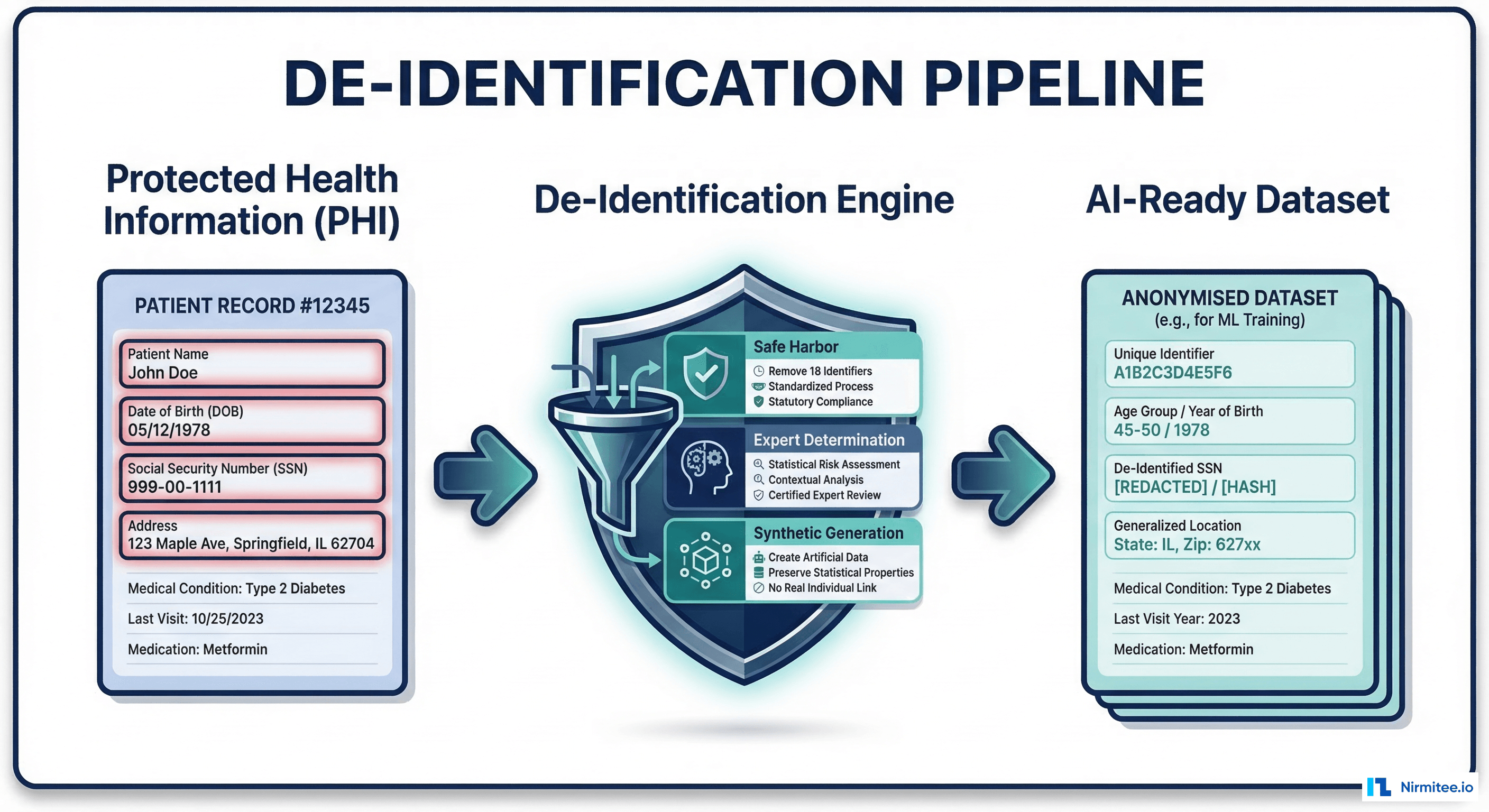
De-identification is the bridge between clinical data locked in EHRs and AI-ready datasets that can safely leave institutional walls. But the primary framework most organizations rely on — HIPAA's Safe Harbor method — was written in 2000, before social media, wearable devices, genomic sequencing, and the re-identification research that has fundamentally changed our understanding of privacy risk.
This guide is for the data engineers, ML teams, and compliance officers who need to actually build de-identification pipelines. We will cover what works, what doesn't, and how to choose between Safe Harbor, Expert Determination, and synthetic data generation based on your specific use case.
The Safe Harbor Method: All 18 Identifiers, with Practical Notes
HIPAA's Safe Harbor method (45 CFR 164.514(b)) is the most commonly used de-identification approach because it is prescriptive: remove or generalize these 18 specific identifiers, and your data is considered de-identified. No statistician required. No risk assessment needed.

Here are all 18 identifiers with the implementation details that matter for engineering teams:
| # | Identifier | What to Remove/Generalize | Engineering Notes |
|---|---|---|---|
| 1 | Names | Full name, initials, nicknames | Watch for names embedded in clinical notes, not just structured fields |
| 2 | Geographic data | Anything more specific than state | Zip codes: keep first 3 digits only if the 3-digit zip population is >20,000. Otherwise, set to 000. Per 2020 Census, 17 three-digit zips must be zeroed |
| 3 | Dates | All dates except year | Birth dates, admission dates, discharge dates, death dates — all reduced to year only. Ages over 89 must be grouped into a single category ("90+") |
| 4 | Phone numbers | All phone numbers | Check clinical notes for phone numbers documented by clinicians |
| 5 | Fax numbers | All fax numbers | Still common in healthcare — referral documents, pharmacy communications |
| 6 | Email addresses | All email addresses | Patient portal emails, provider contact fields, and free-text notes |
| 7 | Social Security numbers | Full SSN | May appear in insurance fields, billing records, or legacy system imports |
| 8 | Medical record numbers | MRNs, encounter IDs | Replace with pseudonymized IDs (hash + salt) to maintain linkability within datasets |
| 9 | Health plan beneficiary numbers | Insurance IDs, Medicare/Medicaid numbers | Found in claims data, eligibility responses, EOBs |
| 10 | Account numbers | All account numbers | Billing account numbers, guarantor accounts |
| 11 | Certificate/license numbers | Driver's license, professional licenses | Sometimes captured during patient registration |
| 12 | Vehicle identifiers | VIN, license plate numbers | Rare in clinical data, but present in trauma/accident reports |
| 13 | Device identifiers | UDI, serial numbers | Implant records, FHIR Device resources, IoMT device logs |
| 14 | Web URLs | Patient portal URLs, any URLs | Check for URLs in clinical notes referencing patient-specific pages |
| 15 | IP addresses | All IP addresses | Audit logs, patient portal access records, telehealth session data |
| 16 | Biometric identifiers | Fingerprints, voiceprints, retinal scans | Increasingly common with biometric authentication at registration kiosks |
| 17 | Full-face photographs | Any comparable image | Dermatology images, wound care photos — crop or blur facial features |
| 18 | Any other unique identifying number | Catch-all for unique identifiers | Includes study IDs that could link back to identified datasets, genetic accession numbers, biobank specimen IDs |
The Safe Harbor method has a critical advantage: legal certainty. If you remove all 18 identifiers and have no actual knowledge that the remaining data could identify someone, the data is legally de-identified under HIPAA. No expert review. No statistical proof. This makes it the default choice for most health systems and research institutions.
Why Safe Harbor Is Showing Its Age

The Safe Harbor regulation was finalized in August 2000. That is before Facebook (2004), before the iPhone (2007), before consumer genomics (23andMe launched in 2006), and before wearable health devices became ubiquitous. The privacy landscape it was designed for no longer exists.
The Re-Identification Research That Should Worry You
In 2000, Latanya Sweeney's landmark research demonstrated that 87% of the US population could be uniquely identified using just three data points: 5-digit zip code, date of birth, and gender. These are all quasi-identifiers that Safe Harbor allows in some form — zip codes (first 3 digits), birth year, and gender are all permitted in Safe Harbor-compliant datasets.
More recent studies have escalated the concern:
- 2019 — Nature Communications: Researchers at Imperial College London and Université catholique de Louvain showed that 99.98% of Americans could be re-identified in any dataset using 15 demographic attributes, even when the data was "anonymized" (Rocher et al., 2019).
- 2013 — Science: MIT researchers re-identified individuals in a credit card metadata dataset with 90% accuracy using just four spatiotemporal data points (de Montjoye et al., 2015).
- Genetic data: A 2018 study in Science showed that 60% of Americans with European ancestry could be identified through genetic genealogy databases, even if they had never submitted their own DNA (Erlich et al., 2018).
What Safe Harbor Does Not Cover
The 18 identifiers were designed for the data types that existed in 2000. They do not explicitly address:
- Social media correlation: A patient's Twitter/X post about their hospital visit, combined with de-identified admission data, can dramatically narrow identification.
- Wearable device data: Continuous heart rate, GPS, sleep patterns — even without names, the temporal patterns are highly unique
- Genomic data: Genetic sequences are inherently identifying and are not covered by Safe Harbor's 18 identifiers
- Geolocation breadcrumbs: Appointment times correlated with cell tower data or location history
- Rare disease data: In small populations, diagnosis codes alone can identify patients — only 7,000 people in the US have Huntington's disease
- Longitudinal data: Even without direct identifiers, a unique combination of diagnoses, procedures, and visit patterns over time can fingerprint an individual
Safe Harbor is not wrong — it is incomplete for modern data environments, especially when building AI training datasets that combine multiple data sources.
Expert Determination: The Flexible (but Expensive) Alternative
HIPAA's second de-identification method — Expert Determination (45 CFR 164.514(a)) — requires a qualified statistician or scientist to apply statistical and scientific principles and determine that the risk of re-identification is "very small." The expert must document their methods and results.
When Expert Determination Makes Sense
- You need date precision: AI models for sepsis prediction, readmission risk, or disease progression need actual dates, not just years. Expert Determination can allow date retention with other controls.
- Geographic analysis: Social determinants of health research require sub-state geography. An expert can determine acceptable geographic granularity based on population density.
- Small datasets: Safe Harbor is riskier with small datasets, where combinations of permitted attributes (age, gender, zip prefix) might still identify individuals. Expert Determination applies quantitative risk assessment.
- Multi-source data: When combining clinical, claims, and FHIR-based data pipelines, the linkage itself creates re-identification risk that Safe Harbor does not address.
What Expert Determination Costs
A typical Expert Determination engagement runs $50,000 to $200,000, depending on dataset complexity, number of data elements, and the expert's assessment of re-identification risk. The process typically takes 8-16 weeks and includes:
- Data profiling and quasi-identifier analysis
- Population uniqueness assessment
- Risk quantification using k-anonymity, l-diversity, or other privacy models
- Transformation recommendations (suppression, generalization, perturbation)
- Formal certification letter
Organizations like Privacy Analytics (an IQVIA company) and academic groups at institutions like Vanderbilt and Harvard specialize in Expert Determination for healthcare datasets.
Practical Tooling for Data Engineers

If you are building de-identification into your data pipeline — rather than hiring an external service — these are the tools that actually work in production:
Open-Source Libraries
| Tool | Maintainer | Best For | Notes |
|---|---|---|---|
| Presidio | Microsoft | NER-based PHI detection in clinical text | Supports custom recognizers, FHIR integration, multiple NLP backends (spaCy, Stanza, Transformers) |
| scrubadub | LeapBeyond | Quick text scrubbing | Good for structured text, less accurate on clinical notes than Presidio |
| Philter | UCSF | Clinical note de-identification | Purpose-built for clinical text, rule-based + ML hybrid, validated on i2b2 datasets |
| Synthea | MITRE | Fully synthetic patient generation | Generates complete FHIR Bundles with realistic clinical progressions |
De-Identifying a FHIR Patient Resource: Python Example
Here is a practical code snippet for de-identifying a FHIR Patient resource programmatically. This covers the most common Safe Harbor transformations:
import hashlib
import json
from datetime import datetime
def deidentify_fhir_patient(patient_resource: dict, salt: str) -> dict:
"""
De-identify a FHIR Patient resource per Safe Harbor method.
Removes direct identifiers, generalizes quasi-identifiers.
"""
patient = json.loads(json.dumps(patient_resource)) # deep copy
# 1. Remove names (Safe Harbor #1)
patient.pop("name", None)
# 2. Generalize address — keep only state (Safe Harbor #2)
if "address" in patient:
for addr in patient["address"]:
addr.pop("line", None)
addr.pop("city", None)
# Truncate zip to 3 digits; zero out if population < 20K
zip_code = addr.get("postalCode", "")
LOW_POP_PREFIXES = {"036", "059", "063", "102", "203",
"556", "692", "790", "821", "823",
"830", "831", "878", "879", "884",
"890", "893"}
if zip_code:
prefix = zip_code[:3]
addr["postalCode"] = "000" if prefix in LOW_POP_PREFIXES else prefix
addr.pop("district", None)
# 3. Generalize birthDate to year only (Safe Harbor #3)
if "birthDate" in patient:
try:
birth_year = int(patient["birthDate"][:4])
current_year = datetime.now().year
age = current_year - birth_year
if age > 89:
patient["birthDate"] = "1900" # grouped 90+
else:
patient["birthDate"] = str(birth_year)
except (ValueError, IndexError):
patient.pop("birthDate", None)
# 4. Remove telecom — phone, fax, email (Safe Harbor #4, 5, 6)
patient.pop("telecom", None)
# 5. Remove SSN from identifiers (Safe Harbor #7)
# 6. Hash MRN for linkability (Safe Harbor #8)
if "identifier" in patient:
cleaned_ids = []
for ident in patient["identifier"]:
system = ident.get("system", "")
if "ssn" in system.lower() or "social" in system.lower():
continue # remove SSN
if "mrn" in system.lower() or "medical-record" in system.lower():
# Pseudonymize MRN with salted hash
raw = ident.get("value", "")
ident["value"] = hashlib.sha256(
f"{salt}:{raw}".encode()
).hexdigest()[:16]
ident["system"] = "urn:oid:deidentified-mrn"
cleaned_ids.append(ident)
patient["identifier"] = cleaned_ids
# 7. Remove photo (Safe Harbor #17)
patient.pop("photo", None)
# 8. Strip extensions that may contain PHI
patient.pop("extension", None)
return patient
# Example usage
original_patient = {
"resourceType": "Patient",
"id": "example-patient-001",
"name": [{"family": "Smith", "given": ["John", "Michael"]}],
"birthDate": "1985-07-23",
"gender": "male",
"address": [{
"line": ["123 Main St"],
"city": "Boston",
"state": "MA",
"postalCode": "02115"
}],
"telecom": [
{"system": "phone", "value": "617-555-0123"},
{"system": "email", "value": "john.smith@email.com"}
],
"identifier": [
{"system": "urn:oid:mrn", "value": "MRN-12345678"},
{"system": "urn:oid:ssn", "value": "123-45-6789"}
]
}
deidentified = deidentify_fhir_patient(original_patient, salt="project-x-2026")
print(json.dumps(deidentified, indent=2))
# Output: no name, no telecom, no SSN, birth year only,
# zip truncated to "021", MRN hashedThis is a starting point. Production pipelines should also scan clinical notes (using Presidio or Philter) for PHI in free-text fields like Observation.valueString or DiagnosticReport.conclusion.
Synthetic Data: When You Don't Need Real Patients at All
For many AI training scenarios — especially development, testing, and algorithm prototyping — synthetic data eliminates the de-identification problem entirely. No real patients means no PHI, no HIPAA applicability, no risk.
Synthetic Data Tools That Work
- Synthea (MITRE): Generates fully synthetic patient records as FHIR Bundles. Includes realistic disease progressions, medication histories, and care encounters. Used by ONC, CMS, and hundreds of health IT companies for testing. Free and open source.
- Gretel.ai: Uses differential privacy and generative models to create synthetic datasets that preserve statistical properties of real data. Particularly useful when you need synthetic data that mirrors your actual patient population's distributions.
- MDClone: Creates "synthetic twins" — synthetic records that preserve the statistical relationships in real clinical data without exposing any individual patient. Used by health systems like Intermountain Health and the Children's Hospital of Philadelphia.
- MOSTLY AI: Enterprise synthetic data platform with healthcare-specific models. Includes privacy guarantees and utility metrics.
The trade-off with synthetic data is always fidelity versus privacy. Synthea generates realistic but generic patient journeys — useful for pipeline testing but not for training models that need to capture the specific patterns in your patient population. Tools like Gretel and MDClone bridge this gap by learning from real data to generate synthetic records, but the question of whether the synthetic data is "too similar" to the source data requires careful evaluation.
Beyond HIPAA: Privacy Models for AI Training Data

HIPAA defines the legal floor for de-identification, but it does not provide mathematical privacy guarantees. If you are building AI training datasets — especially datasets that will be shared with external partners or used in federated learning — you need to understand the formal privacy models:
k-Anonymity
A dataset satisfies k-anonymity if every combination of quasi-identifiers (age, gender, zip code) appears in at least k records. If k=5, every record "hides" among at least 4 others with the same quasi-identifier values. This is the most intuitive privacy model and is used in many Expert Determination assessments.
Limitation: If all 5 records in a k-anonymous group have the same sensitive value (e.g., all have HIV diagnosis), privacy is still breached. This led to l-diversity.
l-Diversity
Extends k-anonymity by requiring that each equivalence class (group of records with the same quasi-identifiers) has at least l distinct values for the sensitive attribute. This prevents the "homogeneity attack" where all records in a group share the same diagnosis.
t-Closeness
Goes further by requiring that the distribution of the sensitive attribute in each equivalence class is within distance t of the attribute's overall distribution. This prevents the "skewness attack" where the distribution within a group reveals information even when values are diverse.
Differential Privacy
The gold standard for mathematical privacy guarantees. A mechanism satisfies differential privacy if the output of a query on a dataset is statistically indistinguishable whether or not any single individual's record is included. The privacy budget (epsilon) quantifies the maximum information leakage. Apple, Google, and the US Census Bureau all use differential privacy.
For healthcare AI training, differential privacy is increasingly the recommended approach when training models on sensitive data. Libraries like Opacus (PyTorch) and TensorFlow Privacy add differential privacy guarantees directly to model training with minimal code changes.
Decision Framework: Choosing the Right Approach


Choosing between de-identification methods is not an either/or decision — many organizations layer multiple approaches. Here is a practical decision framework:
| Use Case | Recommended Approach | Why |
|---|---|---|
| Sharing data with external researchers | Safe Harbor + k-anonymity verification | Legal certainty of Safe Harbor with quantitative privacy guarantee |
| Training ML models on real clinical patterns | Expert Determination + differential privacy | Preserves clinical nuance while providing mathematical privacy bounds |
| Development and testing environments | Synthetic data (Synthea) | Zero PHI risk, free, generates valid FHIR resources |
| Federated learning across institutions | Differential privacy (Opacus/TF Privacy) | Data never leaves the institution; only model gradients are shared with privacy guarantees |
| Clinical trial data sharing | Expert Determination | Preserves date precision and geographic detail needed for trial analysis |
| Building FHIR-based AI pipelines | Safe Harbor for structured resources + Presidio for narrative text | Handles both coded data and unstructured clinical notes |
| Population health analytics | Safe Harbor with aggregation | Aggregate statistics (cohort-level) reduce re-identification risk to near zero |
A Layered Strategy for Production
The most robust approach combines multiple methods:
- Structured data: Apply Safe Harbor transformations programmatically (the Python code above covers FHIR resources)
- Clinical notes: Run through Presidio or Philter for NER-based PHI detection, then manual review of flagged content
- Quantitative validation: Measure k-anonymity and l-diversity on the output dataset; ensure k ≥ 5 for most use cases
- Model training: Apply differential privacy during model training as a second layer of protection
- Ongoing monitoring: Re-assess risk when adding new data sources or linking datasets
Compliance Considerations Beyond HIPAA
If your AI training data includes patients from outside the US, or if your models will be deployed internationally, HIPAA is just the starting point:
- GDPR (EU): Does not recognize Safe Harbor-style de-identification. Requires either anonymization (irreversible, assessed against "all means reasonably likely") or pseudonymization (still treated as personal data). Article 89 provides research exemptions but with strict safeguards.
- State laws: California's CCPA/CPRA, Washington's My Health My Data Act, and other state privacy laws may impose additional requirements beyond HIPAA.
- 21st Century Cures Act: While focused on information blocking, the Cures Act's data sharing requirements intersect with de-identification when health systems share data through TEFCA or other networks.
From predictive models to clinical AI, our Healthcare AI Solutions practice helps healthcare organizations deploy AI that delivers real outcomes. We also offer specialized Custom Healthcare Software Development services. Talk to our team to get started.
Frequently Asked Questions
Is de-identified data still protected under HIPAA?
No. Once data is properly de-identified under either Safe Harbor or Expert Determination, it is no longer considered PHI and is not subject to HIPAA restrictions. However, you must maintain the de-identification methodology documentation, and if you retain a re-identification key, the key itself is PHI.
Can I use Safe Harbor for genomic data?
Safe Harbor's 18 identifiers do not explicitly address genomic sequences, which are inherently identifying. For genomic data, Expert Determination is strongly recommended, and many experts consider genomic data impossible to fully de-identify. Synthetic genomic data or federated approaches may be the only viable path.
What is the re-identification risk threshold for Expert Determination?
HIPAA requires the risk to be "very small" but does not define a specific threshold. In practice, most experts target a re-identification risk below 0.04 (1 in 25) to 0.09 (1 in 11), depending on the sensitivity of the data and the anticipated recipient. The HHS guidance on de-identification provides additional context.
How do I de-identify medical images (X-rays, CT scans)?
DICOM images contain PHI in both the pixel data (burned-in text overlays with patient name, MRN) and the metadata headers. Tools like DicomAnonymizer handle metadata scrubbing. For burned-in PHI, OCR-based detection (using Tesseract + Presidio) or deep learning approaches are needed. The RSNA CTP (Clinical Trial Processor) is widely used for radiology image de-identification.
Building De-Identification Into Your Architecture
De-identification should not be an afterthought or a one-time export process. For organizations serious about leveraging clinical data for AI, it needs to be a first-class component of your data architecture — a pipeline stage that runs continuously as new data flows in.
At Nirmitee, we build healthcare data infrastructure with privacy as a core architectural concern, not a bolt-on. Our EHR and integration platforms are designed with FHIR-native data pipelines where de-identification can be applied at the resource level before data ever leaves the clinical system boundary. If you are building an AI-ready data infrastructure and need help getting the de-identification layer right, we would welcome the conversation.
The bottom line: Safe Harbor's 18 identifiers are still the right starting point, but they are not sufficient for modern healthcare AI. Layer Expert Determination, differential privacy, or synthetic data generation on top — and build the de-identification step into your pipeline, not your export process.