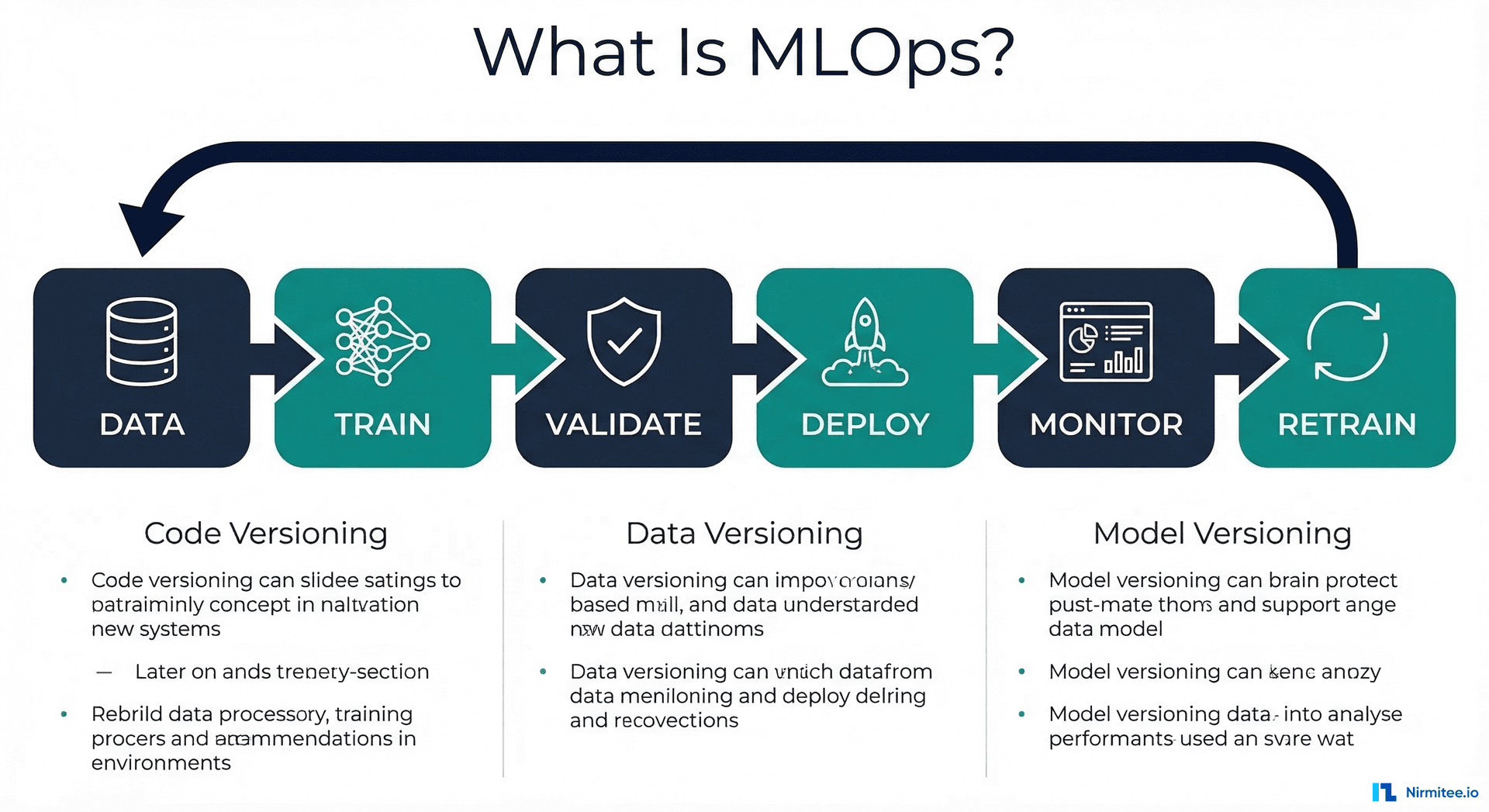
Why MLflow Is the Foundation of Every Healthcare MLOps Stack
If you ask any ML engineer what tool they use for experiment tracking, the answer is almost always MLflow. With over 18,000 GitHub stars and adoption by organizations from startups to Fortune 500 companies, MLflow has become the de facto standard for managing the machine learning lifecycle. But in healthcare, MLflow is not just convenient — it is essential. The combination of experiment reproducibility, model versioning, and audit trail capabilities maps directly to HIPAA, FDA, and clinical governance requirements.
This guide covers everything a healthcare engineering team needs to set up MLflow properly: from installation and experiment tracking to model registry workflows, HIPAA-safe practices, and production deployment. We include complete code examples that you can adapt for your clinical ML projects. For foundational context on what MLOps is and why healthcare needs it, see our introduction to MLOps for healthcare developers.

MLflow's Four Components
MLflow is composed of four integrated components, each serving a distinct role in the ML lifecycle:
1. MLflow Tracking
The experiment tracking server logs every training run with its parameters, metrics, and artifacts. This is your experiment journal — instead of keeping notes in a spreadsheet or naming notebooks model_v3_final_FINAL_v2.ipynb, every run is automatically recorded with its full context.
2. MLflow Model Registry
The model registry provides centralized model storage with versioning and stage transitions. Models move through stages — None, Staging, Production, Archived — with approval workflows at each transition. In healthcare, this maps directly to clinical review board approval gates.
3. MLflow Projects
Projects define reproducible runs by packaging code with environment specifications (conda.yaml or Dockerfile). Any team member can re-execute a training run and get identical results — critical for FDA audit requirements.
4. MLflow Models
The model packaging format supports multiple ML frameworks (scikit-learn, PyTorch, TensorFlow, XGBoost) with a unified serving interface. This means your deployment pipeline works the same regardless of the underlying framework.
Setting Up MLflow for Healthcare: Self-Hosted with PostgreSQL
For healthcare organizations, self-hosting MLflow is strongly preferred over using managed cloud services. The reason is simple: experiment metadata — parameter names, metric values, run descriptions — can inadvertently contain PHI if not carefully managed. Keeping the tracking server within your security boundary gives you full control over access and encryption.
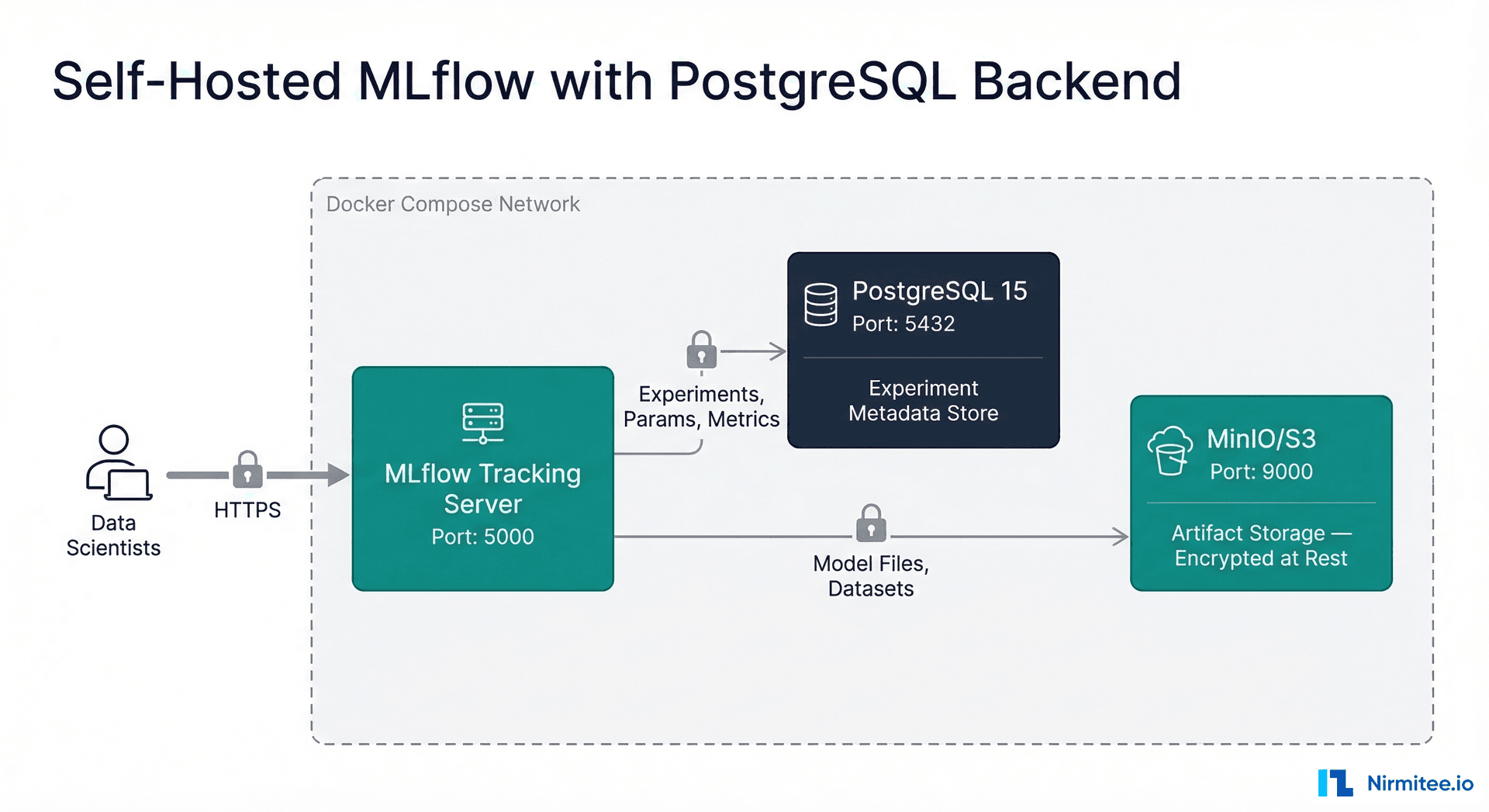
# docker-compose.yml — Self-hosted MLflow with PostgreSQL backend
version: "3.8"
services:
mlflow:
image: ghcr.io/mlflow/mlflow:2.12.1
container_name: mlflow-tracking
ports:
- "5000:5000"
environment:
- MLFLOW_BACKEND_STORE_URI=postgresql://mlflow:${DB_PASSWORD}@postgres:5432/mlflow
- MLFLOW_DEFAULT_ARTIFACT_ROOT=s3://mlflow-artifacts/
- AWS_ACCESS_KEY_ID=${MINIO_ACCESS_KEY}
- AWS_SECRET_ACCESS_KEY=${MINIO_SECRET_KEY}
- MLFLOW_S3_ENDPOINT_URL=http://minio:9000
command: >
mlflow server
--host 0.0.0.0
--port 5000
--backend-store-uri postgresql://mlflow:${DB_PASSWORD}@postgres:5432/mlflow
--default-artifact-root s3://mlflow-artifacts/
--serve-artifacts
depends_on:
postgres:
condition: service_healthy
minio:
condition: service_started
restart: unless-stopped
postgres:
image: postgres:15-alpine
container_name: mlflow-postgres
environment:
POSTGRES_DB: mlflow
POSTGRES_USER: mlflow
POSTGRES_PASSWORD: ${DB_PASSWORD}
volumes:
- pgdata:/var/lib/postgresql/data
healthcheck:
test: ["CMD-SHELL", "pg_isready -U mlflow"]
interval: 5s
timeout: 5s
retries: 5
restart: unless-stopped
minio:
image: minio/minio:latest
container_name: mlflow-minio
ports:
- "9000:9000"
- "9001:9001"
environment:
MINIO_ROOT_USER: ${MINIO_ACCESS_KEY}
MINIO_ROOT_PASSWORD: ${MINIO_SECRET_KEY}
volumes:
- minio_data:/data
command: server /data --console-address ":9001"
restart: unless-stopped
volumes:
pgdata:
driver: local
minio_data:
driver: local
# .env file (keep this secure, never commit to Git)
DB_PASSWORD=your_secure_password_here
MINIO_ACCESS_KEY=mlflow_minio_access
MINIO_SECRET_KEY=your_secure_minio_secret
# Start the stack
docker compose up -d
# Create the MinIO bucket
docker compose exec minio mc alias set local http://localhost:9000 $MINIO_ACCESS_KEY $MINIO_SECRET_KEY
docker compose exec minio mc mb local/mlflow-artifacts
# Verify MLflow is running
curl http://localhost:5000/health
Experiment Tracking: The HIPAA-Safe Way
Experiment tracking is where most healthcare teams unknowingly introduce compliance risks. The rule is simple but frequently violated: never log patient identifiers as experiment parameters or tags.

# Complete MLflow experiment tracking setup for healthcare
import mlflow
import mlflow.sklearn
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import (
roc_auc_score, precision_recall_curve,
classification_report, confusion_matrix
)
import numpy as np
import hashlib
from datetime import datetime
# Point to self-hosted tracking server
mlflow.set_tracking_uri("http://mlflow-server:5000")
mlflow.set_experiment("sepsis-early-warning-system")
def train_and_log_model(X_train, y_train, X_test, y_test, data_version, feature_set_name):
with mlflow.start_run(run_name=f"gbm-{data_version}-{datetime.now().strftime('%Y%m%d')}"):
# --- SAFE: Log aggregate parameters, never patient IDs ---
mlflow.log_params({
"model_type": "GradientBoostingClassifier",
"n_estimators": 200,
"max_depth": 6,
"learning_rate": 0.1,
"data_version": data_version, # e.g., "v2.3"
"feature_set": feature_set_name, # e.g., "vitals+labs+demo"
"train_size": len(X_train), # Aggregate count, not IDs
"test_size": len(X_test),
"positive_rate": f"{y_train.mean():.4f}",
"training_date": datetime.now().isoformat(),
"data_hash": hashlib.sha256( # Hash, not content
str(X_train.shape).encode()
).hexdigest()[:12]
})
# Train
model = GradientBoostingClassifier(
n_estimators=200, max_depth=6, learning_rate=0.1, random_state=42
)
model.fit(X_train, y_train)
# Evaluate
y_pred_proba = model.predict_proba(X_test)[:, 1]
y_pred = (y_pred_proba >= 0.5).astype(int)
# --- SAFE: Log aggregate metrics ---
mlflow.log_metrics({
"auc_roc": roc_auc_score(y_test, y_pred_proba),
"sensitivity": recall_score(y_test, y_pred),
"specificity": specificity_score(y_test, y_pred),
"ppv": precision_score(y_test, y_pred),
"npv": npv_score(y_test, y_pred),
"f1": f1_score(y_test, y_pred),
})
# Log confusion matrix as artifact (aggregate, no patient IDs)
cm = confusion_matrix(y_test, y_pred)
np.savetxt("/tmp/confusion_matrix.csv", cm, delimiter=",",
header="Predicted_Neg,Predicted_Pos", comments="")
mlflow.log_artifact("/tmp/confusion_matrix.csv")
# Log model with signature
from mlflow.models import infer_signature
signature = infer_signature(X_test, y_pred_proba)
mlflow.sklearn.log_model(
model,
"sepsis-model",
signature=signature,
registered_model_name="sepsis-early-warning"
)
print(f"Run logged: AUC={roc_auc_score(y_test, y_pred_proba):.4f}")
What to NEVER Log
- Patient IDs, MRNs, or names as parameters or tags
- Raw patient data files as artifacts (use de-identified or aggregated data only)
- Model prediction outputs with patient identifiers attached
- Screenshots or exports containing PHI
- Free-text run descriptions that mention specific patients
What You SHOULD Log
- Data version identifiers (e.g., "v2.3-post-covid-update")
- Aggregate statistics (cohort size, positive rate, feature missing rates)
- Hashed dataset identifiers for traceability
- Feature set names and preprocessing pipeline versions
- All performance metrics at clinically relevant thresholds
Model Registry: Clinical Approval Workflows
The MLflow Model Registry maps naturally to healthcare's clinical review and approval process. Each model version moves through stages that correspond to healthcare governance gates:

# Model Registry operations for healthcare workflow
from mlflow import MlflowClient
client = MlflowClient(tracking_uri="http://mlflow-server:5000")
# Register a new model version from the best experiment run
model_uri = "runs:/abc123def456/sepsis-model"
result = client.create_model_version(
name="sepsis-early-warning",
source=model_uri,
run_id="abc123def456",
description="GBM trained on v2.3 dataset, AUC 0.89, approved by clinical review"
)
print(f"Registered version: {result.version}")
# Transition to Staging (triggers shadow deployment)
client.transition_model_version_stage(
name="sepsis-early-warning",
version=result.version,
stage="Staging",
archive_existing_versions=False
)
# Add clinical review notes as model version tags
client.set_model_version_tag(
name="sepsis-early-warning",
version=result.version,
key="clinical_review_status",
value="approved"
)
client.set_model_version_tag(
name="sepsis-early-warning",
version=result.version,
key="clinical_reviewer",
value="CMIO-Dr-Johnson"
)
client.set_model_version_tag(
name="sepsis-early-warning",
version=result.version,
key="review_date",
value="2026-02-15"
)
client.set_model_version_tag(
name="sepsis-early-warning",
version=result.version,
key="fda_documentation_id",
value="DOC-2026-SEP-V23"
)
# After shadow validation succeeds: promote to Production
client.transition_model_version_stage(
name="sepsis-early-warning",
version=result.version,
stage="Production",
archive_existing_versions=True # Archive previous production version
)

RBAC: Access Control for Clinical Teams
MLflow's open-source version has limited built-in RBAC. For healthcare deployments, you need to add an access control layer. The recommended approaches are:

Option 1: Reverse Proxy with OAuth (Recommended)
# nginx.conf - OAuth2 proxy in front of MLflow
upstream mlflow {
server mlflow-tracking:5000;
}
server {
listen 443 ssl;
server_name mlflow.internal.hospital.org;
ssl_certificate /etc/ssl/certs/mlflow.crt;
ssl_certificate_key /etc/ssl/private/mlflow.key;
# OAuth2 authentication
auth_request /oauth2/auth;
error_page 401 = /oauth2/sign_in;
location / {
proxy_pass http://mlflow;
proxy_set_header X-User $remote_user;
proxy_set_header X-Groups $http_x_groups;
# Audit logging - log every access
access_log /var/log/nginx/mlflow_audit.log audit_format;
}
location /oauth2/ {
proxy_pass http://oauth2-proxy:4180;
}
}
Option 2: MLflow with Databricks (Managed)
If your organization uses Databricks, Managed MLflow provides enterprise RBAC out of the box — workspace-level permissions, experiment-level ACLs, and model registry approval workflows. This is the easiest path but requires a Databricks subscription and data residency considerations under HIPAA.
Encrypted Artifact Storage
Model artifacts (trained model files, preprocessing pipelines, evaluation datasets) must be encrypted at rest. When using S3-compatible storage:
# Configure MLflow to use encrypted S3 artifact storage
import boto3
# S3 bucket policy enforcing encryption
bucket_policy = {
"Version": "2012-10-17",
"Statement": [
{
"Sid": "DenyUnencryptedUploads",
"Effect": "Deny",
"Principal": "*",
"Action": "s3:PutObject",
"Resource": "arn:aws:s3:::mlflow-artifacts/*",
"Condition": {
"StringNotEquals": {
"s3:x-amz-server-side-encryption": "aws:kms"
}
}
}
]
}
# When logging artifacts, MLflow respects the bucket's encryption policy
# No code changes needed in your training scripts
FDA Audit Trail with MLflow
For models classified as Software as a Medical Device (SaMD), the FDA requires comprehensive documentation of the model development and validation process. MLflow provides most of this automatically — but you need to structure it deliberately.
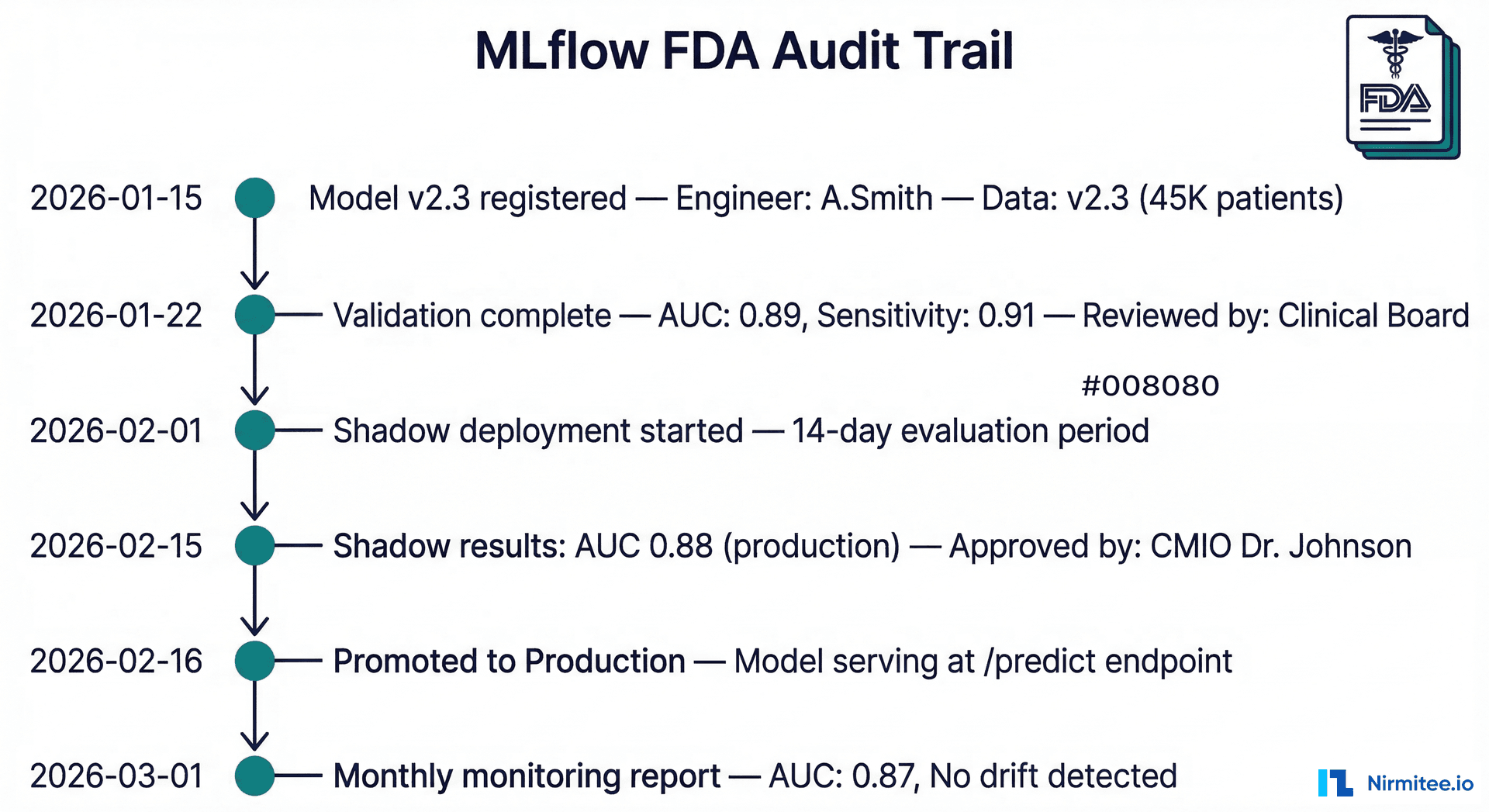

What the FDA Wants to See
| FDA Requirement | MLflow Feature | Implementation |
|---|---|---|
| Algorithm description | Run parameters + tags | Log model type, architecture, hyperparameters |
| Training data description | Run parameters | Log data version, size, date range, demographics |
| Performance evaluation | Run metrics | Log AUC, sensitivity, specificity, calibration |
| Validation methodology | Run tags + artifacts | Log CV strategy, test set description, holdout approach |
| Version history | Model Registry | All versions preserved with transition timestamps |
| Change control plan | Registry stage transitions | Document what triggers retraining and re-validation |
# Generate FDA-ready audit report from MLflow
def generate_fda_audit_report(model_name, version):
client = MlflowClient()
# Get model version details
mv = client.get_model_version(name=model_name, version=version)
run = client.get_run(mv.run_id)
report = {
"document_id": f"FDA-AUDIT-{model_name}-v{version}",
"generated_at": datetime.now().isoformat(),
"model_name": model_name,
"model_version": version,
"algorithm": {
"type": run.data.params.get("model_type"),
"framework": "scikit-learn",
"hyperparameters": {k: v for k, v in run.data.params.items()
if k not in ["data_version", "feature_set"]}
},
"training_data": {
"version": run.data.params.get("data_version"),
"size": run.data.params.get("train_size"),
"positive_rate": run.data.params.get("positive_rate"),
"date": run.data.params.get("training_date")
},
"performance": {
"auc_roc": run.data.metrics.get("auc_roc"),
"sensitivity": run.data.metrics.get("sensitivity"),
"specificity": run.data.metrics.get("specificity"),
"ppv": run.data.metrics.get("ppv"),
},
"approvals": {
"clinical_review": mv.tags.get("clinical_review_status"),
"reviewer": mv.tags.get("clinical_reviewer"),
"review_date": mv.tags.get("review_date"),
},
"stage_history": [
{"stage": "None", "timestamp": mv.creation_timestamp},
{"stage": mv.current_stage, "timestamp": mv.last_updated_timestamp}
]
}
return report
Serving Models from MLflow
MLflow provides built-in model serving, but for production healthcare deployments, you will typically export the model and serve it through a custom FastAPI wrapper with health checks, input validation, and audit logging.
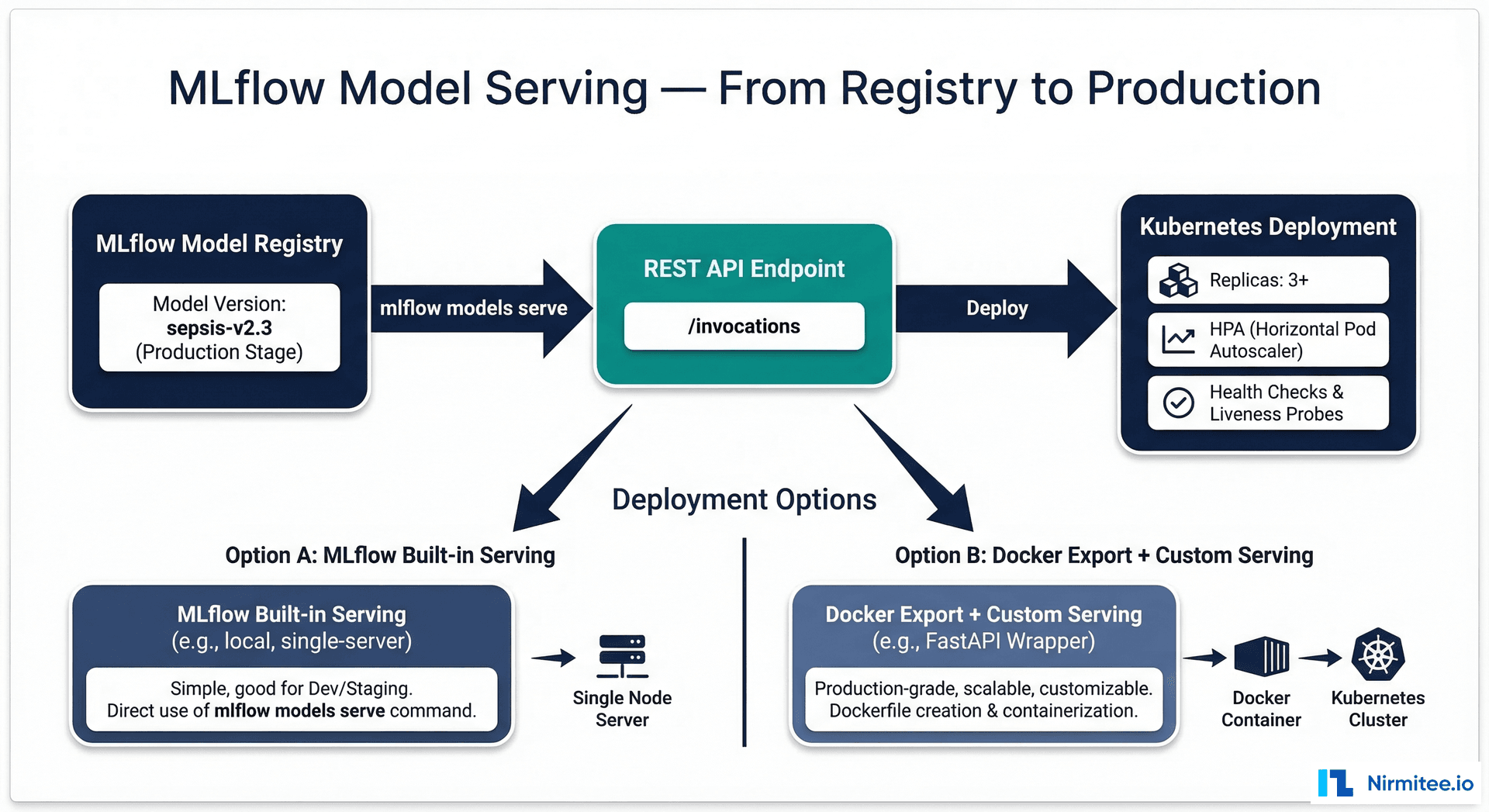
# Production model serving with MLflow + FastAPI
import mlflow.pyfunc
from fastapi import FastAPI, HTTPException
from pydantic import BaseModel, validator
import logging
# Audit logger
audit_logger = logging.getLogger("audit")
audit_handler = logging.FileHandler("/var/log/model-audit.log")
audit_logger.addHandler(audit_handler)
app = FastAPI(title="Sepsis Early Warning - Production")
# Load model from MLflow registry
MODEL_NAME = "sepsis-early-warning"
MODEL_STAGE = "Production"
model = mlflow.pyfunc.load_model(f"models:/{MODEL_NAME}/{MODEL_STAGE}")
class PredictionInput(BaseModel):
heart_rate: float
systolic_bp: float
temperature: float
wbc_count: float = None
lactate: float = None
creatinine: float = None
age: int
encounter_id: str # For audit trail, not for model input
@validator('heart_rate')
def validate_heart_rate(cls, v):
if not 20 <= v <= 300:
raise ValueError('Heart rate out of physiological range')
return v
@app.post("/predict")
def predict(input_data: PredictionInput):
import pandas as pd
# Prepare features (exclude encounter_id - it is metadata, not a feature)
features = pd.DataFrame([{
"heart_rate": input_data.heart_rate,
"sbp": input_data.systolic_bp,
"temperature": input_data.temperature,
"wbc": input_data.wbc_count,
"lactate": input_data.lactate,
"creatinine": input_data.creatinine,
"age": input_data.age,
}])
# Predict
risk_score = float(model.predict(features)[0])
# Audit log (encounter_id for traceability, no patient name/MRN)
audit_logger.info(
f"prediction|encounter={input_data.encounter_id}|"
f"score={risk_score:.4f}|model={MODEL_NAME}|stage={MODEL_STAGE}"
)
return {
"risk_score": round(risk_score, 4),
"risk_level": "HIGH" if risk_score > 0.7 else "MEDIUM" if risk_score > 0.3 else "LOW",
"model_version": MODEL_NAME,
}
Frequently Asked Questions
Is MLflow HIPAA compliant out of the box?
No. MLflow itself is a tool, not a HIPAA-compliant platform. HIPAA compliance depends on how you deploy and configure it. Self-hosting within your HIPAA-covered infrastructure, adding encryption at rest for the PostgreSQL backend and artifact storage, implementing RBAC via a reverse proxy, and training your team never to log PHI as experiment parameters — these configurations make your MLflow deployment HIPAA compliant. The tool provides the capabilities; compliance is an implementation responsibility.
Should we use managed MLflow (Databricks) or self-hosted?
Both are valid for healthcare. Managed MLflow on Databricks provides enterprise features (RBAC, SSO, audit logging) out of the box and operates under a BAA. Self-hosted gives you complete control over data residency, which some health systems require. The decision typically comes down to: (1) Does your BAA with Databricks cover experiment metadata? (2) Does your security team require all ML infrastructure to remain on-premises? Start with self-hosted if unsure — you can always migrate to managed later.
How does MLflow compare to Weights and Biases (W&B)?
W&B offers a more polished UI and better visualization capabilities. MLflow offers a more complete lifecycle tool (registry, projects, serving) and is fully open source with no vendor lock-in. For healthcare, MLflow's self-hosting capability is often the deciding factor — W&B's cloud offering requires sending experiment data to their servers, which complicates HIPAA compliance. W&B does offer a self-hosted option (W&B Server) but it requires a commercial license.
Can MLflow handle multi-tenant access for different clinical departments?
MLflow supports multiple experiments and registered models, which can be organized by department. However, native multi-tenancy with access isolation requires either Databricks Managed MLflow (workspace-level isolation) or a custom RBAC layer via reverse proxy that maps user groups to experiment and model permissions. For organizations with strict departmental data boundaries, consider running separate MLflow instances per department.
How do we migrate existing experiments from notebooks to MLflow?
Start with your next experiment — do not try to backfill historical work. Set up the MLflow tracking server, add import mlflow and tracking calls to your training scripts, and begin logging from your next training run forward. Historical results can be manually logged via the MLflow API if needed for audit purposes, but the priority is capturing future work systematically. Most teams are fully integrated within 1-2 sprint cycles.