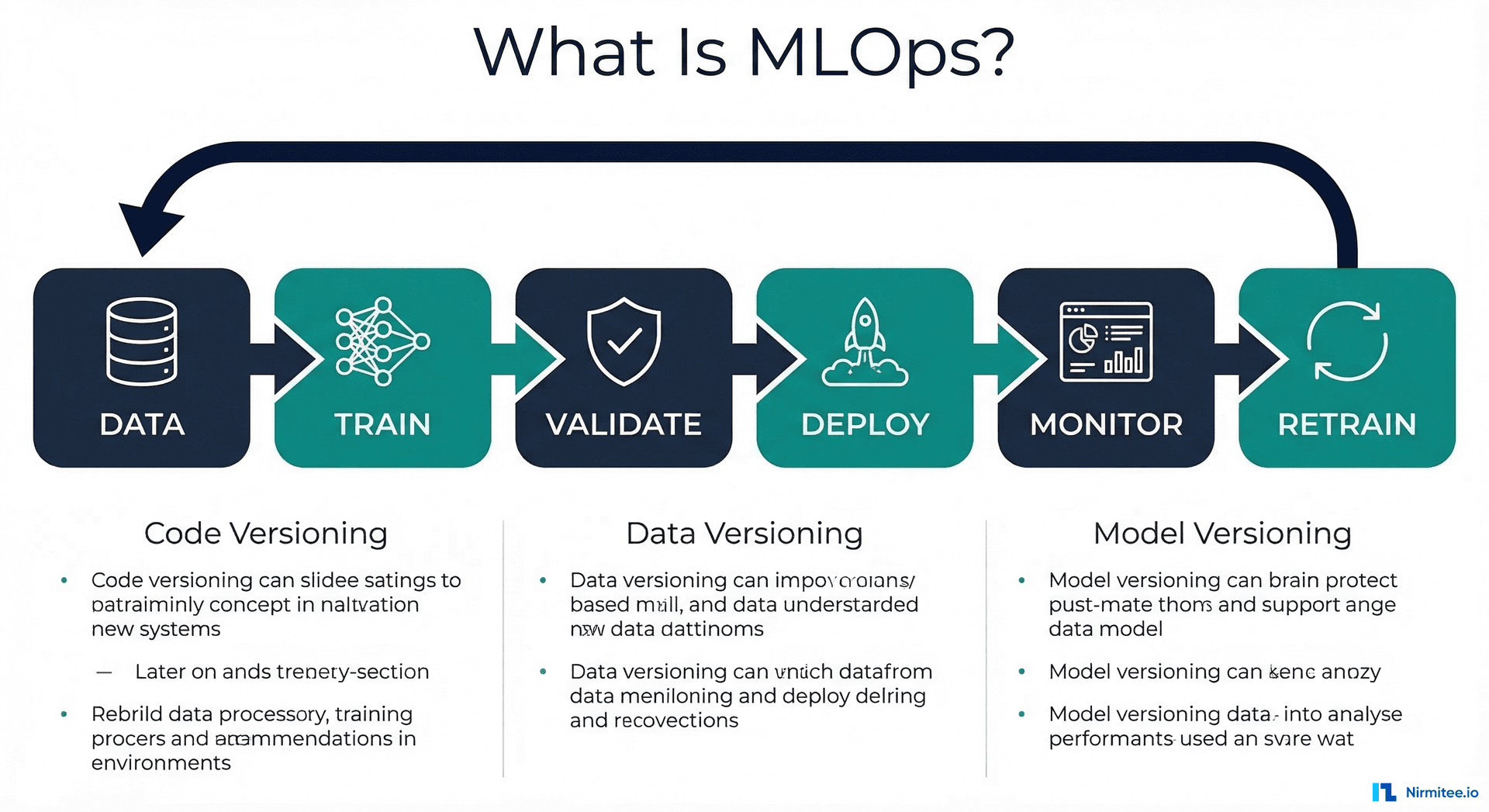
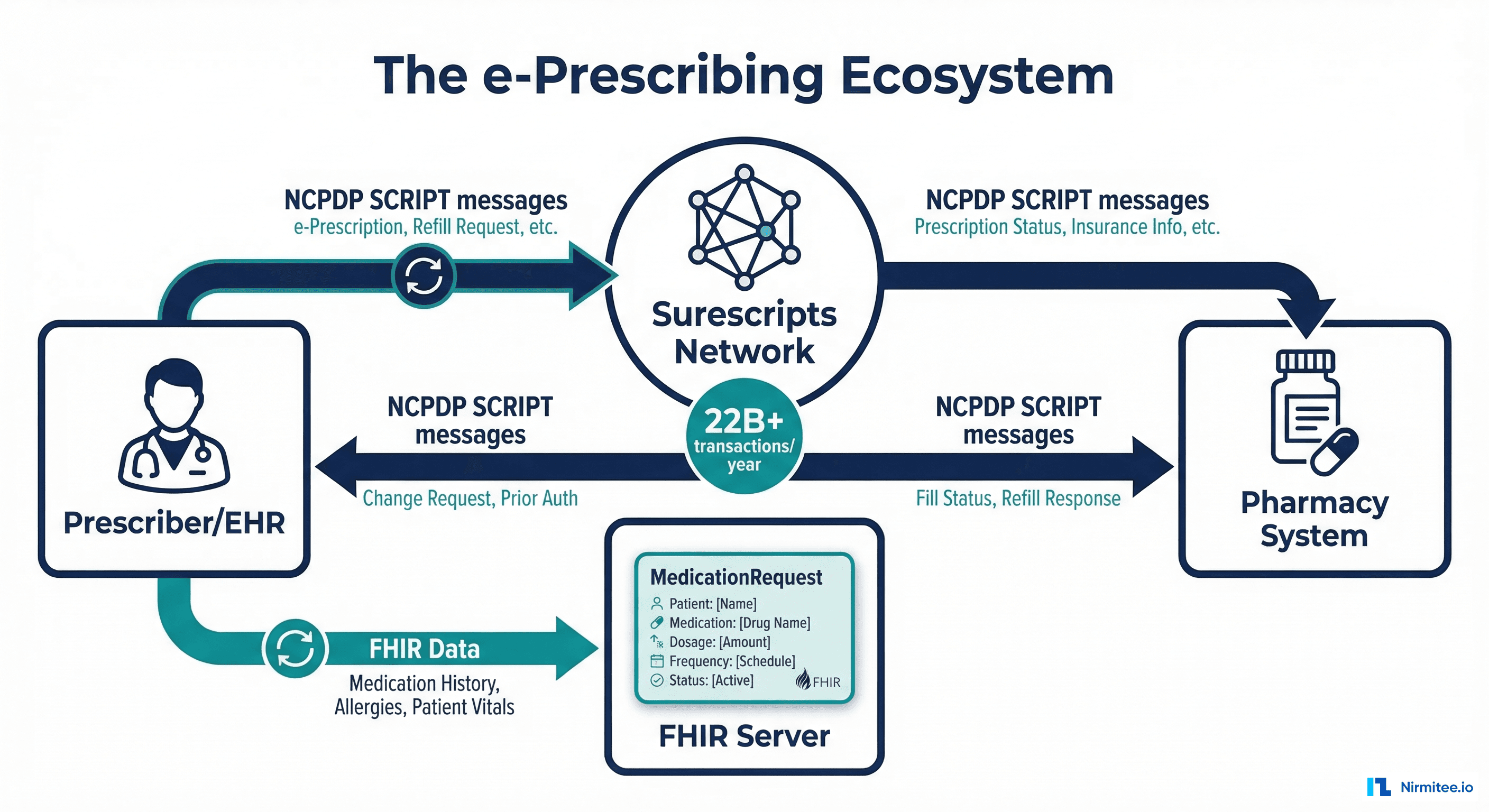
Machine learning models in healthcare face a unique paradox. They are built to save lives, accelerate diagnoses, and reduce clinician burnout -- but the majority of them never make it to production, and those that do often degrade silently within months. A 2025 study by Gartner found that only 53% of ML projects move from prototype to production, and in healthcare that number drops below 40% due to regulatory, safety, and data complexity barriers.
MLOps -- the discipline of operationalizing machine learning -- solves this gap. But healthcare MLOps is fundamentally different from MLOps in e-commerce, fintech, or ad-tech. The stakes are not "lower click-through rate" but "missed sepsis diagnosis." The data is not user behavior logs but protected health information governed by HIPAA. The deployment target is not an API endpoint serving ads but a clinical decision support system integrated into an EHR workflow.
This guide covers the complete MLOps lifecycle for healthcare, from FHIR data extraction through production monitoring, with architecture diagrams, tool comparisons, and implementation checklists. Whether you are deploying your first clinical AI model or scaling an existing ML platform, this is the reference your engineering team needs.
Why Healthcare MLOps Is Different

Before diving into the lifecycle, you need to understand the four forces that make healthcare MLOps categorically different from general-purpose MLOps.
Regulatory Burden
Every clinical AI model is potentially a Software as a Medical Device (SaMD) under FDA oversight. The FDA's AI/ML framework requires documented data lineage, validated training pipelines, bias analysis, and predetermined change control plans. Your MLOps platform must produce regulatory-grade audit trails, not just experiment logs.
Data Sensitivity
Training data is PHI. Every pipeline stage -- extraction, preprocessing, feature engineering, model training -- must comply with HIPAA's minimum necessary standard. De-identification is not optional. BAAs must cover every tool in your stack. A data leak is not a PR problem; it is a federal violation carrying penalties up to $1.5 million per incident category.
Distribution Shift Is Constant
Healthcare data shifts constantly. Seasonal disease patterns change the prevalence of conditions your model predicts. EHR system upgrades alter how data is recorded. New treatment protocols change outcomes your model was trained on. COVID-19 demonstrated this dramatically: sepsis prediction models trained on pre-pandemic ICU data degraded by 15-30% in accuracy within months of the pandemic onset.
Organizational Complexity
Healthcare ML involves clinicians, data scientists, ML engineers, compliance officers, and IT security -- all with different priorities. Research by McKinsey estimates that 70% of production ML failures in healthcare are organizational, not technical. Your MLOps platform must bridge these stakeholder groups with shared visibility into model performance, data quality, and compliance status.
The Healthcare MLOps Lifecycle: Eight Stages
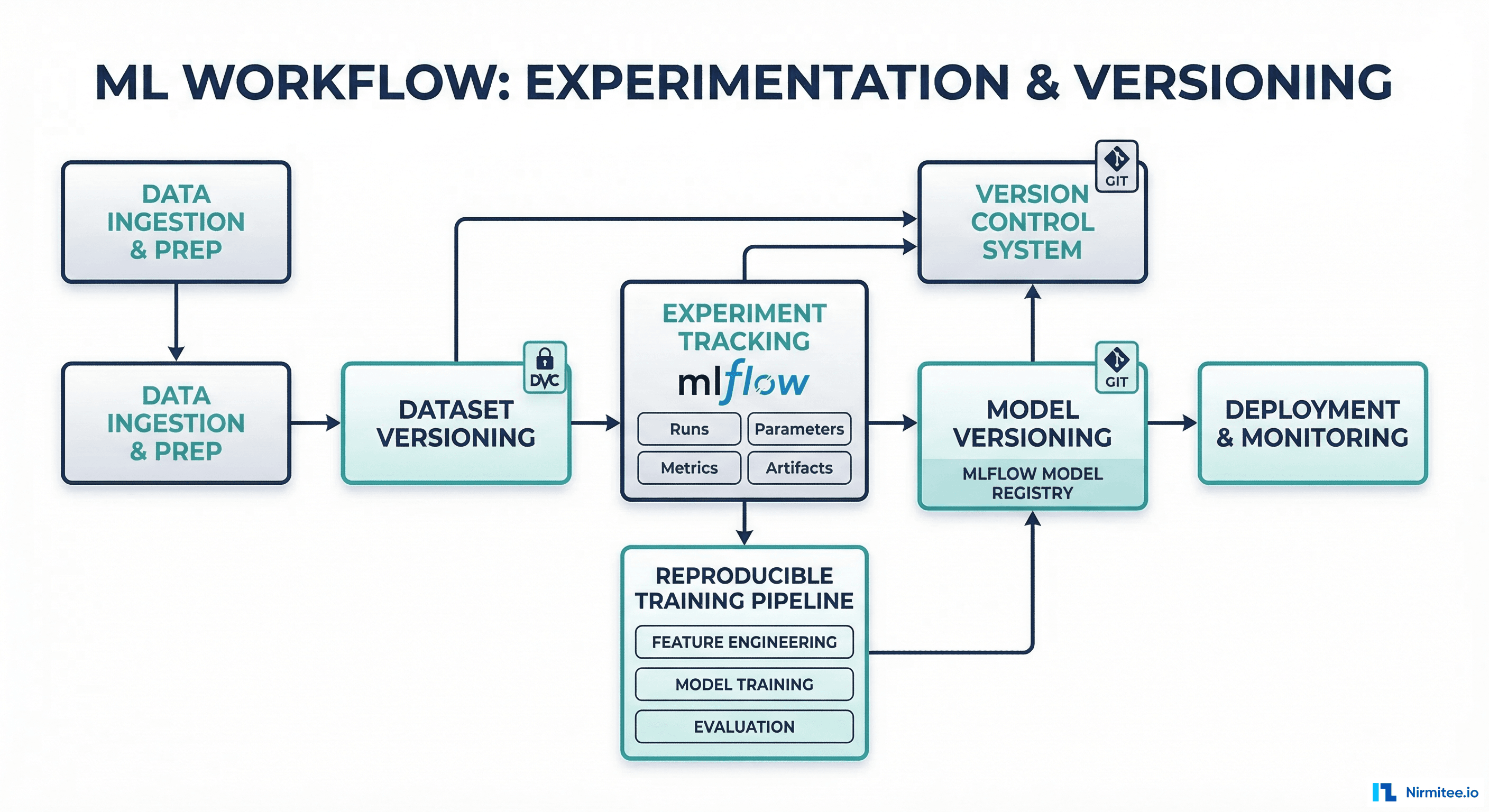
The healthcare MLOps lifecycle has eight stages. Each stage has healthcare-specific requirements that general MLOps frameworks underestimate or ignore entirely.
Stage 1: Data Pipeline -- FHIR Extraction and De-Identification
Clinical data lives in EHR systems, typically accessible via FHIR R4 APIs or bulk data exports. The pipeline starts with FHIR Bulk $export, which produces NDJSON files containing Patient, Observation, Condition, MedicationRequest, and other resources.
import requests
import ndjson
from typing import List, Dict
import hashlib
class FHIRDataExtractor:
def __init__(self, fhir_base_url: str, auth_token: str):
self.base_url = fhir_base_url
self.headers = {
"Authorization": f"Bearer {auth_token}",
"Accept": "application/fhir+json",
"Prefer": "respond-async"
}
def kick_off_bulk_export(self, resource_types: List[str]) -> str:
"""Initiate FHIR Bulk $export and return content-location."""
type_param = ",".join(resource_types)
resp = requests.get(
f"{self.base_url}/$export?_type={type_param}",
headers=self.headers
)
resp.raise_for_status()
return resp.headers["Content-Location"]
def deidentify_patient(self, patient: Dict) -> Dict:
"""Safe Harbor de-identification for patient resources."""
# Hash the patient ID for linkage without exposure
patient["id"] = hashlib.sha256(
patient["id"].encode()
).hexdigest()[:16]
# Remove direct identifiers
patient.pop("name", None)
patient.pop("telecom", None)
patient.pop("address", None)
# Generalize dates (year only for Safe Harbor)
if "birthDate" in patient:
patient["birthDate"] = patient["birthDate"][:4]
# Remove zip codes beyond first 3 digits
# (Safe Harbor allows first 3 if population > 20,000)
return patient
def extract_features(self, resources: List[Dict]) -> Dict:
"""Convert FHIR resources to ML feature vectors."""
features = {
"age_bucket": None,
"condition_count": 0,
"medication_count": 0,
"lab_values": {},
"encounter_frequency": 0
}
for r in resources:
rtype = r.get("resourceType")
if rtype == "Condition":
features["condition_count"] += 1
elif rtype == "MedicationRequest":
features["medication_count"] += 1
elif rtype == "Observation":
code = r.get("code", {}).get("coding", [{}])[0].get("code")
value = r.get("valueQuantity", {}).get("value")
if code and value:
features["lab_values"][code] = value
return featuresCritical consideration: de-identification must happen before data leaves the HIPAA-covered environment. Never send identifiable PHI to a training pipeline running outside your BAA-covered infrastructure. For organizations building end-to-end ML pipelines, this stage defines the compliance posture for every subsequent step.
Stage 2: Feature Engineering
Clinical features differ fundamentally from typical ML features. Time-series vital signs, ICD-10 code hierarchies, medication interaction networks, and lab value trajectories all require domain-specific engineering. A feature store (Feast, Tecton, or SageMaker Feature Store) is essential for consistency between training and inference.
| Feature Category | Examples | FHIR Source | Engineering Notes |
|---|---|---|---|
| Demographics | Age bucket, sex, race/ethnicity | Patient | Use age buckets (18-29, 30-44, etc.) for de-identification |
| Diagnoses | Active condition count, chronic disease flags | Condition | Map ICD-10 to CCS categories for feature reduction |
| Medications | Polypharmacy count, drug class flags | MedicationRequest | Use ATC classification hierarchy for grouping |
| Lab Trends | Creatinine slope, A1C trajectory | Observation | Compute 30/60/90-day slopes, handle missing values |
| Utilization | ED visits (6mo), readmission history | Encounter | Time-windowed aggregates with encounter type filtering |
| Vitals | BP trend, BMI change, heart rate variability | Observation (vitals) | Resample to regular intervals, impute gaps |
Stage 3: Model Training -- Versioned and Reproducible
Every training run must be fully reproducible. This is not a best practice -- it is a regulatory requirement for clinical AI. MLflow is the most widely adopted experiment tracking tool in healthcare ML, and for good reason: it is open-source, self-hostable (critical for HIPAA), and integrates with every major ML framework.
import mlflow
import mlflow.sklearn
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import cross_val_score
import json
# Configure MLflow for healthcare (self-hosted, encrypted)
mlflow.set_tracking_uri("https://mlflow.internal.hospital.org")
mlflow.set_experiment("sepsis-prediction-v3")
with mlflow.start_run(run_name="gbm-baseline-v3.1") as run:
# Log data version and lineage
mlflow.log_param("data_version", "fhir-export-2026-03-01")
mlflow.log_param("deident_method", "safe_harbor_v2")
mlflow.log_param("feature_store_version", "feast-0.35")
mlflow.log_param("cohort_size", 45000)
mlflow.log_param("positive_rate", 0.08)
# Train model
model = GradientBoostingClassifier(
n_estimators=500,
max_depth=6,
learning_rate=0.05,
subsample=0.8
)
model.fit(X_train, y_train)
# Log clinical metrics (not just accuracy)
mlflow.log_metric("auroc", 0.89)
mlflow.log_metric("sensitivity", 0.82) # Critical for clinical
mlflow.log_metric("specificity", 0.85)
mlflow.log_metric("ppv", 0.34)
mlflow.log_metric("npv", 0.98)
mlflow.log_metric("f1", 0.48)
# Log fairness metrics by demographic group
fairness = compute_fairness_metrics(model, X_test, y_test, demographics)
for group, metrics in fairness.items():
mlflow.log_metric(f"auroc_{group}", metrics["auroc"])
mlflow.log_metric(f"fpr_{group}", metrics["fpr"])
# Log model with signature
mlflow.sklearn.log_model(model, "sepsis-gbm")Stage 4: Validation -- Clinical and Statistical

Healthcare model validation has two layers. Statistical validation (AUROC, sensitivity, specificity, calibration) confirms the model performs technically. Clinical validation confirms the model's predictions are clinically meaningful, actionable, and safe. Both must pass before deployment.
Clinical validation requires a formal review by domain experts -- typically physicians, nurses, and pharmacists who will interact with the model's outputs. This review asks: Does the model's alert threshold match clinical workflow? Are false positives at an acceptable rate for the care setting? Does the model perform equitably across patient subgroups?
from dataclasses import dataclass
from typing import Dict, List
@dataclass
class ClinicalValidationReport:
model_version: str
validation_date: str
cohort_description: str
statistical_metrics: Dict[str, float]
fairness_metrics: Dict[str, Dict[str, float]]
clinical_review: Dict[str, str]
approval_status: str # "approved", "conditional", "rejected"
conditions: List[str] # Conditions for deployment
reviewers: List[str] # Clinical reviewers
def generate_validation_report(model, X_val, y_val, demographics):
report = ClinicalValidationReport(
model_version="sepsis-gbm-v3.1",
validation_date="2026-03-15",
cohort_description="45K adult ICU admissions, 2024-2025",
statistical_metrics={
"auroc": compute_auroc(model, X_val, y_val),
"sensitivity_at_90spec": compute_sens_at_spec(model, X_val, y_val, 0.90),
"calibration_slope": compute_calibration(model, X_val, y_val),
"brier_score": compute_brier(model, X_val, y_val)
},
fairness_metrics=compute_fairness_metrics(model, X_val, y_val, demographics),
clinical_review={
"workflow_fit": "Integrates into nursing early warning score",
"alert_fatigue_risk": "Moderate - recommend 0.7 threshold",
"false_positive_tolerance": "Acceptable at current specificity"
},
approval_status="conditional",
conditions=["Deploy in shadow mode for 30 days before activation"],
reviewers=["Dr. Smith (ICU)", "RN Johnson (Nursing Informatics)"]
)
return reportStage 5: Deployment -- Shadow, Canary, Production

Healthcare AI deployment follows a three-phase rollout that is more conservative than typical software deployment. This is non-negotiable for patient safety.
Shadow Mode (2-4 weeks): The model runs in production on real data but its predictions are logged, not displayed to clinicians. Compare shadow predictions against actual outcomes. This catches data pipeline issues, feature engineering bugs, and performance gaps between validation and production data.
Canary Deployment (2-4 weeks): The model's predictions are shown to a small group of clinicians (typically one unit or department) with explicit labeling as "AI-assisted" and easy feedback mechanisms. Collect clinician feedback on alert quality, timing, and workflow integration.
Production Rollout: Full deployment with continued monitoring. Even at this stage, maintain a kill switch that can disable the model within minutes if issues are detected.
Stage 6: Monitoring -- Drift, Fairness, and Accuracy
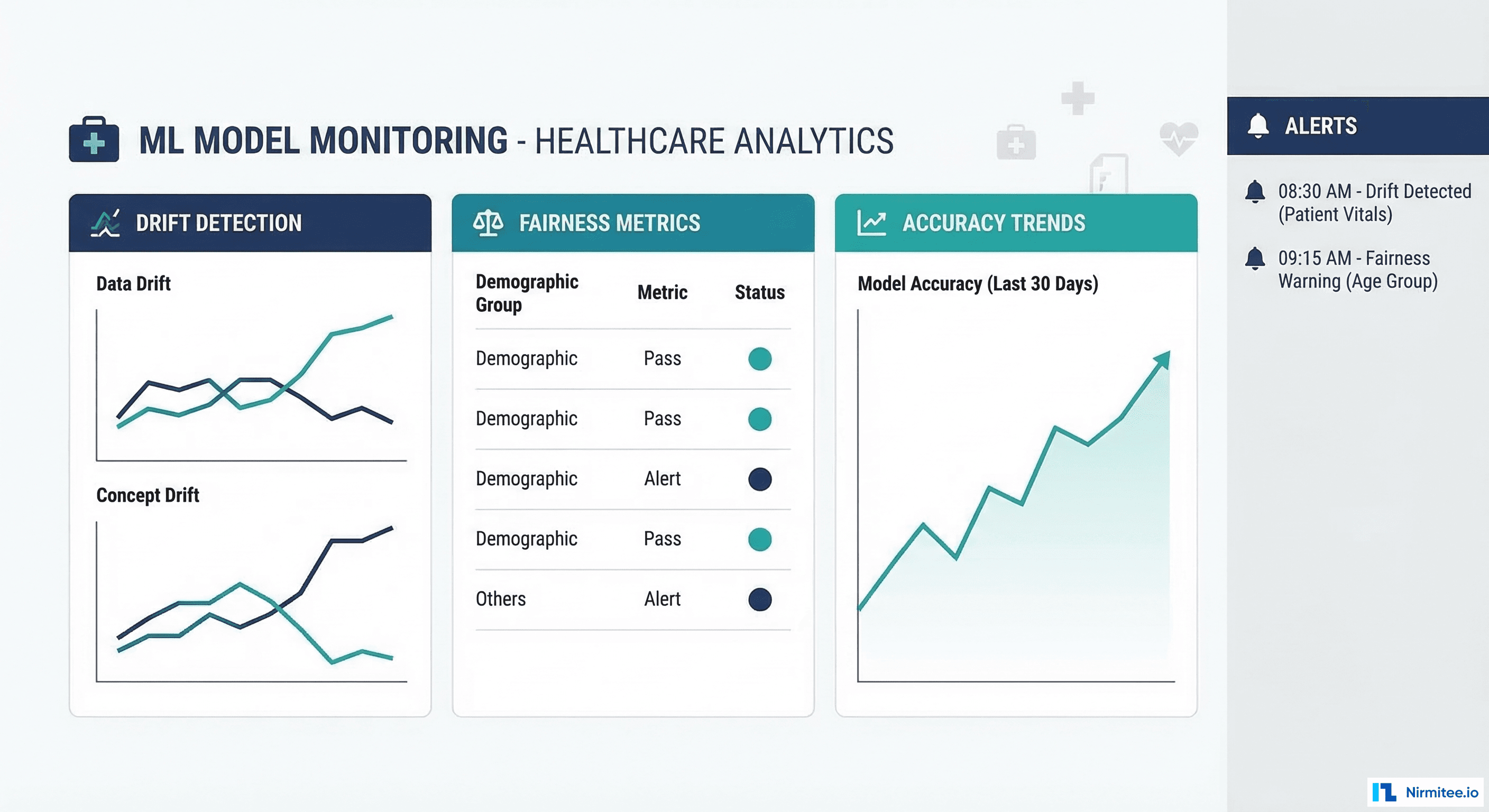
Production monitoring for clinical AI requires three monitoring dimensions running simultaneously. Data drift monitoring uses statistical tests (Population Stability Index, Kolmogorov-Smirnov test, Jensen-Shannon divergence) to detect when incoming data distributions diverge from training data. Fairness monitoring tracks performance metrics disaggregated by demographic groups to ensure the model does not develop disparate performance over time. Accuracy monitoring compares model predictions against confirmed outcomes (which may arrive days or weeks after prediction). For a deep dive on drift specifically, see our guide on detecting model drift in clinical AI.
Stage 7: Retraining Triggers
Automated retraining should be triggered by drift detection thresholds, not by calendar schedules. When the PSI for any critical feature exceeds 0.2, or when rolling AUROC drops below the validated threshold by more than 0.03, the retraining pipeline should activate. Under the FDA's Predetermined Change Control Plan (PCCP) framework, these triggers and the retraining procedure must be documented and approved in advance.
Stage 8: Governance and Audit
Every stage produces artifacts that must be retained for regulatory audit. Data lineage documents, training configurations, validation reports, deployment approvals, monitoring logs, and retraining decisions form a complete audit trail. This is where many teams fail -- they build the ML pipeline but not the governance layer. Tools like MLflow Model Registry, DVC for data versioning, and custom audit logging fill this gap.
Tool Comparison: MLOps Platforms for Healthcare
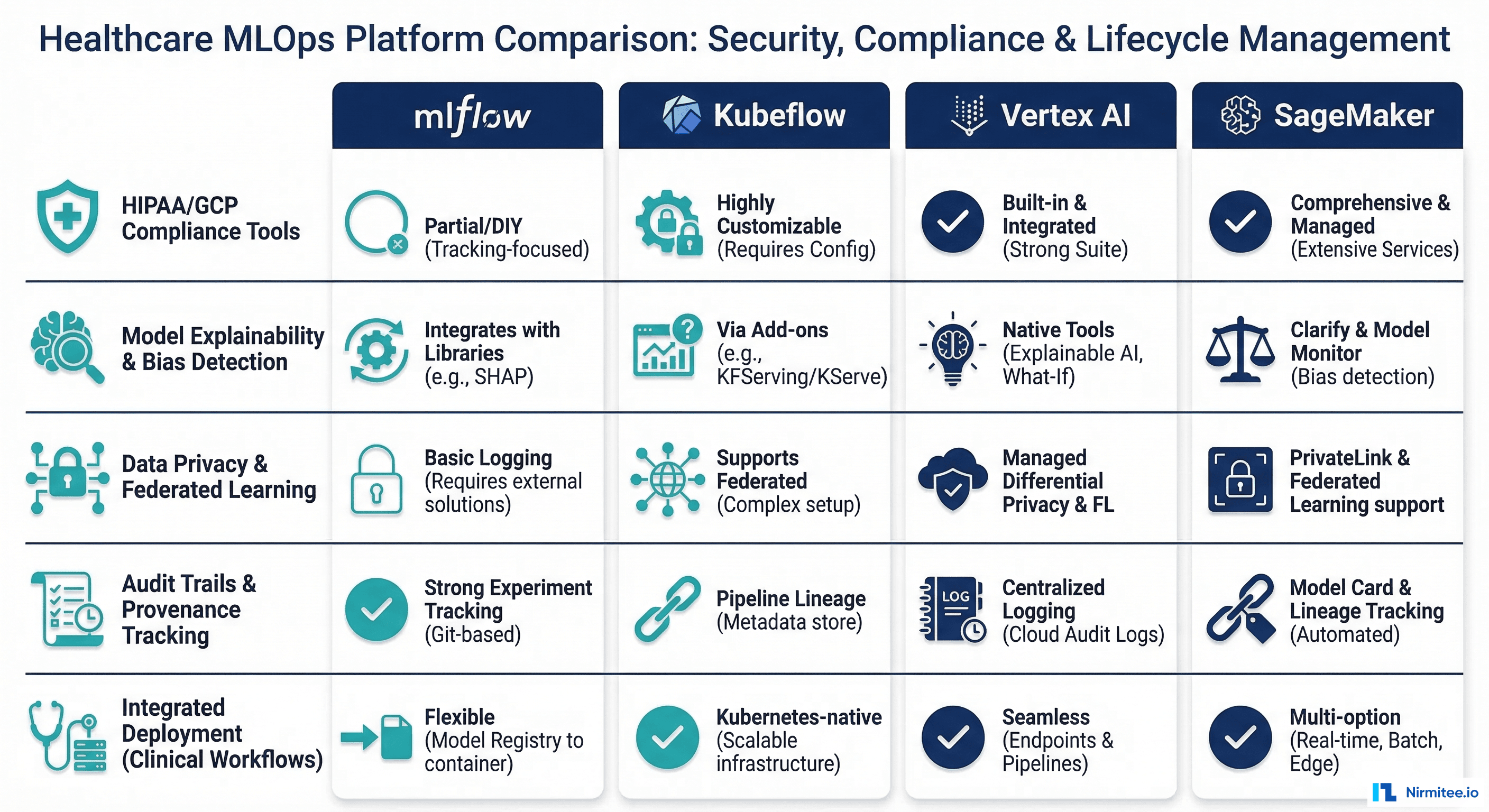
Four platforms dominate healthcare MLOps. Each has distinct strengths and tradeoffs for regulated environments.
| Capability | MLflow | Kubeflow | Vertex AI (GCP) | SageMaker (AWS) |
|---|---|---|---|---|
| Self-Hosted Option | Yes (open-source) | Yes (Kubernetes) | No (GCP only) | No (AWS only) |
| HIPAA BAA Available | Via infra provider | Via infra provider | Yes (GCP BAA) | Yes (AWS BAA) |
| Experiment Tracking | Excellent | Good (via Katib) | Excellent | Excellent |
| Model Registry | Built-in | Limited | Built-in | Built-in |
| Pipeline Orchestration | Limited (needs Airflow) | Excellent (Argo-based) | Excellent | Good (Step Functions) |
| Feature Store | No (use Feast) | No (use Feast) | Built-in | Built-in |
| Model Monitoring | Community plugins | Limited | Built-in | Built-in (Model Monitor) |
| Drift Detection | Via Evidently/WhyLabs | Manual setup | Built-in | Built-in |
| Cost | Free (infra costs) | Free (infra costs) | Per-use pricing | Per-use pricing |
| Best For | Teams wanting full control | Kubernetes-native orgs | GCP-committed orgs | AWS-committed orgs |
Recommendation for healthcare: For organizations that need maximum control over data residency and compliance, MLflow + Kubeflow on self-managed Kubernetes provides the most flexibility. For teams that want managed infrastructure with less operational overhead, Vertex AI and SageMaker both offer HIPAA-eligible environments with comprehensive MLOps tooling. The deciding factor is usually which cloud provider your organization already has a BAA with.
The 70% Problem: Why Most Failures Are Organizational

The technical pipeline described above solves 30% of the problem. The remaining 70% is organizational. Here are the most common organizational failure modes in healthcare MLOps and how to address them.
Failure Mode 1: No Clinical Champion
Every clinical AI project needs a physician or nurse champion who understands both the clinical problem and the model's capabilities. Without this person, data scientists build models that are technically sound but clinically irrelevant -- predicting outcomes that clinicians do not act on, alerting at times when intervention is impossible, or using features that are only available retrospectively.
Failure Mode 2: Validation Theater
Some organizations perform statistical validation (AUROC, sensitivity) but skip clinical validation. The model looks good on paper but fails in practice because it does not fit the clinical workflow. A sepsis alert that fires 6 hours after the physician has already started antibiotics is technically correct but clinically useless.
Failure Mode 3: No Feedback Loop
Clinicians interact with model outputs daily but have no mechanism to report errors, false positives, or missed predictions. Without this feedback, you cannot improve the model and you cannot detect degradation. Build structured feedback collection into the clinical interface from day one.
Failure Mode 4: Compliance as Afterthought
Teams that build the ML pipeline first and add compliance documentation later discover that their pipeline cannot produce the audit artifacts regulators require. Compliance requirements should define pipeline architecture, not be retrofitted onto it. See our guide on FDA SaMD compliance for engineers for what regulators actually require.
Implementation Checklist
Use this checklist to evaluate your organization's readiness for healthcare MLOps. Each item represents a prerequisite that, if missing, will cause pipeline failures in production.
| Category | Checklist Item | Priority |
|---|---|---|
| Data | FHIR R4 API access with Bulk $export capability | Critical |
| Data | Automated de-identification pipeline (Safe Harbor or Expert Determination) | Critical |
| Data | Data versioning system (DVC, LakeFS, or cloud-native) | High |
| Infrastructure | HIPAA-eligible compute environment (on-prem or cloud with BAA) | Critical |
| Infrastructure | Container orchestration (Kubernetes) for pipeline execution | High |
| Infrastructure | Encrypted storage for training data and model artifacts | Critical |
| Tooling | Experiment tracking (MLflow or equivalent) | Critical |
| Tooling | Feature store for training/serving consistency | High |
| Tooling | Model registry with approval workflows | High |
| Tooling | Drift detection framework (Evidently, NannyML, or built-in) | High |
| Governance | Clinical validation protocol with defined reviewers | Critical |
| Governance | Model card template for documentation | High |
| Governance | Incident response plan for model failures | Critical |
| Organization | Identified clinical champion for each model | Critical |
| Organization | Cross-functional team (DS + MLE + clinician + compliance) | Critical |
| Organization | Clinician feedback mechanism integrated into EHR | High |
Frequently Asked Questions
How long does it take to build a healthcare MLOps pipeline from scratch?
For a well-resourced team (2-3 ML engineers, 1 data engineer, clinical informatics support), expect 4-6 months to build a production-grade pipeline for a single model. This includes FHIR data extraction, de-identification, feature engineering, training infrastructure, validation framework, and monitoring. The first model takes the longest; subsequent models leveraging the same infrastructure can be deployed in 4-8 weeks.
Can we use open-source tools for HIPAA-compliant MLOps?
Yes, but the tools themselves are not what makes you HIPAA-compliant -- the infrastructure they run on is. MLflow, Kubeflow, Evidently, and Feast are all open-source tools widely used in healthcare. They must run on HIPAA-eligible infrastructure (on-premises or cloud with BAA), with encryption at rest and in transit, access controls, and audit logging. The tool choice does not determine compliance; the deployment architecture does.
What metrics should we track for clinical AI in production?
At minimum: (1) discrimination metrics (AUROC, AUPRC) computed on rolling windows, (2) calibration (predicted probabilities vs observed rates), (3) alert rate (alerts per patient-day), (4) clinician response rate (what percentage of alerts lead to clinical action), (5) fairness metrics disaggregated by race, sex, age, and insurance status, (6) data drift scores for top features. Track all metrics weekly and set automated thresholds for escalation.
How do we handle model retraining under FDA oversight?
The FDA's Predetermined Change Control Plan (PCCP) framework, finalized in late 2024, allows manufacturers to document planned model updates in advance. If your retraining procedure, triggers, performance thresholds, and validation protocol are described in an approved PCCP, you can retrain without a new regulatory submission. Changes outside the PCCP require a new 510(k) or De Novo submission. This makes PCCP documentation one of the most important artifacts in your MLOps pipeline.
Should we build or buy our MLOps platform?
For most healthcare organizations, a hybrid approach works best. Use managed cloud services (SageMaker, Vertex AI) for compute and basic pipeline orchestration, but build custom components for healthcare-specific needs: FHIR data extraction, de-identification, clinical validation workflows, and regulatory audit trails. No off-the-shelf MLOps platform fully addresses healthcare's unique requirements, but building everything from scratch is unnecessarily expensive.
What is the biggest mistake teams make with healthcare MLOps?
Treating it as a purely technical initiative. The most common failure pattern is a data science team building an excellent model with robust infrastructure that never gets adopted because clinicians were not involved in defining the problem, validating the outputs, or designing the workflow integration. Start with the clinical workflow, work backward to the model, and build the pipeline to serve that workflow.