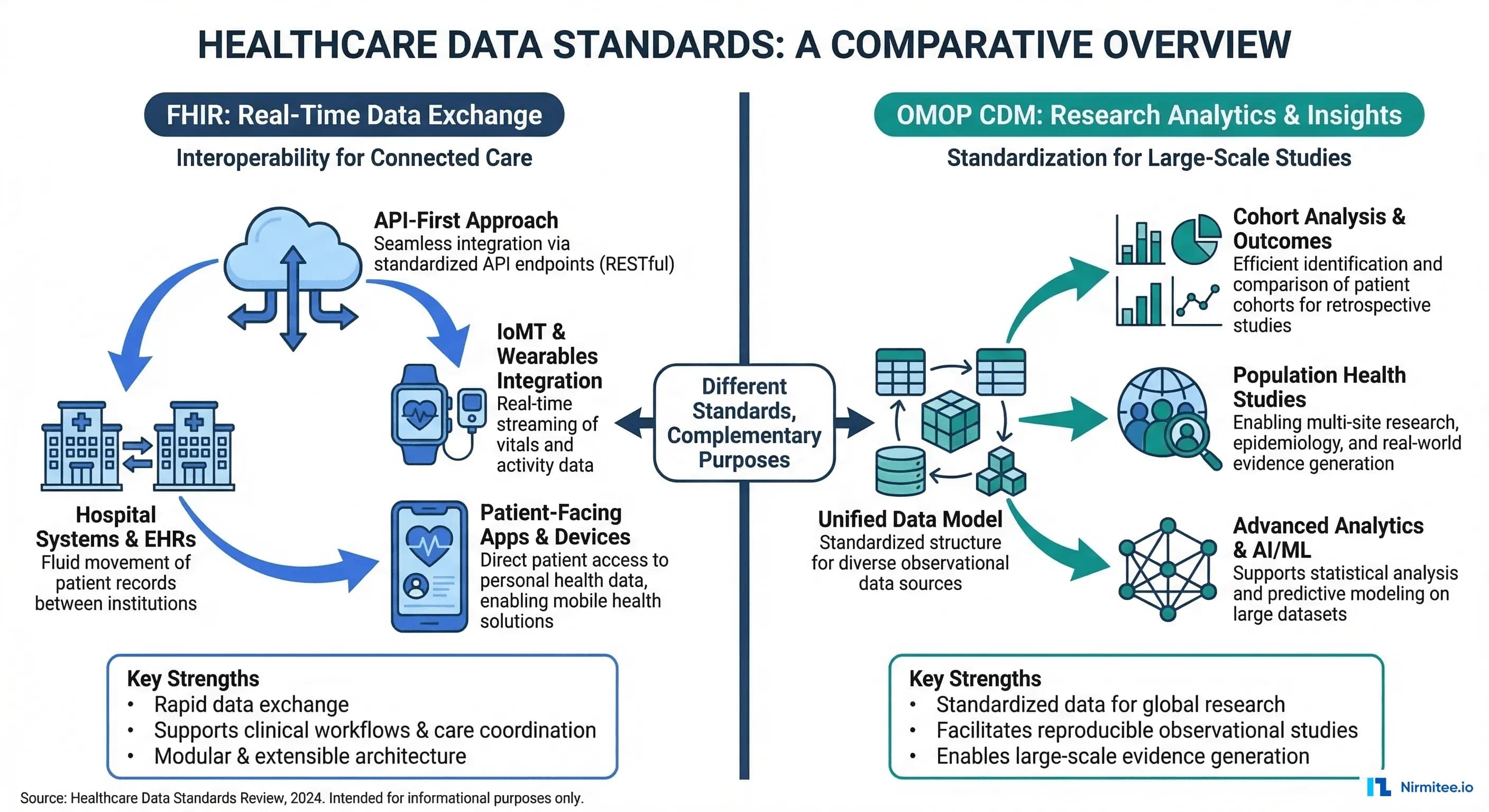
OMOP CDM, maintained by the OHDSI (Observational Health Data Sciences and Informatics) community, is the research and analytics standard. It powers drug safety studies, clinical trial feasibility, population health analysis, and real-world evidence generation across a federated network of 810+ data sources representing 2.5 billion patient records in 30+ countries.
This article provides a detailed, practical guide to both standards — their architectures, their differences, how to map between them, and the ETL pipeline that bridges clinical operations with research analytics. If you are building a healthcare data platform, you need both.
FHIR: The Interoperability Standard for Real-Time Exchange
What FHIR Does
FHIR defines a resource-based data model with RESTful APIs for exchanging clinical data between systems in real time. Each clinical concept — a patient, a diagnosis, a medication — is represented as a FHIR Resource in JSON or XML format.
Key characteristics of FHIR:
- Resource-based model: 150+ resource types (Patient, Condition, Observation, MedicationRequest, Encounter, etc.)
- RESTful API: Standard CRUD operations plus search, batch, and transaction support
- Multiple vocabularies: SNOMED CT, LOINC, ICD-10, RxNorm, CPT — each resource can use any coded system
- Real-time focus: Designed for synchronous API calls, webhooks (FHIR Subscriptions), and event-driven workflows
- Extensibility: Profiles and extensions allow customization without breaking the base spec
FHIR Resource Example: Condition
{
"resourceType": "Condition",
"id": "example-diabetes",
"clinicalStatus": {
"coding": [{
"system": "http://terminology.hl7.org/CodeSystem/condition-clinical",
"code": "active"
}]
},
"verificationStatus": {
"coding": [{
"system": "http://terminology.hl7.org/CodeSystem/condition-ver-status",
"code": "confirmed"
}]
},
"category": [{
"coding": [{
"system": "http://terminology.hl7.org/CodeSystem/condition-category",
"code": "encounter-diagnosis"
}]
}],
"code": {
"coding": [{
"system": "http://snomed.info/sct",
"code": "44054006",
"display": "Type 2 diabetes mellitus"
}, {
"system": "http://hl7.org/fhir/sid/icd-10-cm",
"code": "E11.9",
"display": "Type 2 diabetes mellitus without complications"
}]
},
"subject": {
"reference": "Patient/12345"
},
"onsetDateTime": "2023-06-15",
"recordedDate": "2023-06-15"
}Notice how a single Condition can carry multiple coding systems (SNOMED + ICD-10) simultaneously. This flexibility is powerful for exchange but creates complexity for analytics, which OMOP solves.
When to Use FHIR
- EHR integrations: Pulling/pushing data to Epic, Oracle Health (Cerner), MEDITECH via standardized APIs
- Patient-facing apps: Patient Access APIs per CMS requirements
- Clinical Decision Support: CDS Hooks that fire in real time during clinical workflows
- Data exchange: Payer-provider data exchange, TEFCA, care coordination
- Regulatory compliance: ONC Cures Act, CMS Interoperability Rule, USCDI requirements
OMOP CDM: The Research Standard for Population Analytics
What OMOP CDM Does
OMOP CDM (currently v5.4) defines a relational database schema optimized for large-scale observational research. Unlike FHIR's resource-based approach, OMOP uses a traditional star schema with standardized vocabulary tables that normalize all clinical codes into a single concept hierarchy.
Key characteristics of OMOP CDM:
- Relational model: 39 tables organized into clinical data, health system data, health economics data, and vocabulary tables
- Standardized vocabulary: All source codes (ICD-10, SNOMED, LOINC, RxNorm) mapped to standard OMOP Concept IDs
- Retrospective focus: Designed for querying historical data across populations, not real-time API calls
- SQL-native: Query with standard SQL — no custom query languages or API pagination
- Federated network: Same schema across 810+ institutions enables multi-site studies without sharing patient data
The OMOP Vocabulary System
OMOP's most powerful feature is its vocabulary layer. Every clinical concept from any source system gets mapped to a standard concept. This means "Type 2 diabetes" coded as ICD-10 E11.9, SNOMED 44054006, or a proprietary EHR code all map to the same OMOP standard concept ID (201826). When you query condition_concept_id = 201826, you get every diabetic patient regardless of which coding system their source EHR used.
-- OMOP vocabulary mapping example
-- Find all standard concept IDs for Type 2 Diabetes
SELECT
c.concept_id,
c.concept_name,
c.vocabulary_id,
c.concept_code,
c.standard_concept
FROM concept c
WHERE c.concept_id IN (
-- The standard concept for T2DM
SELECT descendant_concept_id
FROM concept_ancestor
WHERE ancestor_concept_id = 201826 -- Type 2 diabetes mellitus
)
AND c.standard_concept = 'S';
-- This returns ALL descendants: T2DM with nephropathy,
-- T2DM with neuropathy, etc. -- regardless of source coding systemFHIR vs OMOP: Head-to-Head Comparison
| Dimension | FHIR R4/R5 | OMOP CDM v5.4 |
|---|---|---|
| Primary Purpose | Real-time data exchange between systems | Retrospective research and analytics |
| Data Model | Resource-based (JSON/XML documents) | Relational (star schema, SQL tables) |
| Vocabulary Handling | Multiple systems coexist (SNOMED + ICD + LOINC in same resource) | All mapped to standard OMOP concepts |
| Query Language | REST API with search parameters | Standard SQL |
| Temporal Focus | Current state, real-time events | Historical longitudinal data |
| Network Scale | HL7 community, EHR vendor ecosystems | OHDSI: 810+ data sources, 30+ countries |
| Data Quality | Varies by implementation; no built-in quality framework | Achilles + DQD (Data Quality Dashboard) built in |
| Strengths | Interoperability, app ecosystem, regulatory compliance | Large-scale analytics, federated research, standardized vocabulary |
| Governance | HL7 International (standards body) | OHDSI (open science community) |
| Typical Users | App developers, integration engineers, EHR vendors | Epidemiologists, clinical researchers, data scientists |
The FHIR-to-OMOP Resource Mapping
Bridging FHIR and OMOP requires mapping FHIR resources to OMOP tables. This is not a simple 1:1 mapping — FHIR's nested, document-style resources must be flattened into OMOP's relational tables, and all coded values must be translated through the OMOP vocabulary layer.
Core Mapping Table
| FHIR Resource | OMOP CDM Table | Key Transformations |
|---|---|---|
| Patient | person | gender → gender_concept_id, birthDate → year/month/day_of_birth, race/ethnicity extensions → concept_ids |
| Condition | condition_occurrence | code.coding → condition_concept_id (via vocabulary mapping), onsetDateTime → condition_start_date |
| MedicationRequest | drug_exposure | medication → drug_concept_id (RxNorm → OMOP), dosageInstruction → quantity, daysSupply |
| Observation (labs) | measurement | code → measurement_concept_id, valueQuantity → value_as_number, unit → unit_concept_id |
| Observation (vitals) | measurement | Same as labs; component observations (BP systolic/diastolic) expand to multiple measurement rows |
| Observation (social) | observation | Smoking status, alcohol use → observation_concept_id |
| Procedure | procedure_occurrence | code → procedure_concept_id, performedDateTime → procedure_date |
| Encounter | visit_occurrence | class → visit_concept_id (inpatient/outpatient/ED), period → visit_start/end_date |
| AllergyIntolerance | observation | code → observation_concept_id, mapped as allergy observations |
| Immunization | drug_exposure | vaccineCode → drug_concept_id, occurrenceDateTime → drug_exposure_start_date |
| DiagnosticReport | measurement (results linked) | Results reference Observations, which map to measurements; report metadata may go to note table |
| Claim/Explanation of Benefit | cost, payer_plan_period | Payment amounts → cost table, coverage period → payer_plan_period |
Vocabulary Mapping: The Critical Step
The most complex part of the FHIR-to-OMOP transformation is vocabulary mapping. FHIR resources carry source codes (ICD-10, SNOMED, LOINC), but OMOP requires mapping each to a standard concept. Here is how the mapping chain works:
# Python: FHIR code to OMOP concept mapping
import psycopg2
def map_fhir_code_to_omop(fhir_system, fhir_code, conn):
"""Map a FHIR coding system + code to OMOP standard concept."""
FHIR_TO_OMOP_VOCAB = {
"http://snomed.info/sct": "SNOMED",
"http://hl7.org/fhir/sid/icd-10-cm": "ICD10CM",
"http://hl7.org/fhir/sid/icd-10": "ICD10",
"http://loinc.org": "LOINC",
"http://www.nlm.nih.gov/research/umls/rxnorm": "RxNorm",
"http://www.ama-assn.org/go/cpt": "CPT4",
}
omop_vocab = FHIR_TO_OMOP_VOCAB.get(fhir_system)
if not omop_vocab:
return {"concept_id": 0, "concept_name": "No matching concept"}
cursor = conn.cursor()
# Step 1: Find the source concept
cursor.execute(
"SELECT concept_id, concept_name, standard_concept "
"FROM concept "
"WHERE vocabulary_id = %s AND concept_code = %s "
"AND invalid_reason IS NULL",
(omop_vocab, fhir_code)
)
source = cursor.fetchone()
if not source:
return {"concept_id": 0, "concept_name": "Source code not found"}
source_id, source_name, is_standard = source
# Step 2: If already standard, return it
if is_standard == 'S':
return {"concept_id": source_id, "concept_name": source_name}
# Step 3: Map to standard concept via concept_relationship
cursor.execute(
"SELECT c.concept_id, c.concept_name "
"FROM concept_relationship cr "
"JOIN concept c ON c.concept_id = cr.concept_id_2 "
"WHERE cr.concept_id_1 = %s "
"AND cr.relationship_id = 'Maps to' "
"AND cr.invalid_reason IS NULL "
"AND c.standard_concept = 'S'",
(source_id,)
)
standard = cursor.fetchone()
if standard:
return {"concept_id": standard[0], "concept_name": standard[1]}
return {"concept_id": 0, "concept_name": f"No standard mapping for {source_name}"}
# Example usage:
# ICD-10 E11.9 (Type 2 DM) -> OMOP standard concept 201826
# result = map_fhir_code_to_omop(
# "http://hl7.org/fhir/sid/icd-10-cm", "E11.9", conn
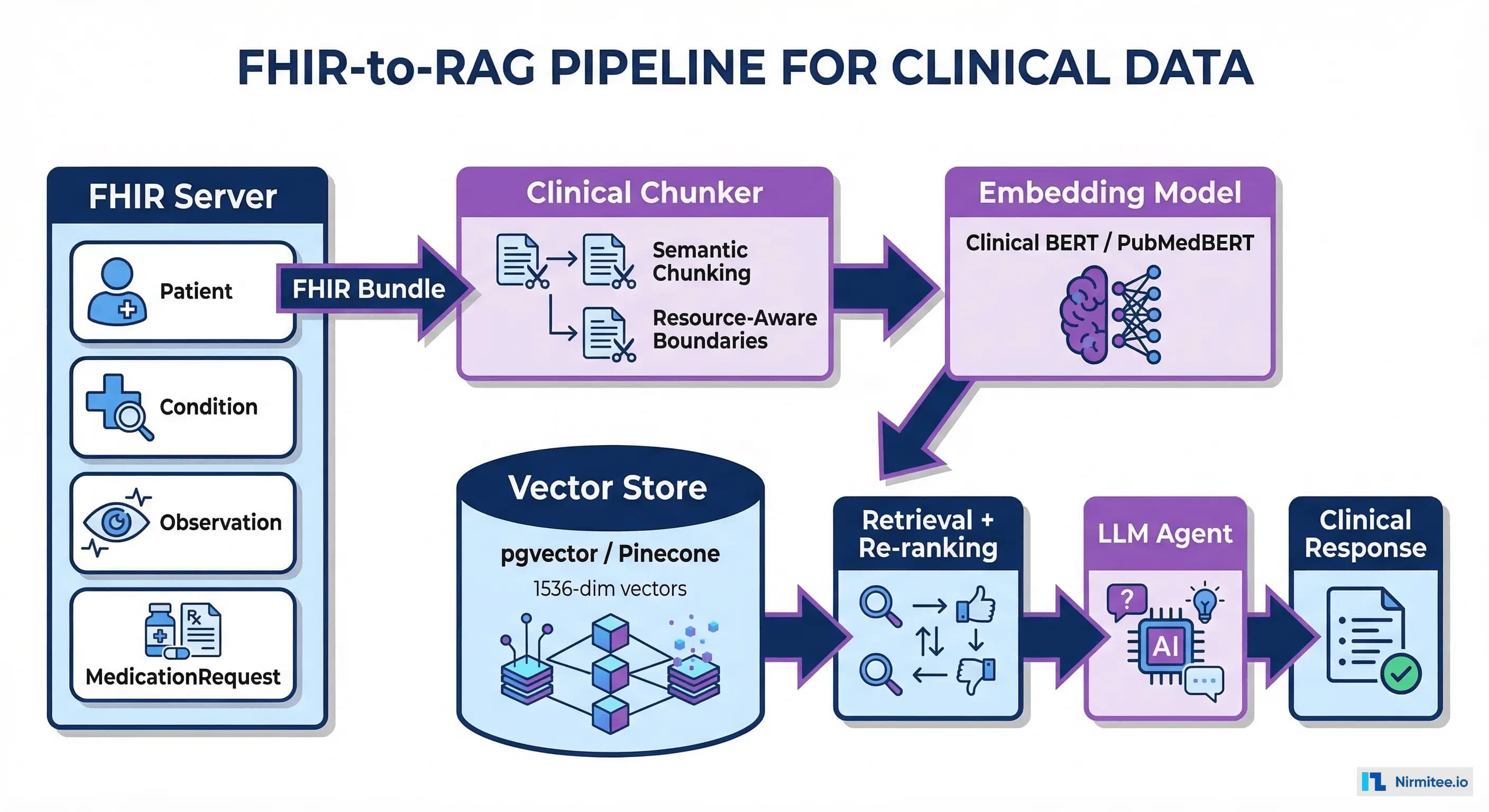
# )Building the FHIR-to-OMOP ETL Pipeline
Architecture Overview
The production architecture for bridging FHIR and OMOP follows a three-layer approach:
- Extract: Pull FHIR data via Bulk Data Export ($export), individual REST calls, or FHIR Subscriptions for real-time updates
- Transform: Flatten FHIR JSON resources, map vocabularies to OMOP standard concepts, apply data quality rules
- Load: Insert into OMOP CDM tables in PostgreSQL, SQL Server, or cloud data warehouses
ETL Implementation: FHIR Condition to OMOP condition_occurrence
# FHIR-to-OMOP ETL: Transform FHIR Condition resources
# to OMOP CDM condition_occurrence table.
# Production pipeline using Apache Spark for scale.
from pyspark.sql import SparkSession
from pyspark.sql.functions import (
col, explode, when, lit, to_date, coalesce
)
from pyspark.sql.types import LongType
spark = SparkSession.builder \
.appName("FHIR-to-OMOP-Conditions") \
.config("spark.sql.extensions",
"io.delta.sql.DeltaSparkSessionExtension") \
.getOrCreate()
# Load FHIR Condition NDJSON (from Bulk Export)
fhir_conditions = spark.read.json(
"s3://ehr-data-lake/bronze/fhir/Condition/*.ndjson"
)
# Load OMOP vocabulary tables (pre-loaded)
concept = spark.read.parquet("s3://omop-vocab/concept/")
concept_relationship = spark.read.parquet(
"s3://omop-vocab/concept_relationship/"
)
person = spark.read.parquet("s3://omop-cdm/person/")
# Step 1: Extract structured fields from FHIR JSON
conditions_flat = fhir_conditions.select(
col("id").alias("fhir_condition_id"),
col("subject.reference").alias("patient_ref"),
col("code.coding").alias("codings"),
col("onsetDateTime").alias("onset_date"),
col("abatementDateTime").alias("abatement_date"),
col("recordedDate").alias("recorded_date"),
col("clinicalStatus.coding")[0]["code"].alias("clinical_status"),
col("verificationStatus.coding")[0]["code"]
.alias("verification_status"),
col("category")[0]["coding"][0]["code"].alias("category_code")
)
# Step 2: Explode codings, pick preferred vocabulary
# Priority: SNOMED > ICD-10 > ICD-9
conditions_coded = conditions_flat \
.withColumn("coding", explode("codings")) \
.withColumn("code_system", col("coding.system")) \
.withColumn("code_value", col("coding.code")) \
.withColumn("vocab_priority",
when(col("code_system") == "http://snomed.info/sct", 1)
.when(col("code_system").contains("icd-10"), 2)
.when(col("code_system").contains("icd-9"), 3)
.otherwise(99)
)
# Step 3: Map to OMOP standard concepts
conditions_mapped = conditions_coded \
.join(
concept.alias("src"),
(col("code_value") == col("src.concept_code")) &
(col("src.vocabulary_id").isin("SNOMED","ICD10CM","ICD9CM")),
"left"
) \
.join(
concept_relationship.alias("cr"),
(col("src.concept_id") == col("cr.concept_id_1")) &
(col("cr.relationship_id") == "Maps to"),
"left"
) \
.join(
concept.alias("std"),
(col("cr.concept_id_2") == col("std.concept_id")) &
(col("std.standard_concept") == "S"),
"left"
)
# Step 4: Build OMOP condition_occurrence table
condition_occurrence = conditions_mapped.select(
col("fhir_condition_id"),
col("patient_ref"),
coalesce(col("std.concept_id"), lit(0))
.cast(LongType()).alias("condition_concept_id"),
to_date(coalesce(col("onset_date"), col("recorded_date")))
.alias("condition_start_date"),
to_date(col("abatement_date"))
.alias("condition_end_date"),
lit(32817).cast(LongType())
.alias("condition_type_concept_id"),
col("clinical_status")
.alias("condition_status_source_value"),
col("code_value").alias("condition_source_value"),
col("src.concept_id").cast(LongType())
.alias("condition_source_concept_id")
)
# Write to OMOP CDM (Delta Lake)
condition_occurrence.write \
.format("delta") \
.mode("overwrite") \
.option("overwriteSchema", "true") \
.save("s3://omop-cdm/condition_occurrence/")The OHDSI Tool Ecosystem
One of OMOP's biggest advantages over building analytics on raw FHIR data is the mature OHDSI tool ecosystem that runs natively on OMOP CDM. These are not toy tools — they power published peer-reviewed research and FDA drug safety analyses.
Key OHDSI Tools
| Tool | Purpose | Key Capabilities |
|---|---|---|
| ATLAS | Cohort definition and analysis | Visual cohort builder, incidence rate analysis, characterization studies — no SQL required |
| Achilles | Data characterization | Automated profiling: patient demographics, condition prevalence, data density, temporal coverage |
| Data Quality Dashboard | Data quality assessment | 3,400+ automated data quality checks aligned to Kahn Framework (conformance, completeness, plausibility) |
| CohortDiagnostics | Cohort evaluation | Evaluate phenotype definitions: sensitivity/specificity tradeoffs, incidence trends, cohort overlap |
| PatientLevelPrediction | ML prediction models | Regularized regression, random forests, deep learning — with validated prediction framework |
| CohortMethod | Causal inference | Propensity score matching, stratification, IPTW for comparative effectiveness research |
| SelfControlledCaseSeries | Drug safety studies | Self-controlled study designs for adverse event detection (used in COVID vaccine safety studies) |
| ARES | Data quality reporting | Interactive data quality reports, source-to-concept mapping coverage, temporal data patterns |
ATLAS Cohort Definition Example
With OMOP + ATLAS, defining a research cohort that would take days of custom SQL takes minutes. Here is the equivalent SQL for a common cohort — patients newly diagnosed with Type 2 Diabetes who were prescribed Metformin within 90 days:
-- OMOP CDM: Cohort definition
-- New T2DM patients started on Metformin within 90 days
WITH first_diabetes AS (
SELECT
co.person_id,
MIN(co.condition_start_date) AS index_date
FROM condition_occurrence co
WHERE co.condition_concept_id IN (
-- T2DM and all descendants
SELECT descendant_concept_id
FROM concept_ancestor
WHERE ancestor_concept_id = 201826
)
GROUP BY co.person_id
),
metformin_within_90d AS (
SELECT
de.person_id,
de.drug_exposure_start_date,
fd.index_date
FROM drug_exposure de
INNER JOIN first_diabetes fd
ON de.person_id = fd.person_id
WHERE de.drug_concept_id IN (
-- Metformin and all formulations
SELECT descendant_concept_id
FROM concept_ancestor
WHERE ancestor_concept_id = 1503297
)
AND de.drug_exposure_start_date
BETWEEN fd.index_date
AND fd.index_date + INTERVAL '90 days'
)
SELECT
fd.person_id,
fd.index_date AS cohort_start_date,
p.gender_concept_id,
p.year_of_birth,
EXTRACT(YEAR FROM fd.index_date) - p.year_of_birth AS age_at_index
FROM first_diabetes fd
INNER JOIN metformin_within_90d m
ON fd.person_id = m.person_id
INNER JOIN person p
ON fd.person_id = p.person_id
INNER JOIN observation_period op
ON fd.person_id = op.person_id
AND fd.index_date >= op.observation_period_start_date + INTERVAL '365 days'
AND fd.index_date <= op.observation_period_end_date;In ATLAS, this same cohort is defined visually with drag-and-drop concept sets — no SQL needed. The tool generates the SQL and executes it across any OMOP-compliant database.
Bridging FHIR and OMOP: The Dual-Standard Architecture
Why You Need Both
A learning health system — one that continuously improves care based on its own data — requires both standards working together. Here is the pattern:
- FHIR powers the operational layer: EHR integrations, SMART on FHIR apps, patient portals, care coordination
- OMOP powers the analytical layer: Population health dashboards, quality measures, research studies
- The ETL bridge: Continuous pipeline that transforms FHIR events into OMOP tables
- Insights flow back: Analytics results (risk scores, cohort flags) are written back to FHIR as Observations or RiskAssessments, available in real-time clinical workflows
Production Architecture
# docker-compose.yml -- FHIR + OMOP Development Stack
version: '3.8'
services:
# FHIR Server (HAPI FHIR)
hapi-fhir:
image: hapiproject/hapi:latest
ports:
- "8080:8080"
environment:
- hapi.fhir.default_encoding=json
- hapi.fhir.subscription.resthook_enabled=true
- spring.datasource.url=jdbc:postgresql://fhir-db:5432/hapi
depends_on:
- fhir-db
fhir-db:
image: postgres:16
environment:
POSTGRES_DB: hapi
POSTGRES_USER: hapi
POSTGRES_PASSWORD: hapi_secret
volumes:
- fhir-data:/var/lib/postgresql/data
# OMOP CDM Database
omop-db:
image: postgres:16
environment:
POSTGRES_DB: omop
POSTGRES_USER: omop
POSTGRES_PASSWORD: omop_secret
volumes:
- omop-data:/var/lib/postgresql/data
- ./omop-ddl:/docker-entrypoint-initdb.d
# OHDSI WebAPI + ATLAS
ohdsi-webapi:
image: ohdsi/webapi:latest
ports:
- "8081:8080"
environment:
- DATASOURCE_URL=jdbc:postgresql://omop-db:5432/omop
- DATASOURCE_USERNAME=omop
- DATASOURCE_PASSWORD=omop_secret
depends_on:
- omop-db
atlas:
image: ohdsi/atlas:latest
ports:
- "8082:8080"
depends_on:
- ohdsi-webapi
# ETL Pipeline (Airflow)
airflow:
image: apache/airflow:2.8.1
ports:
- "8083:8080"
volumes:
- ./dags:/opt/airflow/dags
environment:
- FHIR_SERVER_URL=http://hapi-fhir:8080/fhir
- OMOP_DB_URL=postgresql://omop:omop_secret@omop-db:5432/omop
volumes:
fhir-data:
omop-data:Airflow DAG for Continuous FHIR-to-OMOP Sync
# dags/fhir_to_omop_etl.py
# Airflow DAG: Continuous FHIR-to-OMOP ETL Pipeline
# Runs hourly, extracts new/updated FHIR resources,
# transforms to OMOP CDM, loads into PostgreSQL.
from airflow import DAG
from airflow.operators.python import PythonOperator
from airflow.utils.dates import days_ago
from datetime import timedelta
import requests
import psycopg2
default_args = {
'owner': 'data-engineering',
'depends_on_past': False,
'retries': 3,
'retry_delay': timedelta(minutes=5),
}
dag = DAG(
'fhir_to_omop_etl',
default_args=default_args,
description='Continuous FHIR to OMOP CDM sync',
schedule_interval='@hourly',
start_date=days_ago(1),
catchup=False,
tags=['fhir', 'omop', 'etl'],
)
def extract_fhir_resources(resource_type, **context):
"""Extract FHIR resources updated since last run."""
fhir_url = "http://hapi-fhir:8080/fhir"
last_updated = context['prev_execution_date_success'] or '2024-01-01'
resources = []
url = f"{fhir_url}/{resource_type}?_lastUpdated=gt{last_updated}"
while url:
resp = requests.get(url, headers={"Accept": "application/fhir+json"})
bundle = resp.json()
resources.extend(
[e['resource'] for e in bundle.get('entry', [])]
)
next_link = next(
(l['url'] for l in bundle.get('link', [])
if l['relation'] == 'next'), None
)
url = next_link
context['ti'].xcom_push(
key=f'{resource_type}_resources', value=resources
)
return len(resources)
# Define tasks for each resource type
for resource, omop_table in [
('Condition', 'condition_occurrence'),
('MedicationRequest', 'drug_exposure'),
('Observation', 'measurement'),
('Procedure', 'procedure_occurrence'),
('Encounter', 'visit_occurrence'),
]:
extract = PythonOperator(
task_id=f'extract_{resource.lower()}',
python_callable=extract_fhir_resources,
op_args=[resource],
dag=dag,
)
# transform and load tasks follow same pattern
# extract >> transform >> loadExisting FHIR-to-OMOP Tools
You do not have to build the entire ETL from scratch. Several open-source projects handle FHIR-to-OMOP conversion:
| Tool | Approach | Best For |
|---|---|---|
| ETL-Synthea | Reference ETL for Synthea synthetic data | Learning OMOP ETL patterns, development/testing |
| NACHC fhir-to-omop | Java-based FHIR-to-OMOP converter | Production-grade conversion with vocabulary support |
| Google FHIR Data Pipes | Apache Beam pipeline for FHIR to analytics formats | Cloud-scale ETL with Parquet/BigQuery output |
| HL7 FHIR-OMOP IG | Official HL7 Implementation Guide for FHIR/OMOP mapping | Reference specification for mapping standards |
| Usagi | Semi-automated vocabulary mapping tool | Mapping custom/proprietary codes to OMOP standard concepts |
Data Quality: Achilles and the DQD
Before running any research on your OMOP CDM, you must validate data quality. The OHDSI community provides two essential tools:
Running Achilles
# R: Run Achilles characterization on your OMOP CDM
library(Achilles)
connectionDetails <- createConnectionDetails(
dbms = "postgresql",
server = "omop-db/omop",
user = "omop",
password = "omop_secret",
port = 5432
)
achilles(
connectionDetails = connectionDetails,
cdmDatabaseSchema = "cdm",
resultsDatabaseSchema = "results",
vocabDatabaseSchema = "cdm",
sourceName = "Hospital_FHIR_Source",
cdmVersion = "5.4",
numThreads = 4,
optimizeAtlasCache = TRUE
)
# Generates 200+ analysis tables covering:
# - Demographics distributions
# - Condition prevalence over time
# - Drug utilization patterns
# - Measurement value distributions
# - Visit type breakdowns
# - Data density and completeness metricsRunning the Data Quality Dashboard
# R: Run DQD -- 3,400+ automated quality checks
library(DataQualityDashboard)
DataQualityDashboard::executeDqChecks(
connectionDetails = connectionDetails,
cdmDatabaseSchema = "cdm",
resultsDatabaseSchema = "results",
vocabDatabaseSchema = "cdm",
cdmSourceName = "Hospital_FHIR_Source",
numThreads = 4,
outputFolder = "./dqd_results",
checkLevels = c("TABLE", "FIELD", "CONCEPT"),
checkNames = c() # Run all checks
)
# Generates an interactive HTML report with:
# - Conformance checks (data types, values within range)
# - Completeness checks (% of NULL fields per table)
# - Plausibility checks (age at death > 0, dates in valid range)
# - Pass/fail for each of 3,400+ individual checksThese tools are what make OMOP production-ready for research. Running analytics on a data quality-validated OMOP CDM gives you confidence that your results reflect reality, not data errors.
Real-World Use Cases
1. COVID-19 Vaccine Safety (OHDSI Network Study)
In 2021-2022, the OHDSI network ran the largest vaccine safety study in history across 13 databases in 8 countries, covering 126 million vaccinated individuals. Because all sites used OMOP CDM, the same analysis code ran at each site without sharing patient data. Results were published in the British Medical Journal.
2. FDA Sentinel Initiative
The FDA's Sentinel System uses OMOP CDM to monitor drug safety across a network covering 100+ million lives. Active surveillance queries run across distributed OMOP databases to detect adverse drug events in near real-time.
3. Clinical Trial Feasibility
Pharmaceutical companies use OMOP CDM + ATLAS to assess clinical trial feasibility before committing to site selection. By defining inclusion/exclusion criteria in ATLAS and running them across network sites, they can estimate patient counts in weeks instead of months.