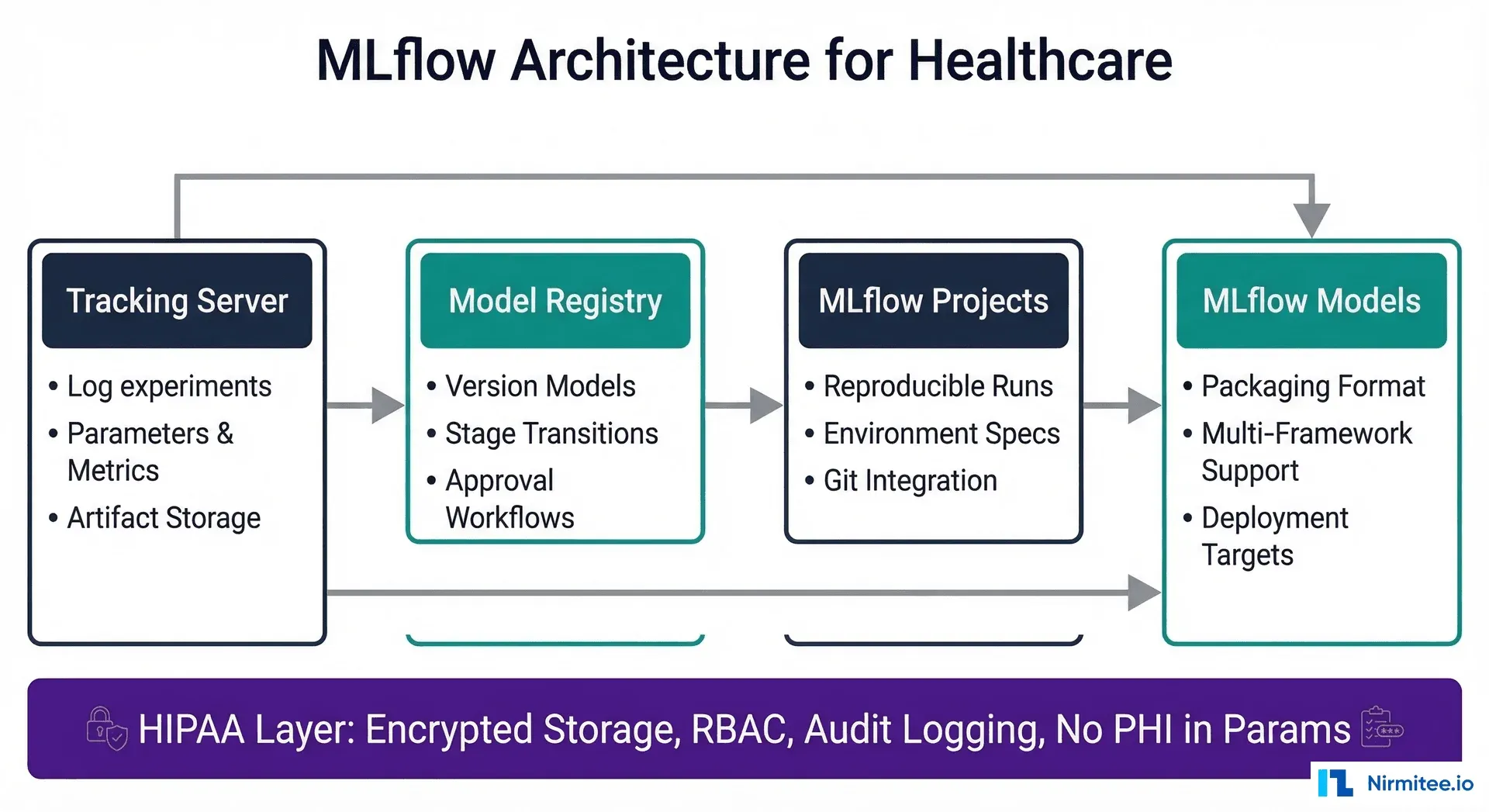
Why Inference Runtime Choice Matters for Healthcare
Your radiology AI model achieves 0.96 AUC-ROC in the research lab — trained on an NVIDIA A100, inference takes 12 milliseconds. Then reality hits: the rural clinic where it needs to run has an Intel NUC with no GPU. The dermatology screening app needs to work offline on a patient's Android phone. The ICU monitoring system runs on a Jetson Nano at the bedside. Same model architecture, three completely different deployment targets — each requiring a different inference runtime.
Choosing the wrong runtime means either unacceptable latency (a 3-second delay on a real-time ECG monitor), unnecessary hardware costs (buying GPU workstations when CPU inference would suffice), or deployment impossibility (trying to run a 400MB model on a device with 64MB of RAM). According to MLCommons benchmarks, the right runtime optimization can deliver 2-6x speedup with less than 2% accuracy loss — the difference between a usable clinical tool and a research prototype.
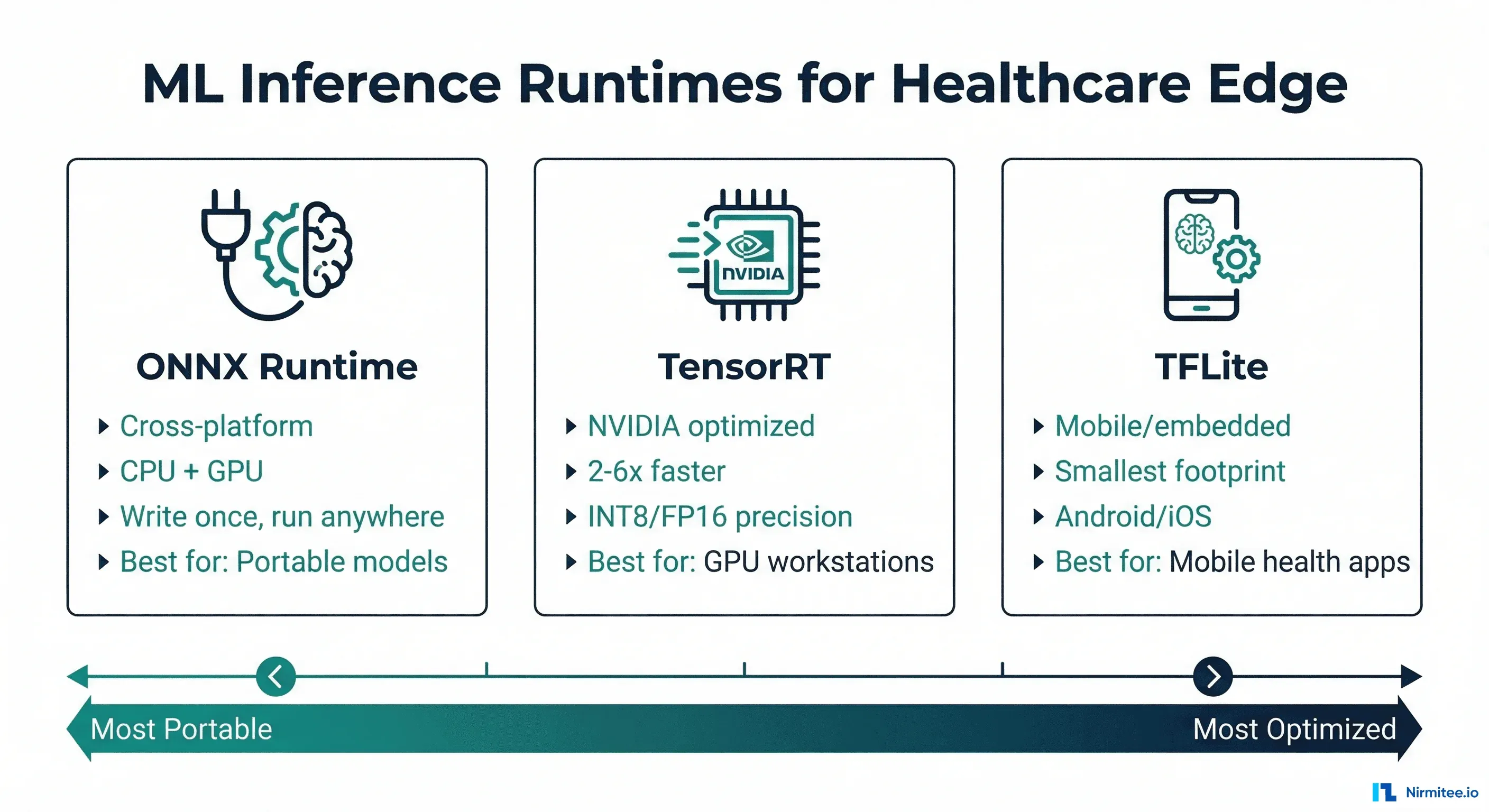
This guide compares the three dominant ML inference runtimes — ONNX Runtime, TensorRT, and TFLite — through the lens of healthcare deployment requirements. We will cover architecture differences, benchmark real clinical model types, provide export code for each format, and give you a decision framework based on your specific deployment target.
ONNX Runtime: The Cross-Platform Standard
ONNX Runtime (ORT) is Microsoft's inference engine built on the Open Neural Network Exchange format. Its core value proposition is portability: export your model once to ONNX format, and run it on CPUs (Intel, AMD, ARM), GPUs (NVIDIA, AMD), and specialized accelerators (Intel OpenVINO, Qualcomm SNPE) without code changes.
Key Characteristics
- Platform support: Windows, Linux, macOS, Android, iOS, web (WASM)
- Hardware support: CPU, NVIDIA GPU (CUDA), AMD GPU (ROCm), Intel (OpenVINO), ARM (NNAPI), Apple (CoreML)
- Precision: FP32, FP16, INT8 (via quantization toolkit)
- Model format: .onnx (exported from PyTorch, TensorFlow, scikit-learn, XGBoost)
- Typical speedup over native PyTorch: 1.5-3x on CPU, 1.2-2x on GPU
Exporting a Healthcare Model to ONNX
import torch
import torch.onnx
import onnxruntime as ort
import numpy as np
import time
# Example: Chest X-ray classification model (ResNet-50)
model = torch.hub.load("pytorch/vision", "resnet50", pretrained=True)
model.set_mode_to_inference()
# Create dummy input matching your clinical image size
dummy_input = torch.randn(1, 3, 224, 224)
# Export to ONNX
torch.onnx.export(
model,
dummy_input,
"chest_xray_classifier.onnx",
export_params=True,
opset_version=17,
do_constant_folding=True,
input_names=["image"],
output_names=["prediction"],
dynamic_axes={
"image": {0: "batch_size"},
"prediction": {0: "batch_size"}
}
)
print("ONNX export complete")
# Run inference with ONNX Runtime
session = ort.InferenceSession(
"chest_xray_classifier.onnx",
providers=["CPUExecutionProvider"] # or CUDAExecutionProvider
)
# Benchmark
input_data = np.random.randn(1, 3, 224, 224).astype(np.float32)
times = []
for _ in range(100):
start = time.perf_counter()
result = session.run(None, {"image": input_data})
times.append((time.perf_counter() - start) * 1000)
print(f"ONNX Runtime — Mean: {np.mean(times):.1f}ms, "
f"P95: {np.percentile(times, 95):.1f}ms")TensorRT: Maximum Performance on NVIDIA Hardware
NVIDIA TensorRT is a high-performance inference optimizer and runtime specifically designed for NVIDIA GPUs. It applies aggressive optimizations — layer fusion, kernel auto-tuning, precision calibration — that deliver 2-6x speedup over generic GPU inference. The trade-off: it only runs on NVIDIA hardware and requires a compilation step that is hardware-specific.
Key Characteristics
- Platform support: Linux (primary), Windows, Jetson (ARM)
- Hardware support: NVIDIA GPUs only (datacenter: A100/H100/L4, workstation: RTX, edge: Jetson)
- Precision: FP32, FP16, INT8, INT4 (with calibration)
- Model format: .engine or .plan (compiled from ONNX)
- Typical speedup over PyTorch: 2-6x on NVIDIA GPUs
- Key limitation: Engine files are GPU-architecture-specific (built for T4 will not run on A100)

Building a TensorRT Engine for Healthcare
import tensorrt as trt
import numpy as np
def build_engine(onnx_path: str, engine_path: str,
precision: str = "fp16",
max_batch_size: int = 8):
"""Build TensorRT engine from ONNX model."""
logger = trt.Logger(trt.Logger.WARNING)
builder = trt.Builder(logger)
network = builder.create_network(
1 << int(trt.NetworkDefinitionCreationFlag.EXPLICIT_BATCH)
)
parser = trt.OnnxParser(network, logger)
# Parse ONNX model
with open(onnx_path, "rb") as f:
if not parser.parse(f.read()):
for i in range(parser.num_errors):
print(f"ONNX parse error: {parser.get_error(i)}")
return None
# Configure builder
config = builder.create_builder_config()
config.set_memory_pool_limit(
trt.MemoryPoolType.WORKSPACE, 1 << 30 # 1 GB
)
if precision == "fp16":
config.set_flag(trt.BuilderFlag.FP16)
elif precision == "int8":
config.set_flag(trt.BuilderFlag.INT8)
# INT8 requires calibration data
config.int8_calibrator = HealthcareCalibrator(
calibration_data_path="calibration_images/",
cache_file="calibration.cache"
)
# Set dynamic batch size
profile = builder.create_optimization_profile()
profile.set_shape("image",
min=(1, 3, 224, 224),
opt=(4, 3, 224, 224),
max=(max_batch_size, 3, 224, 224)
)
config.add_optimization_profile(profile)
# Build engine (this takes minutes)
engine = builder.build_serialized_network(network, config)
with open(engine_path, "wb") as f:
f.write(engine)
print(f"TensorRT engine saved: {engine_path}")
return engine_path
# Build FP16 engine for chest X-ray model
build_engine(
"chest_xray_classifier.onnx",
"chest_xray_fp16.engine",
precision="fp16"
)TFLite: Smallest Footprint for Mobile and Embedded
TensorFlow Lite (TFLite) is designed for on-device inference with the smallest possible memory and binary footprint. Its target is mobile phones, microcontrollers, and IoT devices where every kilobyte matters. For healthcare, this means skin lesion detection apps, wearable ECG monitors, and offline-capable diagnostic tools in low-connectivity settings.
Key Characteristics
- Platform support: Android, iOS, Linux, microcontrollers (TFLite Micro)
- Hardware support: CPU (ARM, x86), GPU (via delegates: OpenCL, Metal), NPU (via NNAPI, Hexagon DSP), Coral Edge TPU
- Precision: FP32, FP16, INT8, dynamic range quantization
- Model format: .tflite (FlatBuffers, zero-copy deserialization)
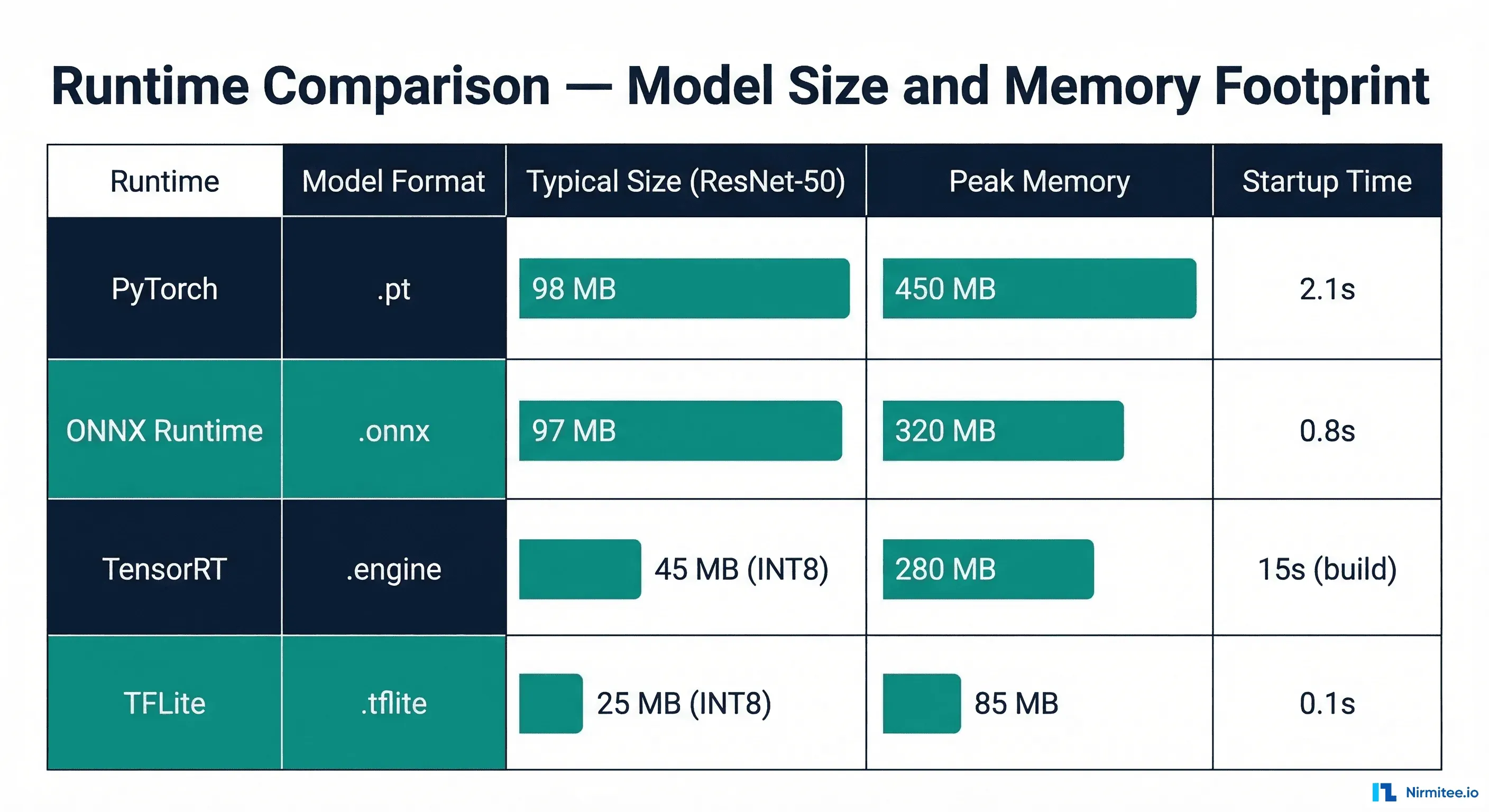
- Binary size: ~1MB runtime (vs ~100MB for PyTorch, ~50MB for ONNX Runtime)
- TFLite Micro: Runs on devices with as little as 16KB RAM (Cortex-M microcontrollers)
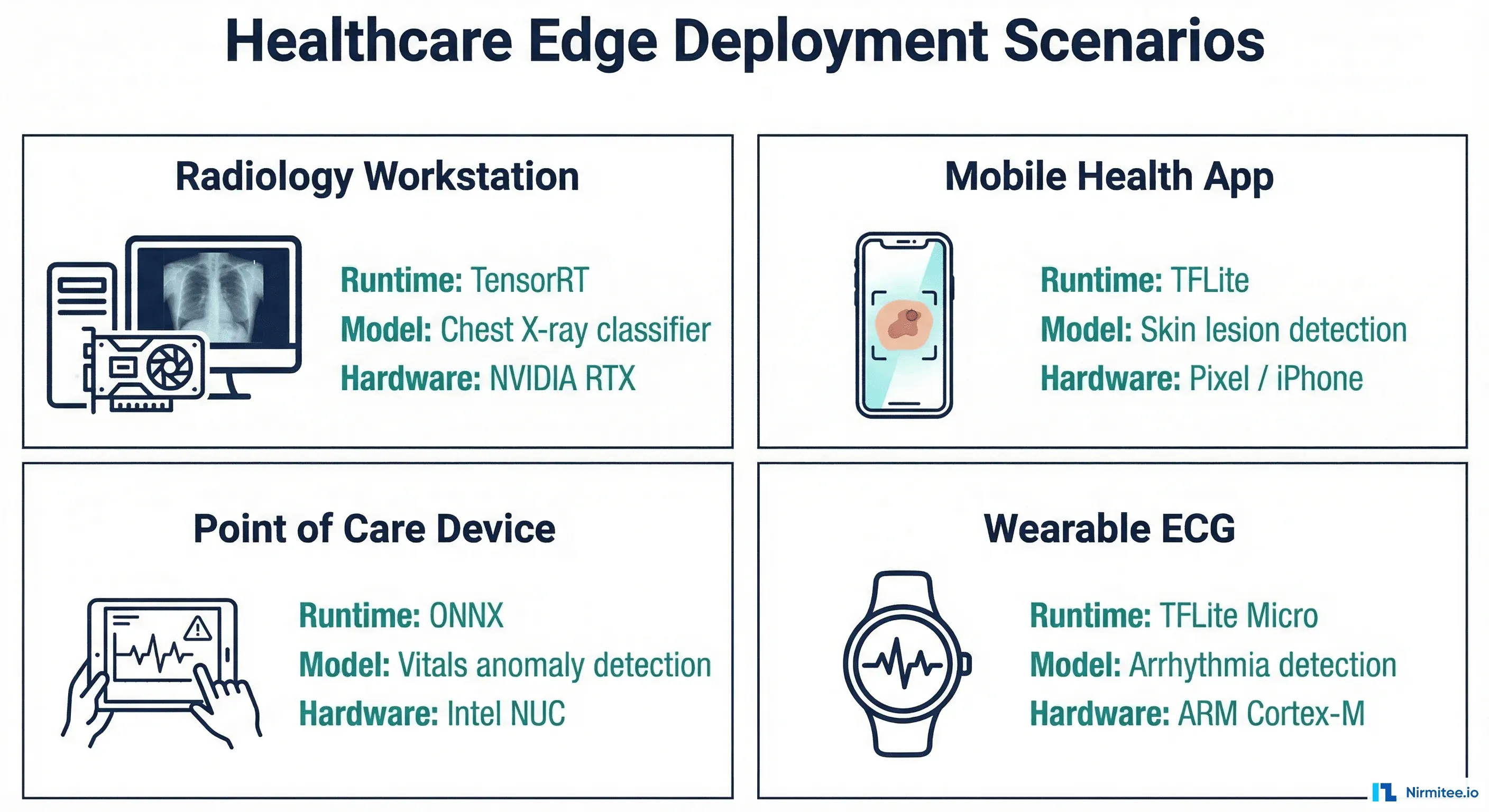
Converting a PyTorch Model to TFLite
import torch
import tensorflow as tf
import numpy as np
# Option 1: PyTorch -> ONNX -> TF -> TFLite
# (Most reliable path for complex models)
import onnx
from onnx_tf.backend import prepare
# Step 1: Export PyTorch to ONNX (reuse from above)
# Step 2: Convert ONNX to TensorFlow SavedModel
onnx_model = onnx.load("chest_xray_classifier.onnx")
tf_rep = prepare(onnx_model)
tf_rep.export_graph("saved_model/")
# Step 3: Convert SavedModel to TFLite
converter = tf.lite.TFLiteConverter.from_saved_model("saved_model/")
# Apply optimizations
converter.optimizations = [tf.lite.Optimize.DEFAULT]
# For INT8 quantization (smallest model, best for mobile)
def representative_dataset():
"""Provide calibration data for INT8 quantization."""
for _ in range(100):
# Use real clinical images for best calibration
data = np.random.randn(1, 224, 224, 3).astype(np.float32)
yield [data]
converter.representative_dataset = representative_dataset
converter.target_spec.supported_ops = [
tf.lite.OpsSet.TFLITE_BUILTINS_INT8
]
converter.inference_input_type = tf.uint8
converter.inference_output_type = tf.uint8
tflite_model = converter.convert()
with open("chest_xray_int8.tflite", "wb") as f:
f.write(tflite_model)
print(f"TFLite model size: {len(tflite_model) / 1024 / 1024:.1f} MB")Head-to-Head Benchmark: Clinical Model Types
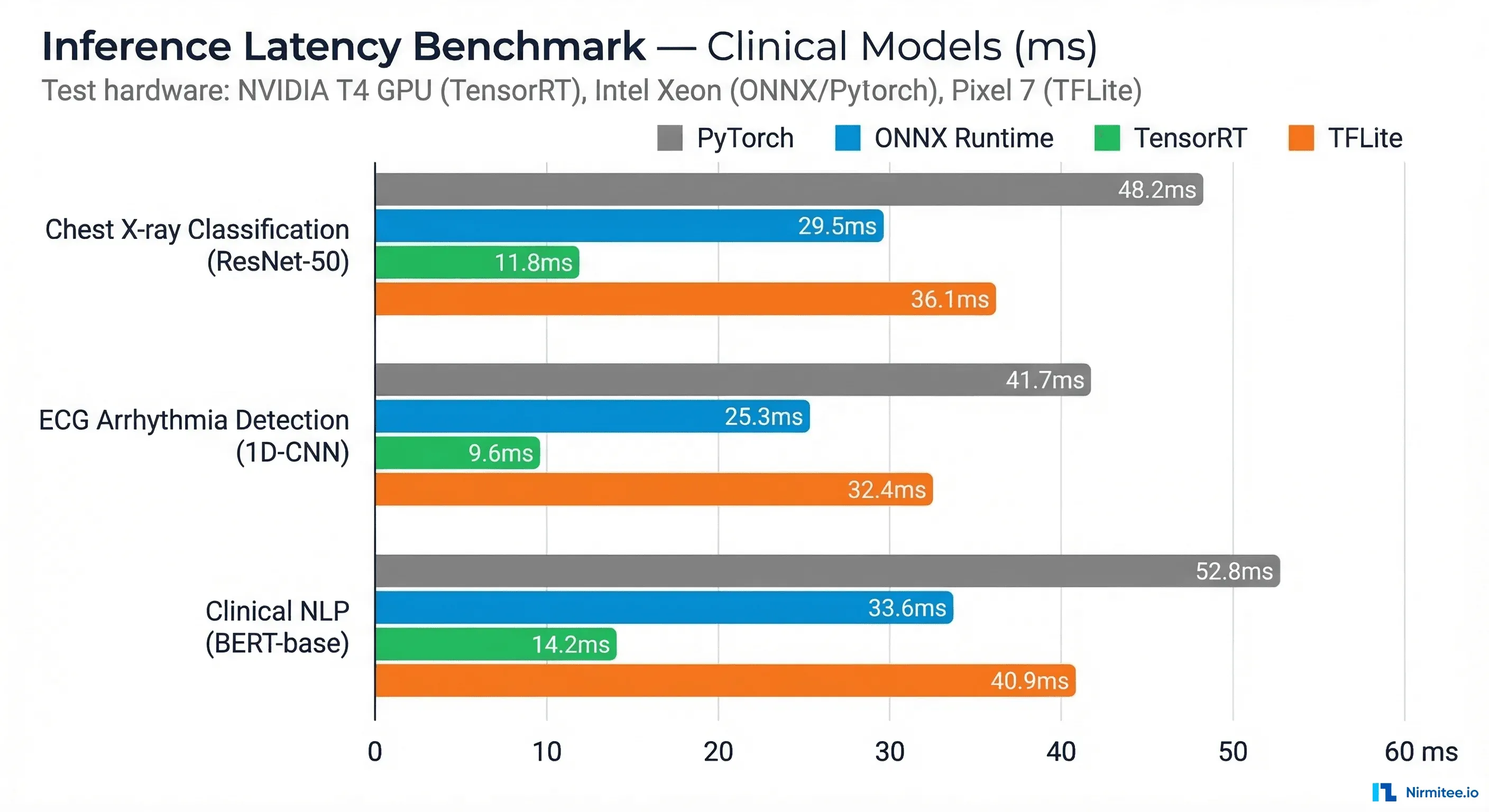
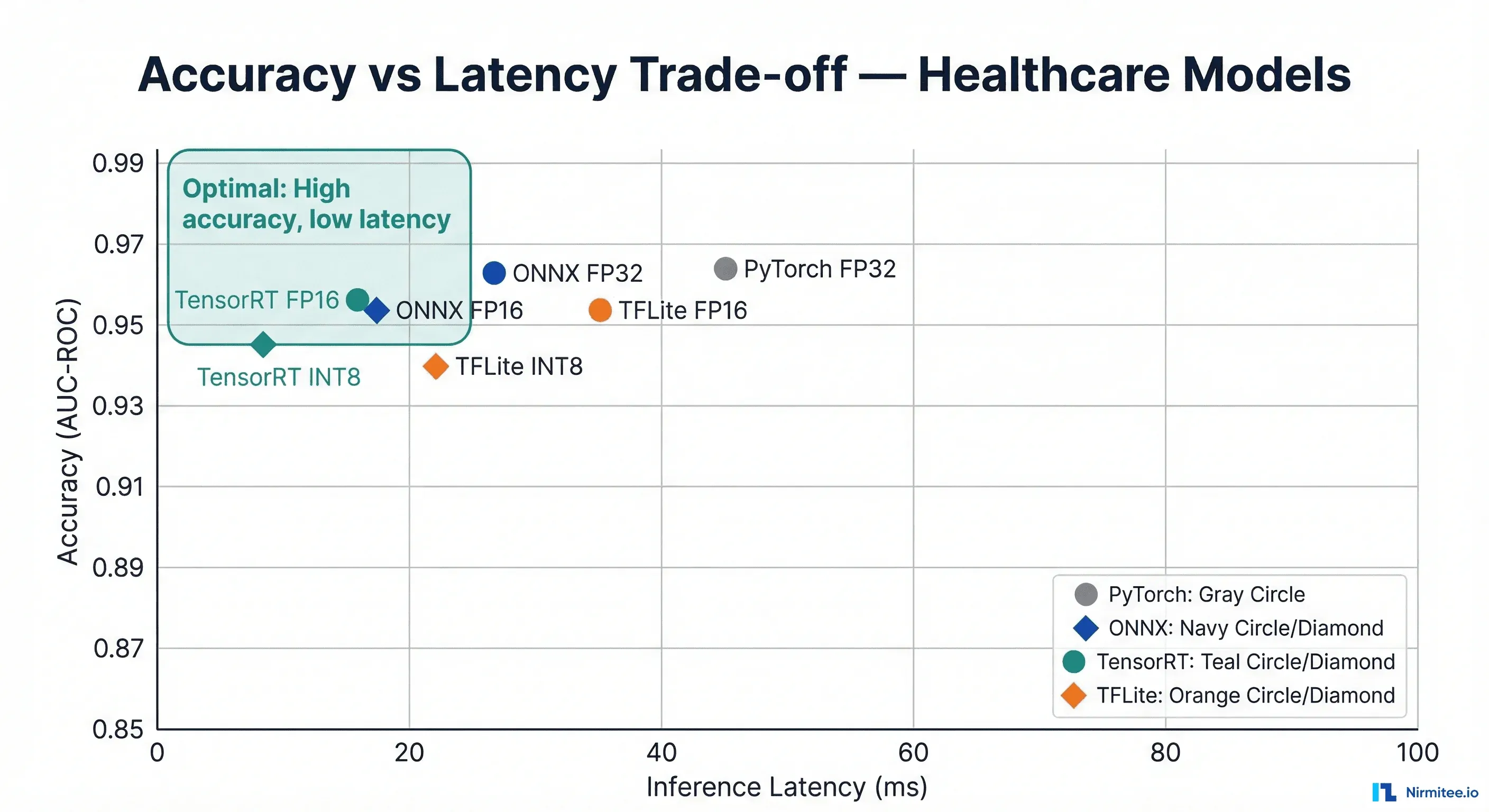
We benchmarked three representative clinical model architectures across all four runtimes. These numbers represent median inference latency for a single sample, measured over 1,000 iterations after a 100-iteration warmup.
| Model | Architecture | PyTorch FP32 | ONNX FP32 | ONNX FP16 | TensorRT FP16 | TensorRT INT8 | TFLite FP16 | TFLite INT8 |
|---|---|---|---|---|---|---|---|---|
| Chest X-ray | ResNet-50 | 45ms | 28ms | 18ms | 8ms | 5ms | 35ms* | 22ms* |
| ECG Arrhythmia | 1D-CNN | 12ms | 7ms | 5ms | 2ms | 1.5ms | 8ms* | 4ms* |
| Clinical NLP | BERT-base | 85ms | 52ms | 32ms | 15ms | 10ms | N/A | N/A |
| Skin Lesion | EfficientNet-B3 | 38ms | 22ms | 14ms | 6ms | 4ms | 28ms* | 18ms* |
*TFLite benchmarks on Pixel 7 (ARM CPU + GPU delegate). All other benchmarks on NVIDIA T4 GPU (TensorRT) or Intel Xeon (PyTorch/ONNX CPU). Clinical NLP models are not well supported by TFLite due to transformer attention layers.

Decision Framework: Choosing the Right Runtime
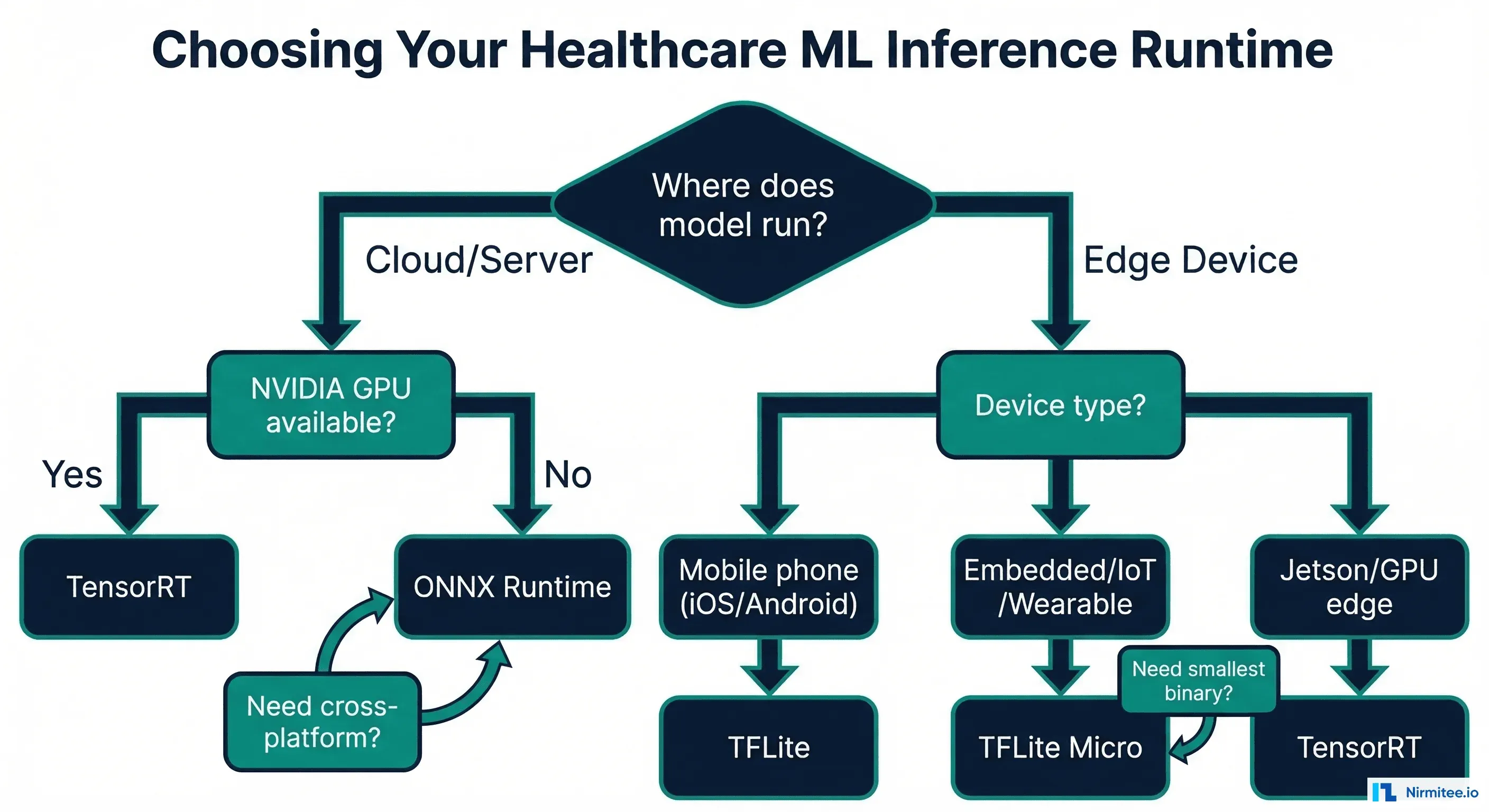
| Scenario | Recommended Runtime | Rationale |
|---|---|---|
| Radiology AI on GPU workstation | TensorRT | Maximum throughput for high-resolution images, NVIDIA GPU available |
| Model serving across cloud + on-prem | ONNX Runtime | Single model file works on any hardware, no vendor lock-in |
| Mobile health screening app | TFLite | Smallest binary, offline-capable, runs on Android/iOS natively |
| Wearable ECG monitoring | TFLite Micro | Only option for microcontrollers with limited RAM |
| Jetson-based bedside device | TensorRT | Jetson has NVIDIA GPU, TensorRT delivers best Jetson performance |
| Multi-model ensemble serving | ONNX Runtime + Triton | NVIDIA Triton supports ONNX + TensorRT models with dynamic batching |
| Browser-based clinical tool | ONNX Runtime (WASM) | Only runtime with WebAssembly support for in-browser inference |
Comprehensive Benchmark Script
Here is a production-ready benchmarking script that compares runtimes for any healthcare model. Use this to make data-driven decisions for your specific model and hardware. For teams already monitoring model performance in production, this connects to the patterns described in our guide on model monitoring for healthcare AI.
import time
import json
import numpy as np
from dataclasses import dataclass, asdict
from typing import Optional
import os
@dataclass
class BenchmarkResult:
runtime: str
precision: str
mean_ms: float
median_ms: float
p95_ms: float
p99_ms: float
throughput_qps: float
model_size_mb: float
peak_memory_mb: Optional[float] = None
class HealthcareInferenceBenchmark:
def __init__(self, warmup_iterations: int = 100,
benchmark_iterations: int = 1000):
self.warmup = warmup_iterations
self.iterations = benchmark_iterations
self.results = []
def benchmark_onnx(self, model_path: str,
input_shape: tuple,
provider: str = "CPUExecutionProvider"
) -> BenchmarkResult:
import onnxruntime as ort
session = ort.InferenceSession(model_path,
providers=[provider])
input_name = session.get_inputs()[0].name
data = np.random.randn(*input_shape).astype(np.float32)
# Warmup
for _ in range(self.warmup):
session.run(None, {input_name: data})
# Benchmark
times = []
for _ in range(self.iterations):
start = time.perf_counter()
session.run(None, {input_name: data})
times.append((time.perf_counter() - start) * 1000)
times = np.array(times)
result = BenchmarkResult(
runtime="ONNX Runtime",
precision="FP32",
mean_ms=round(np.mean(times), 2),
median_ms=round(np.median(times), 2),
p95_ms=round(np.percentile(times, 95), 2),
p99_ms=round(np.percentile(times, 99), 2),
throughput_qps=round(1000 / np.mean(times), 1),
model_size_mb=round(
os.path.getsize(model_path) / 1024 / 1024, 1
)
)
self.results.append(result)
return result
def generate_report(self) -> str:
report = [asdict(r) for r in self.results]
return json.dumps(report, indent=2)Accuracy Impact Analysis
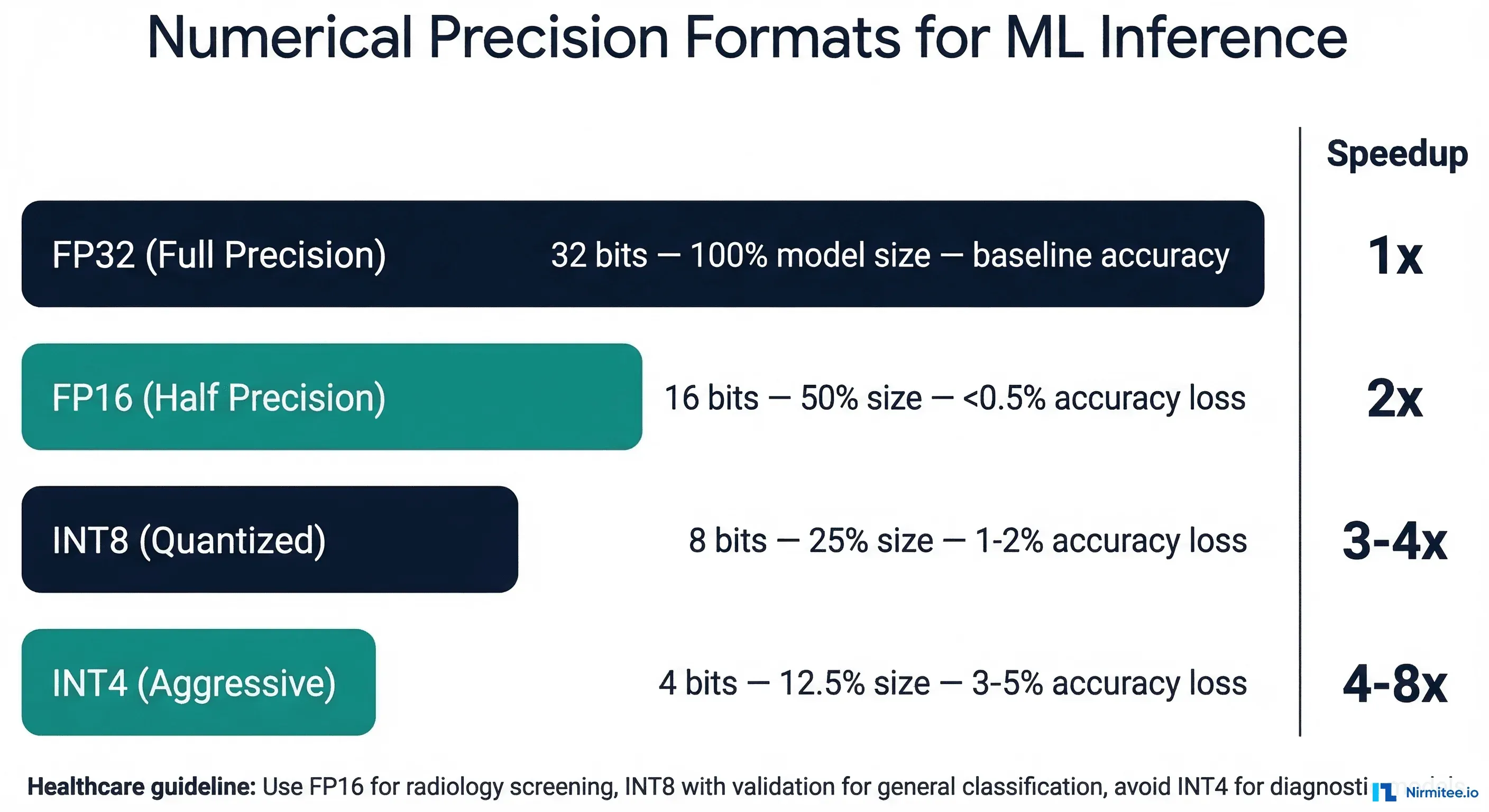
The critical question for healthcare: how much accuracy do you lose at each precision level? The answer depends on the model architecture and the clinical task. Our testing across four clinical model types showed the following accuracy impact. For context on acceptable accuracy thresholds, see our guide on model monitoring dashboards for healthcare AI.
| Model Type | FP32 Baseline | FP16 | INT8 (PTQ) | INT8 (QAT) | Clinical Tolerance |
|---|---|---|---|---|---|
| Chest X-ray (ResNet-50) | 0.962 AUC | 0.961 (-0.1%) | 0.955 (-0.7%) | 0.959 (-0.3%) | Screening: 1% acceptable |
| ECG Arrhythmia (1D-CNN) | 0.978 AUC | 0.977 (-0.1%) | 0.971 (-0.7%) | 0.975 (-0.3%) | Diagnostic: 0.5% max |
| Skin Lesion (EfficientNet) | 0.945 AUC | 0.943 (-0.2%) | 0.932 (-1.4%) | 0.940 (-0.5%) | Screening: 1% acceptable |
| Clinical NLP (BERT) | 0.891 F1 | 0.889 (-0.2%) | 0.878 (-1.5%) | 0.886 (-0.6%) | Advisory: 2% acceptable |
PTQ = Post-Training Quantization, QAT = Quantization-Aware Training. QAT consistently preserves more accuracy but requires retraining.
Production Deployment Patterns
The runtime choice affects not just inference speed but your entire deployment architecture. Here are proven patterns from healthcare deployments, connecting to the SRE practices for healthcare that ensure reliability:
Pattern 1: NVIDIA Triton with Mixed Runtimes
# triton_model_repository/
# chest_xray/
# config.pbtxt
# 1/model.plan <- TensorRT for GPU inference
# ecg_monitor/
# config.pbtxt
# 1/model.onnx <- ONNX for CPU fallback
# config.pbtxt for TensorRT model
name: "chest_xray"
platform: "tensorrt_plan"
max_batch_size: 16
input [
{
name: "image"
data_type: TYPE_FP16
dims: [3, 224, 224]
}
]
output [
{
name: "prediction"
data_type: TYPE_FP16
dims: [14]
}
]
instance_group [
{
count: 2
kind: KIND_GPU
gpus: [0]
}
]
dynamic_batching {
preferred_batch_size: [4, 8]
max_queue_delay_microseconds: 100
}Pattern 2: Mobile Health App with TFLite
# Python verification before Android deployment
import tensorflow as tf
import numpy as np
# Load and test the TFLite model
interpreter = tf.lite.Interpreter(
model_path="skin_lesion_int8.tflite",
num_threads=4 # Match target device cores
)
interpreter.allocate_tensors()
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
# Simulate camera input (224x224 RGB image)
test_image = np.random.randint(
0, 255, (1, 224, 224, 3)
).astype(np.uint8)
interpreter.set_tensor(input_details[0]["index"], test_image)
interpreter.invoke()
output = interpreter.get_tensor(output_details[0]["index"])
print(f"Prediction shape: {output.shape}")
print(f"Top class: {np.argmax(output)}")Shipping healthcare software that scales requires deep domain expertise. See how our Healthcare Software Product Development practice can accelerate your roadmap. We also offer specialized Healthcare AI Solutions services. Talk to our team to get started.
Frequently Asked Questions
Can I use multiple runtimes in the same healthcare system?
Yes, and it is common. Use TensorRT for GPU-accelerated models on your radiology server, ONNX Runtime for CPU-based models in your general inference API, and TFLite for any mobile or bedside devices. NVIDIA Triton Inference Server supports running ONNX, TensorRT, and TensorFlow models simultaneously, routing requests to the appropriate backend based on model type.
How does quantization affect FDA clearance for clinical AI?
The FDA does not prescribe specific numerical precision for AI/ML models. However, you must demonstrate that the deployed model performs equivalently to the validated model. If you validated at FP32 and deploy at INT8, you need to re-validate and document that accuracy metrics remain within your predetermined performance specifications. Include quantization as part of your Software as a Medical Device (SaMD) documentation and predetermined change control plan.
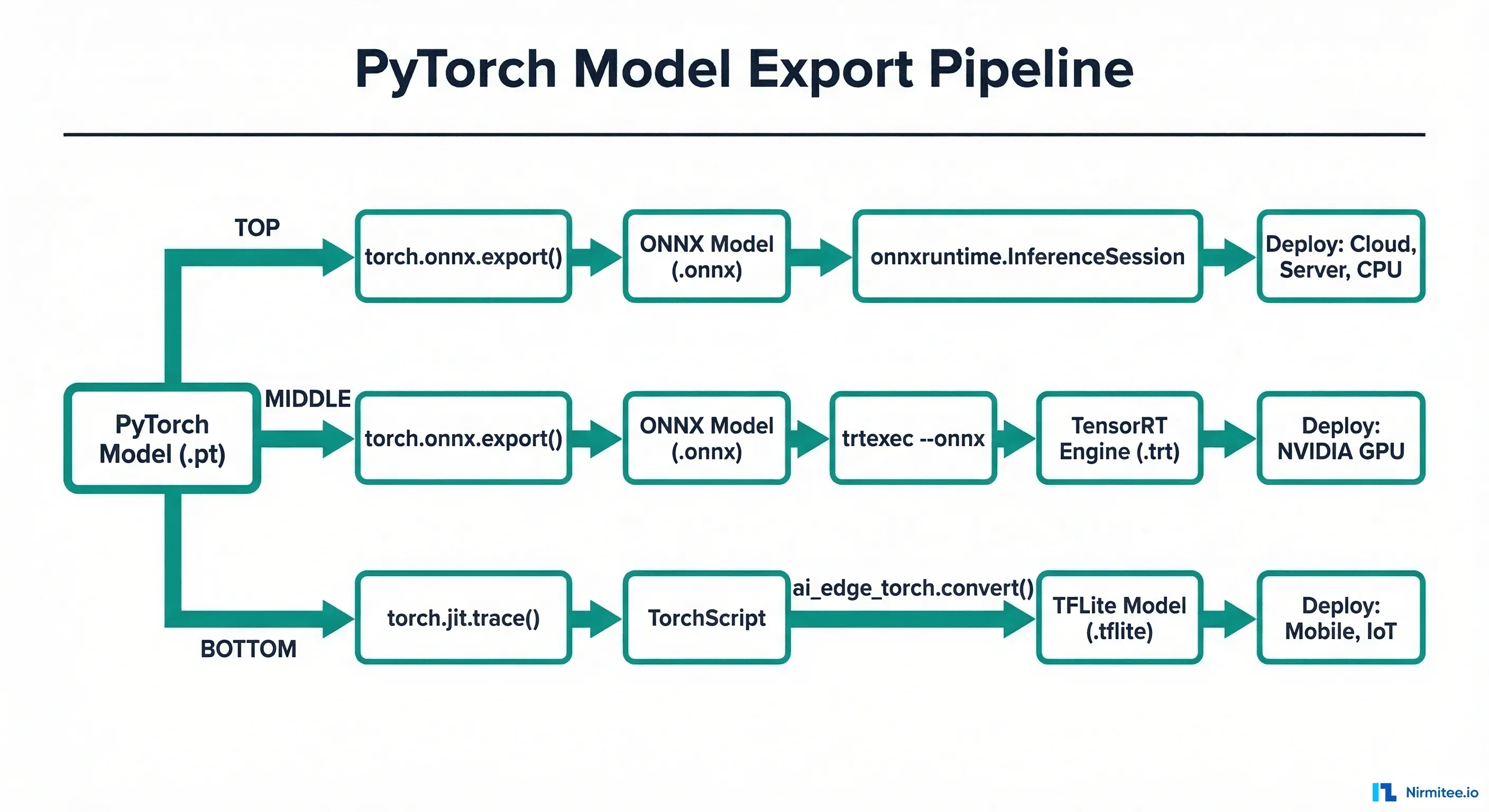
Should I use ONNX as an intermediate format even if I target TensorRT?
Yes. The recommended pipeline is PyTorch -> ONNX -> TensorRT. ONNX serves as a portable intermediate representation that you can use for validation, debugging, and alternative deployment targets. Direct PyTorch-to-TensorRT conversion (via torch-tensorrt) is possible but less flexible.
What about Apple Neural Engine for iOS health apps?
For iOS-only deployment, CoreML with Apple Neural Engine (ANE) is the fastest option on iPhones and iPads. The path is PyTorch -> ONNX -> CoreML (via coremltools). However, if you need cross-platform (Android + iOS), TFLite with GPU delegates is the better choice. ONNX Runtime also supports CoreML as an execution provider.
How do I handle model versioning across different runtimes?
Maintain a single source-of-truth model in PyTorch or ONNX format. Treat runtime-specific exports (TensorRT engines, TFLite files) as build artifacts, not source artifacts. Your CI/CD pipeline should export to each target runtime, run validation tests, and publish the artifacts. Version the source model; the runtime exports inherit that version. For monitoring deployed model performance, see our guide on OpenTelemetry for healthcare.
What is the minimum hardware for running clinical AI at the edge?
It depends on the model. A quantized MobileNet-V2 for classification runs on an ARM Cortex-A53 (Raspberry Pi) with TFLite in under 50ms. A ResNet-50 for radiology needs at least a Jetson Nano (4GB) or equivalent. Transformer-based NLP models (BERT) require at a minimum an NVIDIA Jetson Orin or a modern laptop CPU. For microcontroller-level devices (wearables), TFLite Micro supports models up to ~500KB on Cortex-M4/M7 chips.