Every healthcare AI team hits the same wall. You have a model architecture. You have a training pipeline. You have compute. What you do not have is data -- because real patient data is locked behind HIPAA, IRB approvals, data use agreements, and months of organizational negotiation. The model sits idle while lawyers and compliance officers debate access terms.
Synthetic data changes the equation entirely. Instead of waiting six months for a de-identified dataset from your partner health system -- a dataset that will arrive incomplete, biased toward one demographics, and still carrying re-identification risk -- you generate exactly the patient population you need, with exactly the clinical characteristics your model requires, in exactly the format your pipeline consumes. No PHI. No BAA. No IRB. No waiting.
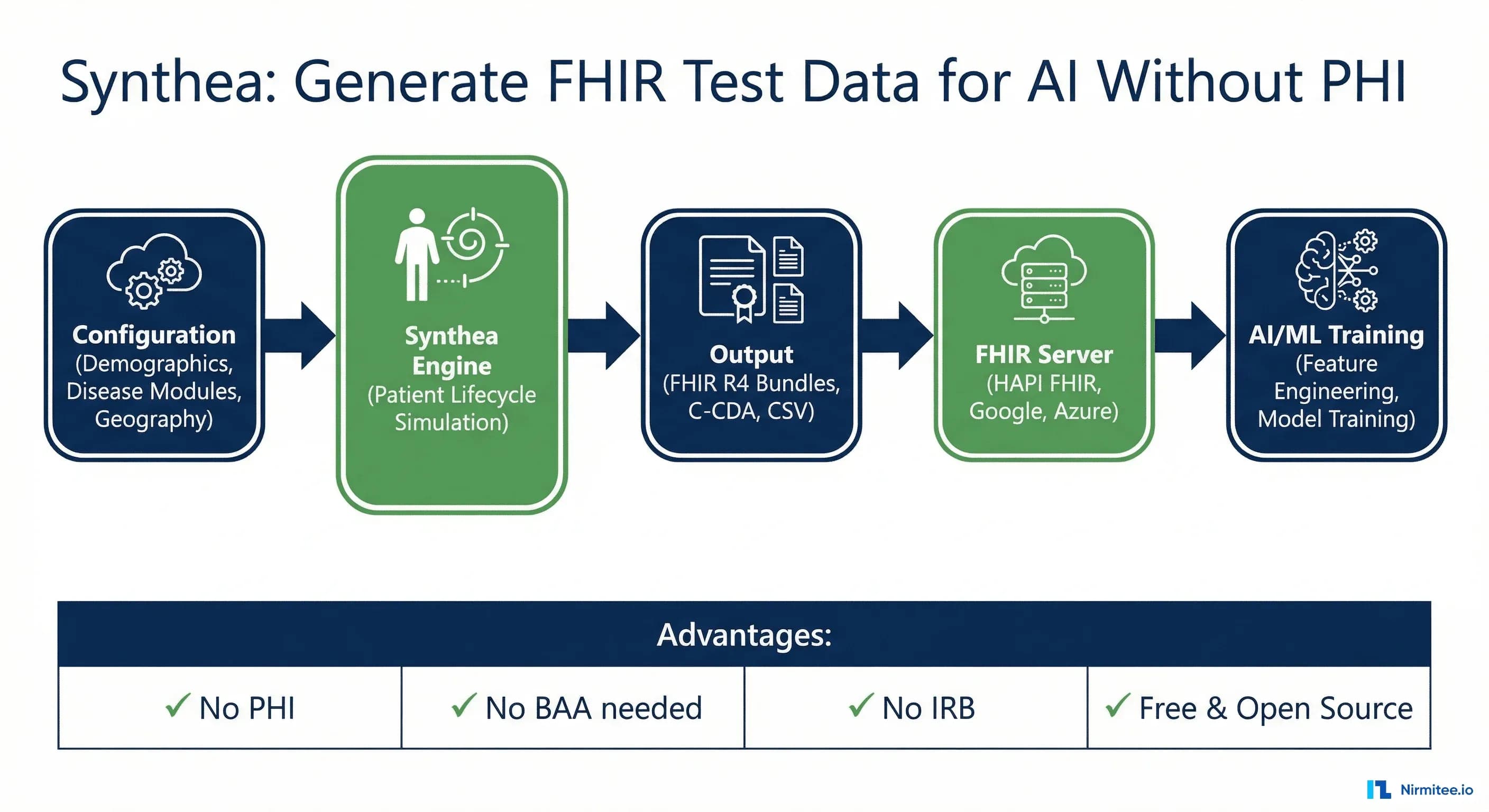
The tool that makes this possible is Synthea, MITRE's open-source synthetic patient generator. It produces clinically realistic patient records -- complete medical histories from birth to death -- and exports them as FHIR R4 Bundles ready for ingestion into any FHIR server or ML pipeline. Over 200 research papers cite Synthea. The MITRE-curated dataset on synthea.mitre.org has been downloaded millions of times. If you are building healthcare AI and you are not using synthetic data in your development workflow, you are leaving months of velocity on the table.
The Healthcare AI Data Problem
A 2024 Nature Medicine review found that the median training dataset for FDA-cleared AI/ML medical devices contains fewer than 1,000 patients. For comparison, ImageNet contains 14 million labeled images. Healthcare AI is operating with orders of magnitude less data than consumer AI, not because the data does not exist, but because accessing it is extraordinarily difficult.
The barriers compound on each other:
HIPAA and PHI Restrictions
The HIPAA Privacy Rule classifies 18 categories of protected health information (PHI). Any dataset containing names, dates, geographic data smaller than a state, or medical record numbers requires a Business Associate Agreement (BAA) with every entity that touches it. For a startup building an AI model, this means legal review of every data partnership, security assessments, and breach notification procedures. Health systems report that data access agreements take 3-9 months to negotiate, and many smaller organizations cannot absorb the compliance cost.
De-Identification Limitations
HIPAA provides two de-identification methods -- Safe Harbor (remove all 18 identifiers) and Expert Determination (statistical proof of low re-identification risk). Both have serious limitations for AI training. Safe Harbor strips dates to year-only, removing temporal patterns that are critical for time-series models. It strips geographic data below the state level, eliminating spatial analysis. It strips ages above 89 into a single bucket, collapsing geriatric populations. Expert Determination is expensive -- a qualified statistical expert charges $50,000-150,000 per dataset evaluation -- and the determination is specific to the data release context, meaning you need a new evaluation for each use case. For more on de-identification approaches and their trade-offs for AI, see our healthcare data de-identification guide.
IRB Timelines and Institutional Friction
Academic medical centers require Institutional Review Board (IRB) approval for any research involving patient data, even retrospective chart review. Initial IRB review takes 4-8 weeks for expedited protocols and 8-16 weeks for full board review. Amendments -- which are inevitable as AI experiments evolve -- add 2-6 weeks each. Multi-site studies require separate IRB approval at each institution, or reliance on a central IRB with institution-specific addenda. An AI team that needs to iterate quickly on training data composition is structurally incompatible with this timeline.
Small, Biased Datasets
Even when data access is secured, the resulting datasets are typically small, skewed toward the demographics of the source institution, and incomplete. A model trained on data from a single academic medical center in Boston will overrepresent commercially insured, English-speaking patients and underrepresent rural, Medicaid, and non-English-speaking populations. The FDA has explicitly flagged training data representativeness as a key concern in AI/ML device evaluations.
Synthetic data does not solve every problem -- models ultimately need validation against real clinical data. But it solves the development velocity problem. You can build, iterate, and stress-test your pipeline on synthetic data today, while the real data access processes run in parallel. When the real data arrives, your pipeline is ready.
What Is Synthea?
Synthea is an open-source synthetic patient generator developed by The MITRE Corporation, a federally funded R&D center. First released in 2017, Synthea simulates the medical history of synthetic patients from birth to death (or present day), generating clinically realistic records that include conditions, medications, procedures, encounters, observations, immunizations, and care plans.
Synthea is not a random data generator. It is a clinically modeled simulation engine. Each disease and condition is represented as a state machine -- a module that defines how a patient progresses through the lifecycle of a condition based on real epidemiological data. A patient with Type 2 Diabetes, for example, does not just get a diagnosis code slapped on their record. Synthea models the progression from prediabetes through diagnosis, medication escalation (metformin, then sulfonylureas, then insulin), complication development (retinopathy, nephropathy, neuropathy), and the associated lab value changes (HbA1c trending upward over years). The clinical trajectories are grounded in published treatment guidelines and population-level statistics.
How the Simulation Engine Works
Synthea's generation pipeline follows this sequence:
- Patient seed -- A new patient is created with demographics drawn from US Census data: name, gender, race/ethnicity, geographic location (down to the city level), and socioeconomic factors
- Lifecycle simulation -- The engine advances the patient through time, month by month, from birth. At each time step, every active disease module evaluates whether its state transitions fire based on the patient's current age, gender, existing conditions, and probabilistic triggers
- Encounter generation -- When a module triggers a clinical event (symptom onset, annual wellness visit, emergency), Synthea generates the appropriate encounter with the provider, facility, and payer context
- Clinical data attachment -- Each encounter generates the associated observations (vitals, labs), procedures, medication orders, and care plans that would occur in that clinical context
- Export -- The complete patient record is serialized into the configured output format: FHIR R4 Bundle, C-CDA document, CSV tables, or all three
The result is a complete, longitudinal patient record that looks like what you would extract from a real EHR -- not a random scatter of codes, but a coherent clinical narrative with temporal consistency, clinical plausibility, and proper coding.
Getting Started: Installation and First Run
Synthea is a Java application. You need Java 11 or higher and Git. Here is the complete setup from zero to generated FHIR data.
Prerequisites
# Check Java version (need 11+)
java -version
# If you don't have Java, install via Homebrew (macOS) or apt (Linux)
# macOS:
brew install openjdk@17
# Ubuntu/Debian:
sudo apt install openjdk-17-jdk
# Verify
java -version
# openjdk version "17.0.10" 2024-01-16
Clone and Build
# Clone the Synthea repository
git clone https://github.com/synthetichealth/synthea.git
cd synthea
# Build the project (uses Gradle wrapper, no separate Gradle install needed)
./gradlew build check test
# This takes 3-5 minutes on first run (downloads dependencies)
# You'll see "BUILD SUCCESSFUL" when complete
Generate Your First Patient Population
# Generate 100 patients in Massachusetts (default state)
./run_synthea -p 100
# Generate 500 patients in California
./run_synthea -p 500 California
# Generate 50 patients in a specific city
./run_synthea -p 50 Texas Austin
# Generate with a specific random seed (for reproducibility)
./run_synthea -p 100 -s 12345 Massachusetts Boston
The -p flag sets the population size, and the -s flag sets the random seed for reproducible generation. When you need the same dataset across team members or CI runs, fix the seed.
Examine the Output
# Default output goes to ./output/fhir/
ls -la output/fhir/ | head -20
# Each file is a FHIR Bundle for one patient
# File naming: {UUID}.json
# -rw-r--r-- 1 user staff 245K Apr 2 10:15 0a1b2c3d-4e5f-6789-abcd-ef0123456789.json
# -rw-r--r-- 1 user staff 189K Apr 2 10:15 1b2c3d4e-5f6a-789b-cdef-0123456789ab.json
# Check total file count
ls output/fhir/*.json | wc -l
# 100
# Examine a single patient bundle
cat output/fhir/$(ls output/fhir/ | head -1) | python3 -m json.tool | head -50
# Count resource types in a bundle
cat output/fhir/$(ls output/fhir/ | head -1) | \
python3 -c "import json,sys,collections; \
d=json.load(sys.stdin); \
c=collections.Counter(e['resource']['resourceType'] for e in d['entry']); \
[print(f'{v:4d} {k}') for k,v in c.most_common()]"
A typical Synthea patient bundle contains 50-500 resources depending on the patient's age and clinical complexity. A 70-year-old patient with diabetes, hypertension, and heart failure will generate significantly more resources than a healthy 25-year-old.
Synthea's Disease Module Architecture
The clinical realism of Synthea comes from its modular disease simulation system. Each clinical condition is modeled as an independent Generic Module Framework (GMF) -- a finite state machine defined in a JSON document that specifies states, transitions, and clinical actions.
As of the latest release, Synthea ships with over 90 disease and wellness modules covering:
- Cardiovascular -- Atrial fibrillation, coronary heart disease, heart failure, stroke, hypertension
- Endocrine -- Type 1 and Type 2 diabetes (with full complication modeling), hypothyroidism, metabolic syndrome
- Respiratory -- Asthma, COPD, pneumonia, COVID-19, lung cancer
- Oncology -- Breast cancer, colorectal cancer, lung cancer (with TNM staging, treatment protocols, and survival curves)
- Orthopedic -- Osteoarthritis, osteoporosis, fractures
- Mental health -- Depression, anxiety, opioid use disorder
- Wellness -- Immunization schedules, preventive screenings, annual wellness visits
- Pediatric -- Childhood growth, ear infections, allergies
Module State Machine Structure
Each module defines a sequence of states that a patient transitions through. Here is a simplified view of the Type 2 Diabetes module's state machine:
{
"name": "Diabetes Type 2",
"states": {
"Initial": {
"type": "Initial",
"direct_transition": "Age Guard"
},
"Age Guard": {
"type": "Guard",
"allow": {
"condition_type": "Age",
"operator": ">=",
"quantity": 18,
"unit": "years"
},
"direct_transition": "Diabetes Prevalence Check"
},
"Diabetes Prevalence Check": {
"type": "Simple",
"complex_transition": [
{
"condition": {
"condition_type": "Attribute",
"attribute": "diabetes_severity",
"operator": "is not nil"
},
"transition": "Terminal"
},
{
"distributions": [
{"transition": "Prediabetes Onset", "distribution": 0.33},
{"transition": "No Diabetes", "distribution": 0.67}
]
}
]
},
"Prediabetes Onset": {
"type": "ConditionOnset",
"codes": [
{
"system": "SNOMED-CT",
"code": "15777000",
"display": "Prediabetes"
}
],
"direct_transition": "Prediabetes_HbA1c"
},
"Prediabetes_HbA1c": {
"type": "Observation",
"category": "laboratory",
"unit": "%",
"codes": [
{
"system": "LOINC",
"code": "4548-4",
"display": "Hemoglobin A1c/Hemoglobin.total in Blood"
}
],
"range": {"low": 5.7, "high": 6.4},
"direct_transition": "Diabetes Progression Delay"
},
"Diabetes Progression Delay": {
"type": "Delay",
"range": {"low": 2, "high": 10, "unit": "years"},
"direct_transition": "Diabetes Onset"
},
"Diabetes Onset": {
"type": "ConditionOnset",
"codes": [
{
"system": "SNOMED-CT",
"code": "44054006",
"display": "Diabetes mellitus type 2"
}
],
"direct_transition": "Metformin Prescription"
}
}
}
The key design principle is that transitions are probabilistic and condition-dependent. A patient does not automatically progress from prediabetes to diabetes -- there is a delay of 2-10 years with probability-weighted transitions. This produces the kind of clinical variation you see in real populations. Some patients progress quickly. Some stabilize. Some develop complications early while others do not. The distributions are calibrated against published epidemiological data from sources like the CDC's National Diabetes Statistics Report.
Creating Custom Modules
You can create custom modules using the Synthea Module Builder, a visual drag-and-drop editor that generates the JSON module definition. This is particularly valuable when you need synthetic data for a condition or workflow that Synthea does not cover out of the box -- rare diseases, custom clinical protocols, or organization-specific care pathways.
To add a custom module, place the JSON file in src/main/resources/modules/ and rebuild:
# Add your custom module
cp my_custom_condition.json synthea/src/main/resources/modules/
# Rebuild
cd synthea
./gradlew build
# Generate patients -- your module will be included in the simulation
./run_synthea -p 100
Output Formats: FHIR R4, C-CDA, and CSV
Synthea supports three primary output formats, configurable via src/main/resources/synthea.properties:
# Export configuration in synthea.properties
# FHIR R4 (default: enabled)
exporter.fhir.export = true
# C-CDA (default: disabled)
exporter.ccda.export = false
# CSV (default: disabled)
exporter.csv.export = false
# Hospital FHIR export (bundled by facility)
exporter.hospital.fhir.export = false
# Practitioner FHIR export
exporter.practitioner.fhir.export = false
# Output directory
exporter.baseDirectory = ./output/
FHIR R4 Bundle Structure
The primary output format -- and the one most useful for AI pipelines -- is FHIR R4. Each patient generates a single Bundle of type transaction, containing all resources for that patient's complete medical history. Here is an abbreviated example showing the structure:
{
"resourceType": "Bundle",
"type": "transaction",
"entry": [
{
"fullUrl": "urn:uuid:7a3d8e2f-1b4c-5d6e-a7f8-9012345abcde",
"resource": {
"resourceType": "Patient",
"id": "7a3d8e2f-1b4c-5d6e-a7f8-9012345abcde",
"name": [
{
"use": "official",
"family": "Rodriguez",
"given": ["Maria", "Elena"],
"prefix": ["Mrs."]
}
],
"gender": "female",
"birthDate": "1958-03-14",
"address": [
{
"line": ["742 Evergreen Terrace"],
"city": "Springfield",
"state": "Massachusetts",
"postalCode": "01103",
"country": "US"
}
],
"maritalStatus": {
"coding": [
{
"system": "http://terminology.hl7.org/CodeSystem/v3-MaritalStatus",
"code": "M",
"display": "Married"
}
]
},
"extension": [
{
"url": "http://hl7.org/fhir/us/core/StructureDefinition/us-core-race",
"extension": [
{
"url": "ombCategory",
"valueCoding": {
"system": "urn:oid:2.16.840.1.113883.6.238",
"code": "2106-3",
"display": "White"
}
}
]
},
{
"url": "http://hl7.org/fhir/us/core/StructureDefinition/us-core-ethnicity",
"extension": [
{
"url": "ombCategory",
"valueCoding": {
"system": "urn:oid:2.16.840.1.113883.6.238",
"code": "2135-2",
"display": "Hispanic or Latino"
}
}
]
}
]
},
"request": {
"method": "POST",
"url": "Patient"
}
},
{
"fullUrl": "urn:uuid:condition-diabetes-001",
"resource": {
"resourceType": "Condition",
"clinicalStatus": {
"coding": [
{
"system": "http://terminology.hl7.org/CodeSystem/condition-clinical",
"code": "active"
}
]
},
"verificationStatus": {
"coding": [
{
"system": "http://terminology.hl7.org/CodeSystem/condition-ver-status",
"code": "confirmed"
}
]
},
"code": {
"coding": [
{
"system": "http://snomed.info/sct",
"code": "44054006",
"display": "Diabetes mellitus type 2"
}
]
},
"subject": {
"reference": "urn:uuid:7a3d8e2f-1b4c-5d6e-a7f8-9012345abcde"
},
"onsetDateTime": "2012-07-22T10:30:00-04:00",
"recordedDate": "2012-07-22T10:30:00-04:00"
},
"request": {
"method": "POST",
"url": "Condition"
}
},
{
"fullUrl": "urn:uuid:obs-hba1c-001",
"resource": {
"resourceType": "Observation",
"status": "final",
"category": [
{
"coding": [
{
"system": "http://terminology.hl7.org/CodeSystem/observation-category",
"code": "laboratory"
}
]
}
],
"code": {
"coding": [
{
"system": "http://loinc.org",
"code": "4548-4",
"display": "Hemoglobin A1c/Hemoglobin.total in Blood"
}
]
},
"subject": {
"reference": "urn:uuid:7a3d8e2f-1b4c-5d6e-a7f8-9012345abcde"
},
"effectiveDateTime": "2025-01-15T09:00:00-04:00",
"valueQuantity": {
"value": 7.8,
"unit": "%",
"system": "http://unitsofmeasure.org",
"code": "%"
}
},
"request": {
"method": "POST",
"url": "Observation"
}
}
]
}
Key characteristics of Synthea's FHIR output that matter for AI/ML pipelines:
- US Core conformance -- Resources include US Core extensions for race, ethnicity, and birth sex, making the data compatible with US Core FHIR server validation
- Standard terminology -- All codes use SNOMED CT, LOINC, RxNorm, ICD-10-CM, and CPT, matching what production EHR systems emit
- Transaction bundles -- Each patient is a single transaction Bundle, ready to POST directly to a FHIR server's transaction endpoint
- Cross-references -- Resources reference each other via UUID URNs, maintaining referential integrity within the bundle
- Temporal consistency -- Observations, encounters, and conditions are properly ordered in time, with realistic intervals between events
CSV Output for Tabular ML
For teams working with traditional ML models (XGBoost, random forests, logistic regression) rather than FHIR-native pipelines, Synthea's CSV output provides pre-flattened tables:
# Enable CSV export
# In synthea.properties:
# exporter.csv.export = true
# After generation, CSV files appear in output/csv/
ls output/csv/
# allergies.csv conditions.csv encounters.csv
# imaging_studies.csv immunizations.csv medications.csv
# observations.csv organizations.csv patients.csv
# payer_transitions.csv payers.csv procedures.csv
# providers.csv supplies.csv
# Examine the patients table
head -5 output/csv/patients.csv
# Id,BIRTHDATE,DEATHDATE,SSN,DRIVERS,PASSPORT,PREFIX,FIRST,LAST,...
# 7a3d8e2f...,1958-03-14,,999-12-3456,S99912345,,Mrs.,Maria,Rodriguez,...
# Count observations
wc -l output/csv/observations.csv
# 247,832 observations.csv
The CSV tables use the patient ID as a foreign key across all tables, making joins straightforward for feature engineering in pandas or SQL.
Configuring Population Parameters
Synthea's default generation produces a population that mirrors Massachusetts demographics. For AI training, you almost always need to customize the population to match your target deployment environment or to create specific clinical distributions. All configuration happens in synthea.properties.
Geographic and Demographic Control
# Generate for a specific state (overrides command-line if set)
generate.default_state = California
# Age distribution
generate.demographics.default_file = geography/demographics.csv
# Gender ratio (default: 0.5)
generate.demographics.gender.male = 0.50
# Population size
generate.default_population = 10000
Synthea ships with demographic data for all 50 US states plus territories, sourced from US Census Bureau data. The demographics files control the distribution of age, gender, race/ethnicity, income, and education level for each geographic area.
Controlling Disease Prevalence
You can override module prevalence rates to create datasets enriched for specific conditions -- critical when training models for rare disease detection or when you need balanced classes:
# Override diabetes prevalence (default follows CDC rates ~10%)
# Increase to 30% for a diabetes-focused AI model
generate.modules.diabetes.prevalence = 0.30
# Increase heart failure prevalence
generate.modules.congestive_heart_failure.prevalence = 0.15
# Control the overall chronic condition probability
generate.max_conditions = 10
Time Range Configuration
# Set the reference date for "today" in the simulation
# Default: current system date
generate.reference_date = 20260101
# Only include patients alive after this date
# Useful for filtering out historical-only records
generate.only_alive_patients = true
# Or generate patients who died (for mortality prediction models)
generate.only_alive_patients = false
generate.only_dead_patients = true
Insurance and Payer Mix
# Payer configuration
# Synthea models insurance transitions based on age and employment
# Patients get Medicare at 65, Medicaid based on income, etc.
# Custom payer file
generate.payers.default_file = payers/custom_payers.csv
# Include cost data in claims
exporter.cost_access_outcomes_report = true
The payer modeling is particularly useful for AI teams building cost prediction, utilization management, or revenue cycle automation models. Synthea generates realistic Claims and ExplanationOfBenefit resources with procedure costs, copays, and payer responsibilities.
Loading Synthea Data into FHIR Servers
Generating synthetic data is step one. Loading it into a FHIR server where your AI pipeline can query it via standard FHIR APIs is step two. Synthea's transaction Bundle format makes this straightforward -- each patient Bundle is a valid FHIR transaction that any compliant server will accept. For a detailed comparison of FHIR server options, see our FHIR data store comparison guide.
HAPI FHIR Server
HAPI FHIR is the most widely used open-source FHIR server. You can spin it up in Docker and load Synthea data in minutes:
# Start HAPI FHIR server in Docker
docker run -d --name hapi-fhir \
-p 8080:8080 \
-e hapi.fhir.default_encoding=json \
-e hapi.fhir.fhir_version=R4 \
hapiproject/hapi:latest
# Wait for startup
sleep 15
# Verify the server is running
curl -s http://localhost:8080/fhir/metadata | \
python3 -c "import json,sys; d=json.load(sys.stdin); \
print(f'FHIR {d[\"fhirVersion\"]} — {d[\"implementation\"][\"description\"]}')"
# FHIR 4.0.1 — HAPI FHIR R4 Server
# Load a single patient bundle
curl -X POST http://localhost:8080/fhir \
-H "Content-Type: application/fhir+json" \
-d @output/fhir/$(ls output/fhir/ | head -1)
# Bulk load all patient bundles
for f in output/fhir/*.json; do
echo "Loading: $f"
curl -s -X POST http://localhost:8080/fhir \
-H "Content-Type: application/fhir+json" \
-d @"$f" > /dev/null
done
# Verify patient count
curl -s "http://localhost:8080/fhir/Patient?_summary=count" | \
python3 -c "import json,sys; print(json.load(sys.stdin)['total'])"
# 100
Parallel Bulk Loading Script
For larger populations (1,000+ patients), sequential loading is slow. Here is a Python script that loads bundles in parallel:
#!/usr/bin/env python3
"""bulk_load_synthea.py — Load Synthea FHIR Bundles into a FHIR server in parallel."""
import json
import sys
import os
import glob
import concurrent.futures
import requests
from pathlib import Path
FHIR_BASE = os.getenv("FHIR_BASE", "http://localhost:8080/fhir")
WORKERS = int(os.getenv("WORKERS", "8"))
SYNTHEA_DIR = sys.argv[1] if len(sys.argv) > 1 else "./output/fhir"
def load_bundle(filepath: str) -> dict:
"""POST a single FHIR transaction Bundle to the server."""
with open(filepath, "r") as f:
bundle = json.load(f)
resp = requests.post(
FHIR_BASE,
json=bundle,
headers={"Content-Type": "application/fhir+json"},
timeout=120,
)
patient_name = "unknown"
for entry in bundle.get("entry", []):
res = entry.get("resource", {})
if res.get("resourceType") == "Patient":
names = res.get("name", [{}])
if names:
given = " ".join(names[0].get("given", []))
family = names[0].get("family", "")
patient_name = f"{given} {family}".strip()
break
return {
"file": os.path.basename(filepath),
"patient": patient_name,
"status": resp.status_code,
"resources": len(bundle.get("entry", [])),
}
def main():
files = sorted(glob.glob(os.path.join(SYNTHEA_DIR, "*.json")))
print(f"Loading {len(files)} bundles into {FHIR_BASE} with {WORKERS} workers...")
success, failed = 0, 0
total_resources = 0
with concurrent.futures.ThreadPoolExecutor(max_workers=WORKERS) as executor:
futures = {executor.submit(load_bundle, f): f for f in files}
for future in concurrent.futures.as_completed(futures):
result = future.result()
if result["status"] == 200:
success += 1
total_resources += result["resources"]
print(f" OK: {result['patient']} ({result['resources']} resources)")
else:
failed += 1
print(f" FAIL [{result['status']}]: {result['file']}")
print(f"\nComplete: {success} loaded, {failed} failed, {total_resources} total resources")
if __name__ == "__main__":
main()
# Run the bulk loader
pip install requests
python3 bulk_load_synthea.py ./output/fhir/
# Loading 1000 bundles into http://localhost:8080/fhir with 8 workers...
# OK: Maria Rodriguez (247 resources)
# OK: James Chen (183 resources)
# ...
# Complete: 1000 loaded, 0 failed, 215,432 total resources
Google Cloud Healthcare API
For cloud-native deployments, Google Cloud Healthcare API provides a managed FHIR R4 store. For a complete guide to setting up and managing Google Cloud FHIR stores, see our Google Cloud Healthcare API production guide.
# Set variables
PROJECT_ID="your-gcp-project"
LOCATION="us-central1"
DATASET_ID="synthea-training-data"
FHIR_STORE_ID="synthea-r4"
# Create the dataset and FHIR store
gcloud healthcare datasets create $DATASET_ID \
--location=$LOCATION
gcloud healthcare fhir-stores create $FHIR_STORE_ID \
--dataset=$DATASET_ID \
--location=$LOCATION \
--version=R4
# Import Synthea data from a GCS bucket
# First, upload the FHIR bundles to GCS
gsutil -m cp output/fhir/*.json gs://$PROJECT_ID-synthea-data/fhir/
# Then import via the Healthcare API's bulk import
gcloud healthcare fhir-stores import gcs $FHIR_STORE_ID \
--dataset=$DATASET_ID \
--location=$LOCATION \
--gcs-uri=gs://$PROJECT_ID-synthea-data/fhir/*.json \
--content-structure=BUNDLE
# Verify
curl -s -H "Authorization: Bearer $(gcloud auth print-access-token)" \
"https://healthcare.googleapis.com/v1/projects/$PROJECT_ID/locations/$LOCATION/datasets/$DATASET_ID/fhirStores/$FHIR_STORE_ID/fhir/Patient?_summary=count"
Azure Health Data Services
# Azure FHIR service setup
az healthcareapis service create \
--resource-group synthea-rg \
--resource-name synthea-fhir \
--kind fhir-R4 \
--location eastus \
--access-policies-object-id $(az ad signed-in-user show --query id -o tsv)
FHIR_URL=$(az healthcareapis service show \
--resource-group synthea-rg \
--resource-name synthea-fhir \
--query "properties.authenticationConfiguration.audience" -o tsv)
# Get access token and load
TOKEN=$(az account get-access-token \
--resource $FHIR_URL --query accessToken -o tsv)
for f in output/fhir/*.json; do
curl -s -X POST "$FHIR_URL" \
-H "Authorization: Bearer $TOKEN" \
-H "Content-Type: application/fhir+json" \
-d @"$f" > /dev/null
echo "Loaded: $(basename $f)"
done
Using Synthetic Data for AI/ML Training
With Synthea data loaded into a FHIR server (or available as local JSON/CSV files), the next step is building training datasets for your ML models. The approach differs depending on whether you are working with structured ML models or LLM-based systems.
Feature Engineering from FHIR Resources
For classical ML models (gradient boosting, neural networks, logistic regression), you need to transform FHIR resources into flat feature vectors. Here is a practical example -- building a dataset for a hospital readmission prediction model:
#!/usr/bin/env python3
"""extract_readmission_features.py — Build readmission prediction dataset from Synthea FHIR."""
import json
import glob
import pandas as pd
from datetime import datetime, timedelta
from collections import defaultdict
def parse_fhir_date(date_str: str) -> datetime:
"""Parse FHIR datetime formats."""
for fmt in ["%Y-%m-%dT%H:%M:%S%z", "%Y-%m-%dT%H:%M:%S", "%Y-%m-%d"]:
try:
return datetime.strptime(date_str[:19], fmt[:len(date_str[:19])+2].rstrip('%z'))
except ValueError:
continue
return None
def extract_patient_features(bundle: dict) -> list[dict]:
"""Extract per-encounter features from a FHIR Bundle."""
resources_by_type = defaultdict(list)
for entry in bundle.get("entry", []):
res = entry.get("resource", {})
resources_by_type[res["resourceType"]].append(res)
patient = resources_by_type["Patient"][0]
birth_date = datetime.strptime(patient["birthDate"], "%Y-%m-%d")
# Extract active conditions as of each encounter
conditions = set()
for cond in resources_by_type.get("Condition", []):
code = cond.get("code", {}).get("coding", [{}])[0].get("code", "")
if code:
conditions.add(code)
# Extract encounter features
encounters = sorted(
resources_by_type.get("Encounter", []),
key=lambda e: e.get("period", {}).get("start", "")
)
rows = []
for i, enc in enumerate(encounters):
enc_class = enc.get("class", {}).get("code", "unknown")
if enc_class != "inpatient":
continue
start = parse_fhir_date(enc.get("period", {}).get("start", ""))
end = parse_fhir_date(enc.get("period", {}).get("end", ""))
if not start or not end:
continue
age_at_encounter = (start - birth_date).days / 365.25
los_days = (end - start).days
# Count medications active at time of encounter
active_meds = sum(
1 for med in resources_by_type.get("MedicationRequest", [])
if med.get("authoredOn", "") <= start.isoformat()
)
# Get latest lab values before this encounter
latest_labs = {}
for obs in resources_by_type.get("Observation", []):
obs_date = obs.get("effectiveDateTime", "")
if obs_date and obs_date <= start.isoformat():
code = obs.get("code", {}).get("coding", [{}])[0].get("code", "")
value = obs.get("valueQuantity", {}).get("value")
if code and value is not None:
if code not in latest_labs or obs_date > latest_labs[code][0]:
latest_labs[code] = (obs_date, value)
# Check for 30-day readmission (label)
readmitted_30d = False
for next_enc in encounters[i+1:]:
next_start = parse_fhir_date(next_enc.get("period", {}).get("start", ""))
if next_start and next_enc.get("class", {}).get("code") == "inpatient":
if (next_start - end).days <= 30:
readmitted_30d = True
break
rows.append({
"patient_id": patient.get("id"),
"age": round(age_at_encounter, 1),
"gender": patient.get("gender", "unknown"),
"los_days": los_days,
"num_conditions": len(conditions),
"num_active_meds": active_meds,
"hba1c": latest_labs.get("4548-4", (None, None))[1], # LOINC: HbA1c
"creatinine": latest_labs.get("2160-0", (None, None))[1], # LOINC: Creatinine
"systolic_bp": latest_labs.get("8480-6", (None, None))[1], # LOINC: Systolic BP
"bmi": latest_labs.get("39156-5", (None, None))[1], # LOINC: BMI
"readmitted_30d": int(readmitted_30d),
})
return rows
# Process all Synthea bundles
all_rows = []
for filepath in glob.glob("./output/fhir/*.json"):
with open(filepath) as f:
bundle = json.load(f)
all_rows.extend(extract_patient_features(bundle))
df = pd.DataFrame(all_rows)
print(f"Dataset shape: {df.shape}")
print(f"Readmission rate: {df['readmitted_30d'].mean():.1%}")
print(f"\nFeature summary:")
print(df.describe().round(2))
df.to_csv("readmission_training_data.csv", index=False)
Building Training Datasets for LLMs
For LLM-based clinical AI -- summarization, question answering, clinical decision support -- the approach shifts from tabular feature extraction to prompt construction from FHIR resources:
def fhir_to_clinical_prompt(bundle: dict) -> str:
"""Convert a FHIR Bundle into a clinical summary prompt for LLM training."""
resources = defaultdict(list)
for entry in bundle.get("entry", []):
res = entry["resource"]
resources[res["resourceType"]].append(res)
patient = resources["Patient"][0]
name = patient["name"][0]
age = 2026 - int(patient["birthDate"][:4])
# Active conditions
conditions = []
for c in resources.get("Condition", []):
status = c.get("clinicalStatus", {}).get("coding", [{}])[0].get("code")
if status == "active":
display = c["code"]["coding"][0]["display"]
onset = c.get("onsetDateTime", "unknown")[:10]
conditions.append(f"- {display} (onset: {onset})")
# Current medications
meds = []
for m in resources.get("MedicationRequest", []):
if m.get("status") == "active":
display = m.get("medicationCodeableConcept", {}).get("coding", [{}])[0].get("display", "Unknown")
meds.append(f"- {display}")
# Recent labs (last 6 months)
recent_labs = []
for obs in resources.get("Observation", []):
if obs.get("category", [{}])[0].get("coding", [{}])[0].get("code") == "laboratory":
value = obs.get("valueQuantity", {})
if value:
lab_name = obs["code"]["coding"][0]["display"]
recent_labs.append(
f"- {lab_name}: {value['value']} {value.get('unit', '')}"
)
prompt = f"""Patient: {name.get('given', [''])[0]} {name.get('family', '')}, {age}yo {patient['gender']}
Active Conditions:
{chr(10).join(conditions[:15]) or '- None documented'}
Current Medications:
{chr(10).join(meds[:10]) or '- None documented'}
Recent Laboratory Results:
{chr(10).join(recent_labs[:20]) or '- None available'}"""
return prompt
This approach lets you build evaluation datasets for LLM output quality testing at scale -- generating thousands of synthetic clinical summaries with known ground truth conditions and medications, then measuring whether your LLM correctly identifies them.
Validation: How Close Is Synthetic Data to Real Data?
The critical question for any AI team using synthetic data: how well do models trained on Synthea data transfer to real clinical data? The research is encouraging but nuanced.
A study published in JAMIA (Journal of the American Medical Informatics Association) compared machine learning models trained on Synthea-generated data against models trained on real EHR data for predicting emergency department utilization. Key findings:
- Models trained on Synthea data achieved AUC scores within 5-12% of models trained on real data for most prediction tasks
- The gap was smallest for conditions where Synthea's disease modules are most detailed (diabetes, cardiovascular disease)
- The gap was largest for conditions with complex social determinants that Synthea does not fully model
- Synthetic-to-real transfer improved significantly when the synthetic population was configured to match the target population's demographics
The practical implication: Synthea data is excellent for pipeline development, integration testing, and initial model architecture exploration. It is not a replacement for real data validation before production deployment. The recommended workflow is:
- Develop on synthetic -- Build your entire pipeline end-to-end using Synthea data. Debug data ingestion, feature engineering, model training, and inference serving.
- Validate on real -- Once the pipeline works, swap in real (de-identified) data for final validation. The pipeline code should not change -- only the data source.
- Monitor drift -- In production, compare model performance metrics against the synthetic baseline to detect distribution shift.
Privacy and Compliance Advantages
The compliance argument for synthetic data is unambiguous. Synthea-generated records contain no real patient information -- they are entirely fabricated from statistical distributions. This eliminates entire categories of regulatory burden. For a deeper analysis of de-identification methods and their limitations, see our healthcare data de-identification guide.
| Dimension | Synthetic Data (Synthea) | De-Identified Data (Safe Harbor) | De-Identified Data (Expert Determination) | Real PHI |
|---|---|---|---|---|
| Contains PHI | No -- never contained real data | Residual risk -- 18 identifiers removed but correlations remain | Low risk -- statistically validated | Yes -- full PHI |
| BAA required | No | Debated -- many orgs require one | Debated -- context-dependent | Yes -- mandatory |
| IRB required | No | Usually exempt or expedited | Usually exempt or expedited | Yes -- full board review |
| Shareable externally | Yes -- freely distributable | With DUA -- restrictions apply | With DUA -- restrictions apply | No -- covered entity only |
| Re-identification risk | Zero -- no real individuals | Non-zero -- linkage attacks possible | Quantified -- typically <0.04% | N/A -- already identified |
| Time to access | Minutes (generate on demand) | Weeks to months | Months (expert engagement) | 3-9 months (legal + compliance) |
| Cost | Free (compute only) | $10K-50K (process + audit) | $50K-150K (expert + audit) | $100K+ (legal + infrastructure) |
| Dataset size control | Unlimited -- generate any size | Limited to source data | Limited to source data | Limited to source data |
| Demographic control | Full -- configurable distributions | None -- reflects source bias | None -- reflects source bias | None -- reflects source bias |
| Reproducibility | Exact -- seed-based generation | Fixed -- one-time extract | Fixed -- one-time extract | Variable -- live system |
Regulatory Context
The regulatory landscape increasingly favors synthetic data. The FDA's guidance on AI/ML-enabled device software functions acknowledges the use of synthetic and augmented data in training datasets, provided that the final validation uses representative real-world data. The ONC's interoperability rules promote the use of synthetic data for testing FHIR API implementations. CMS uses Synthea-generated data in its own Blue Button 2.0 sandbox for developer testing.
For teams building healthcare data lakes and analytics infrastructure, synthetic data enables you to build and test the entire pipeline -- from ingestion to transformation to serving -- without ever touching PHI. See our healthcare data lake architecture guide for the infrastructure patterns.
Frequently Asked Questions
Is Synthea data HIPAA-compliant?
Synthea data does not fall under HIPAA because it contains no protected health information. The records are entirely synthetic -- generated from statistical distributions, not derived from real patient records. There is no real person behind a Synthea record, so HIPAA's privacy protections do not apply. You can share Synthea data freely, store it without a BAA, and use it without encryption requirements (though encryption is still good practice). The HHS de-identification guidance explicitly distinguishes synthetic data from de-identified data -- synthetic data was never identified to begin with.
Can I use Synthea data to train production AI models?
You can and should use Synthea data for pipeline development, architecture validation, and initial model training. However, models destined for clinical deployment must be validated against real (de-identified or IRB-approved) clinical data before production use. The FDA expects validation on data representative of the target population. Synthea data is ideal for the development and testing phases that precede real-data validation.
How clinically accurate is Synthea's data?
Synthea's disease modules are calibrated against published epidemiological data from the CDC, NIH, and peer-reviewed literature. Prevalence rates, complication frequencies, and treatment progressions are grounded in real statistics. However, Synthea cannot capture every clinical nuance -- rare disease presentations and institution-specific practice patterns may not be fully represented. For common conditions like diabetes, hypertension, and COPD, the clinical trajectories are highly realistic. For less common conditions, you may need custom modules.
How large a population can Synthea generate?
Synthea can generate populations of any size, limited only by compute and storage. On a modern laptop, expect approximately 100-200 patients per minute. For large populations (100,000+), run on a server or in CI/CD. A 100,000-patient population generates approximately 20-30 GB of FHIR JSON and takes 8-16 hours on a single machine. You can parallelize by running multiple Synthea instances with different random seeds and geographic regions, then merging the output.
Does Synthea support non-US populations?
Synthea's default configuration is US-centric: US Census demographics, US Core FHIR profiles, US insurance models, and US treatment guidelines. However, the Synthea International project has extended support to several other countries including Canada, the UK, Australia, Finland, and India, with country-specific demographics, healthcare systems, and terminology adaptations. Community contributions continue to expand international support.
Can I modify Synthea's output after generation?
Yes. Synthea's FHIR output is standard JSON. You can post-process bundles with any programming language to add custom extensions, modify coding systems, inject additional resources, or transform the data structure. Common post-processing steps include adding organization-specific identifiers, converting terminology systems (e.g., mapping SNOMED CT to ICD-10 for billing use cases), and filtering resources to match a specific FHIR Implementation Guide profile.
Getting Started Today
The path from zero to a working synthetic data pipeline takes less than a day. Here is the recommended sequence for a team starting from scratch:
- Hour 1: Generate -- Clone Synthea, build, and generate your first 100-patient population. Examine the FHIR output to understand the resource structure.
- Hour 2: Load -- Spin up a HAPI FHIR server in Docker and load your synthetic patients. Verify you can query patients, conditions, and observations via the FHIR REST API.
- Hour 3: Extract -- Write a feature extraction script that pulls the clinical data your model needs from the FHIR server. Build your first training dataset as a CSV or DataFrame.
- Hour 4: Configure -- Customize Synthea's population parameters to match your target deployment: demographics, disease prevalence, population size. Regenerate and reload.
- Day 2+: Iterate -- Integrate Synthea generation into your CI/CD pipeline for reproducible dataset creation. Build out your ML training pipeline against the synthetic data while real data access processes proceed in parallel.
The organizations shipping healthcare AI at scale -- the ones with models in production, not just in demos -- all share one trait: they solved the data pipeline problem before they solved the model problem. Synthetic data with Synthea is how you solve the data pipeline problem without waiting months for PHI access.
Every week you spend waiting for data access is a week your competitors are iterating. Generate the data. Build the pipeline. Ship.
If your team needs help building FHIR-native AI pipelines, integrating Synthea into your development workflow, or standing up the interoperability infrastructure that production healthcare AI requires, talk to our team. We build healthcare data platforms that are AI-ready from day one.