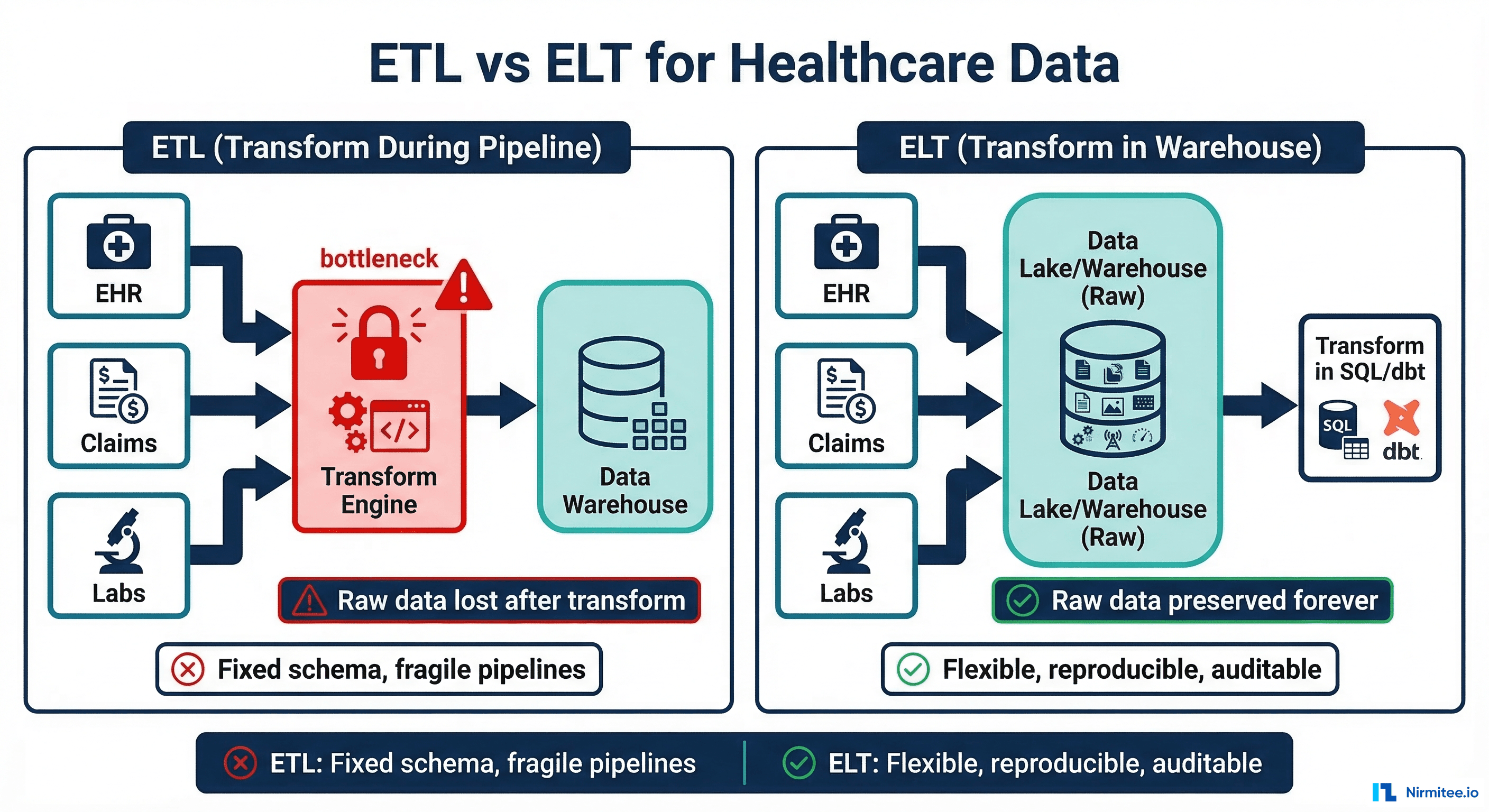
Healthcare data teams face a fundamental architectural decision: should you transform data during the pipeline (ETL) or after loading it into the warehouse (ELT)? For decades, ETL was the default. Extract data from the EHR, transform it in a separate engine like Informatica or Talend, then load the transformed result into the warehouse. But this approach has a fatal flaw for healthcare: it destroys raw data.
When a CMS auditor asks you to prove what a patient record looked like on a specific date, and your ETL pipeline discarded the original FHIR resource after transforming it, you have a compliance problem. ELT (Extract, Load, Transform) solves this by loading raw data first and transforming it inside the warehouse, preserving the originals indefinitely. Combined with dbt for SQL-based transformations and Great Expectations for data quality validation, ELT gives healthcare organizations reproducible, auditable, and flexible data pipelines.
ETL: The Traditional Approach and Its Healthcare Limitations
ETL (Extract, Transform, Load) follows a straightforward pattern: extract data from source systems, transform it in a dedicated processing engine, and load the result into a data warehouse. This approach made sense when warehouse compute was expensive and storage was limited. Transform early to minimize what you store.
How ETL Works in Healthcare
# Traditional ETL: Transform happens BEFORE loading
# Problem: raw FHIR data is lost after transformation
def etl_patient_pipeline():
# 1. EXTRACT from EHR FHIR API
raw_patients = fhir_client.search("Patient", count=1000)
# 2. TRANSFORM in ETL engine (Informatica, Talend, custom)
transformed = []
for patient in raw_patients:
transformed.append({
"patient_id": patient["id"],
"first_name": patient["name"][0]["given"][0],
"last_name": patient["name"][0]["family"],
"dob": patient["birthDate"],
"gender": patient["gender"],
# Extensions, metadata, original structure: LOST
# FHIR meta.versionId: LOST
# FHIR meta.lastUpdated: LOST
# US Core race/ethnicity extensions: LOST
})
# 3. LOAD transformed data into warehouse
warehouse.insert("dim_patient", transformed)
# Raw FHIR resources are gone foreverWhy ETL Fails Healthcare
The problems with ETL in healthcare are not theoretical. They cause real compliance failures and analytics limitations:
- Raw data loss: HIPAA requires organizations to maintain original records. ETL pipelines that discard raw data after transformation create compliance gaps. When an auditor asks for the original HL7v2 message or FHIR resource, it no longer exists.
- Brittle transformations: When Epic releases a new FHIR profile version and adds a field, the ETL pipeline breaks because the transform logic does not expect the new column. The entire pipeline stops until an engineer updates the transformation code, tests it, and redeploys.
- Non-reproducible results: If you discover a bug in your transform logic (a wrong ICD-10 mapping, an incorrect date format), you cannot fix it and re-run because the raw data is gone. You must re-extract from the source system, which may have changed since the original extraction.
- Coupled compute: The ETL engine (often a separate server) must scale with both data volume and transformation complexity. Adding a new transformation requires upgrading the ETL infrastructure.

ELT: Transform-Last Wins for Clinical Data
ELT (Extract, Load, Transform) flips the order. Extract data from source systems, load it raw into the warehouse or data lake, then transform it using the warehouse's own compute engine. The raw data stays forever. Transforms are SQL models that can be re-run, modified, and versioned.
How ELT Works in Healthcare
# ELT: Load raw data FIRST, transform in warehouse
# Raw FHIR data is preserved forever
def elt_patient_pipeline():
# 1. EXTRACT from EHR FHIR API
raw_patients = fhir_client.search("Patient", count=1000)
# 2. LOAD raw JSON directly into warehouse
warehouse.insert("raw_fhir_patients", [
{
"resource_id": p["id"],
"resource_type": "Patient",
"raw_json": json.dumps(p), # FULL resource preserved
"source_system": "epic_prod",
"extracted_at": datetime.utcnow().isoformat(),
"fhir_version": "4.0.1"
}
for p in raw_patients
])
# 3. TRANSFORM happens in warehouse via dbt/SQL
# Runs on schedule, independently of extraction
# Raw data always available for re-transformationELT Advantages for Healthcare: Detailed Breakdown
| Dimension | ETL | ELT | Healthcare Impact |
|---|---|---|---|
| Raw data retention | Discarded after transform | Preserved indefinitely | HIPAA audit compliance, original records always available |
| Schema changes | Pipeline rebuild required | Add new dbt model | FHIR profile updates handled in hours, not weeks |
| Transform bugs | Re-extract from source | Fix SQL, re-run on raw data | ICD-10 mapping errors fixable without source system access |
| Compute cost | ETL server + warehouse | Warehouse only | 30-50% infrastructure cost reduction |
| Time to new analytics | Weeks (ETL development cycle) | Hours (write SQL model) | Clinical researchers self-serve with SQL |
| Data lineage | Opaque ETL engine | Version-controlled SQL | Complete audit trail of every transformation |
| Testing | Manual QA of ETL output | Automated dbt tests | Data quality checks run on every pipeline execution |
For healthcare organizations already struggling with data quality issues, see our guide on healthcare data quality as a prerequisite for AI and analytics.

dbt for Healthcare: SQL-Based Transformations
dbt (data build tool) is the standard transformation layer for ELT architectures. It lets you write transforms as SQL SELECT statements, version them in Git, test them automatically, and generate documentation. For healthcare teams, dbt replaces the opaque ETL engine with transparent, auditable, version-controlled SQL.
Project Structure for Healthcare dbt
healthcare_analytics/
dbt_project.yml
models/
staging/
stg_fhir_patient.sql
stg_fhir_encounter.sql
stg_fhir_observation.sql
stg_fhir_condition.sql
stg_fhir_medication_request.sql
_stg_sources.yml # source definitions
_stg_models.yml # schema tests
intermediate/
int_patient_demographics.sql
int_encounter_timeline.sql
int_lab_results_pivoted.sql
int_active_medications.sql
marts/
mart_patient_360.sql
mart_quality_measures.sql
mart_readmission_risk.sql
mart_ed_utilization.sql
tests/
assert_no_orphaned_encounters.sql
assert_lab_values_physiological.sql
macros/
parse_fhir_reference.sql
extract_coding.sqlStaging Model: Parse FHIR Patient Resources
-- models/staging/stg_fhir_patient.sql
-- Parse raw FHIR Patient JSON into relational columns
-- This is the ONLY place we touch raw data
with raw_patients as (
select
resource_id,
raw_json,
extracted_at,
source_system
from {{ source('fhir_raw', 'raw_fhir_patients') }}
),
parsed as (
select
resource_id as patient_id,
raw_json:id::string as fhir_id,
raw_json:name[0].family::string as family_name,
raw_json:name[0].given[0]::string as given_name,
raw_json:birthDate::date as birth_date,
raw_json:gender::string as gender,
raw_json:address[0].line[0]::string as address_line,
raw_json:address[0].city::string as city,
raw_json:address[0].state::string as state,
raw_json:address[0].postalCode::string as postal_code,
raw_json:telecom[0].value::string as phone,
raw_json:meta.lastUpdated::timestamp as last_updated,
raw_json:meta.versionId::string as version_id,
extracted_at,
source_system
from raw_patients
)
select * from parsedStaging Model: Parse FHIR Encounter Resources
-- models/staging/stg_fhir_encounter.sql
with raw_encounters as (
select * from {{ source('fhir_raw', 'raw_fhir_encounters') }}
),
parsed as (
select
resource_id as encounter_id,
{{ parse_fhir_reference(
'raw_json:subject.reference'
) }} as patient_id,
raw_json:status::string as status,
raw_json:class.code::string as encounter_class,
raw_json:type[0].coding[0].code::string as type_code,
raw_json:type[0].coding[0].display::string as type_display,
raw_json:period.start::timestamp as period_start,
raw_json:period.end::timestamp as period_end,
datediff('hour',
raw_json:period.start::timestamp,
coalesce(
raw_json:period.end::timestamp,
current_timestamp()
)
) as length_of_stay_hours,
raw_json:hospitalization.dischargeDisposition
.coding[0].code::string as discharge_disposition,
extracted_at
from raw_encounters
)
select * from parsedStaging Model: Parse FHIR Observation (Lab Results)
-- models/staging/stg_fhir_observation.sql
with raw_observations as (
select * from {{ source('fhir_raw', 'raw_fhir_observations') }}
),
parsed as (
select
resource_id as observation_id,
{{ parse_fhir_reference(
'raw_json:subject.reference'
) }} as patient_id,
{{ parse_fhir_reference(
'raw_json:encounter.reference'
) }} as encounter_id,
raw_json:status::string as status,
{{ extract_coding(
'raw_json:code', 'http://loinc.org'
) }} as loinc_code,
raw_json:code.coding[0].display::string as display_name,
raw_json:valueQuantity.value::float as value_numeric,
raw_json:valueQuantity.unit::string as value_unit,
raw_json:valueString::string as value_string,
raw_json:effectiveDateTime::timestamp
as effective_datetime,
raw_json:referenceRange[0].low.value::float
as ref_range_low,
raw_json:referenceRange[0].high.value::float
as ref_range_high,
case
when raw_json:valueQuantity.value::float
< raw_json:referenceRange[0].low.value::float
then 'LOW'
when raw_json:valueQuantity.value::float
> raw_json:referenceRange[0].high.value::float
then 'HIGH'
else 'NORMAL'
end as interpretation,
extracted_at
from raw_observations
)
select * from parsedMart Model: Patient 360 View
-- models/marts/mart_patient_360.sql
-- Comprehensive patient view for clinical dashboards
with patients as (
select * from {{ ref('stg_fhir_patient') }}
),
encounter_summary as (
select
patient_id,
count(*) as total_encounters,
count(case when encounter_class = 'ER'
then 1 end) as ed_visits,
count(case when encounter_class = 'IMP'
then 1 end) as inpatient_stays,
max(period_start) as last_encounter_date,
avg(length_of_stay_hours) as avg_los_hours
from {{ ref('stg_fhir_encounter') }}
group by 1
),
lab_summary as (
select
patient_id,
count(*) as total_lab_results,
count(case when interpretation = 'HIGH'
or interpretation = 'LOW'
then 1 end) as abnormal_results,
max(effective_datetime) as last_lab_date
from {{ ref('stg_fhir_observation') }}
where status = 'final'
group by 1
)
select
p.patient_id,
p.given_name,
p.family_name,
p.birth_date,
p.gender,
p.city,
p.state,
e.total_encounters,
e.ed_visits,
e.inpatient_stays,
e.last_encounter_date,
e.avg_los_hours,
l.total_lab_results,
l.abnormal_results,
l.last_lab_date,
datediff('day', e.last_encounter_date,
current_date()) as days_since_last_visit
from patients p
left join encounter_summary e on p.patient_id = e.patient_id
left join lab_summary l on p.patient_id = l.patient_id
dbt Tests for Healthcare Data Quality
dbt tests run automatically on every pipeline execution. They catch data quality issues before bad data reaches dashboards or ML models. For healthcare, this means no lab results with impossible values, no encounters without patients, and no duplicate records reaching clinical reports.
Schema Tests (YAML)
# models/staging/_stg_models.yml
version: 2
models:
- name: stg_fhir_patient
description: "Parsed FHIR Patient resources"
columns:
- name: patient_id
description: "Unique patient identifier"
tests:
- not_null
- unique
- name: birth_date
tests:
- not_null
- name: gender
tests:
- accepted_values:
values: ["male", "female", "other", "unknown"]
- name: stg_fhir_encounter
description: "Parsed FHIR Encounter resources"
columns:
- name: encounter_id
tests:
- not_null
- unique
- name: patient_id
tests:
- not_null
- relationships:
to: ref('stg_fhir_patient')
field: patient_id
- name: status
tests:
- accepted_values:
values:
- planned
- arrived
- triaged
- in-progress
- onleave
- finished
- cancelled
- name: stg_fhir_observation
description: "Parsed FHIR Observation resources"
columns:
- name: observation_id
tests:
- not_null
- unique
- name: patient_id
tests:
- not_null
- relationships:
to: ref('stg_fhir_patient')
field: patient_idCustom Clinical Data Tests
-- tests/assert_birth_date_not_future.sql
-- Fail if any patient has a birth date in the future
select patient_id, birth_date
from {{ ref('stg_fhir_patient') }}
where birth_date > current_date()
-- tests/assert_discharge_after_admit.sql
-- Fail if discharge date is before admission date
select encounter_id, period_start, period_end
from {{ ref('stg_fhir_encounter') }}
where period_end is not null
and period_end < period_start
-- tests/assert_lab_values_physiological.sql
-- Fail if lab values are outside physiological range
select observation_id, loinc_code, value_numeric
from {{ ref('stg_fhir_observation') }}
where (
(loinc_code = '2345-7' and
(value_numeric < 0 or value_numeric > 2000))
or (loinc_code = '718-7' and
(value_numeric < 0 or value_numeric > 30))
or (loinc_code = '6298-4' and
(value_numeric < 0 or value_numeric > 15))
)
Great Expectations for Clinical Data Validation
While dbt tests validate transformed data, Great Expectations validates raw data before it enters the transformation layer. This catches source system issues (corrupt exports, schema changes, missing fields) before they propagate through the pipeline.
import great_expectations as gx
# Create healthcare data validation suite
context = gx.get_context()
# Define expectations for raw FHIR Patient data
patient_suite = context.add_expectation_suite(
"fhir_patient_raw_validation"
)
# Required fields must exist and be non-null
patient_suite.add_expectation(
gx.expectations.ExpectColumnToExist(column="resource_id")
)
patient_suite.add_expectation(
gx.expectations.ExpectColumnValuesToNotBeNull(
column="resource_id"
)
)
patient_suite.add_expectation(
gx.expectations.ExpectColumnValuesToNotBeNull(
column="raw_json"
)
)
# JSON structure validation
patient_suite.add_expectation(
gx.expectations.ExpectColumnValuesToMatchJsonSchema(
column="raw_json",
json_schema={
"type": "object",
"required": ["resourceType", "id", "name",
"birthDate", "gender"],
"properties": {
"resourceType": {
"type": "string",
"enum": ["Patient"]
},
"id": {"type": "string"},
"birthDate": {
"type": "string",
"pattern": "^\\d{4}-\\d{2}-\\d{2}$"
}
}
}
)
)
# Data freshness: fail if data is older than 6 hours
patient_suite.add_expectation(
gx.expectations.ExpectColumnMaxToBeBetween(
column="extracted_at",
min_value="{{current_time - 6h}}",
max_value="{{current_time}}"
)
)
# Run validation checkpoint
checkpoint = context.add_or_update_checkpoint(
name="fhir_patient_checkpoint",
validations=[
{"batch_request": patient_batch_request,
"expectation_suite_name": "fhir_patient_raw_validation"}
]
)
results = checkpoint.run()
if not results.success:
# Alert data engineering team
send_alert("FHIR Patient validation failed", results)
When to Use ETL vs ELT in Healthcare
While ELT wins for most modern healthcare data architectures, there are specific scenarios where ETL still makes sense:
| Scenario | Recommended Approach | Reasoning |
|---|---|---|
| Cloud-native data warehouse (Snowflake, BigQuery, Databricks) | ELT | Warehouse has abundant compute; dbt transforms are efficient |
| Legacy on-premise warehouse with limited compute | ETL | Warehouse cannot handle heavy transforms; pre-process to reduce load |
| FHIR data from multiple EHR vendors | ELT | Raw data preservation enables vendor-specific parsing in SQL |
| HL7v2 to FHIR transformation | ETL (at the edge) | Protocol translation is best done in Mirth Connect before loading |
| PHI de-identification before loading | ETL | De-identify before data reaches the warehouse for defense in depth |
| Real-time clinical dashboards | ELT + streaming | Stream raw events to warehouse, transform with materialized views |
| Regulatory reporting (CMS, HEDIS) | ELT | Reproducible transforms; re-run on corrected raw data |
For healthcare organizations handling HL7v2 to FHIR protocol translation, see our guide on building a FHIR facade over legacy HL7v2 systems. For organizations evaluating FHIR implementation readiness, check our 47-item FHIR implementation checklist.

dbt Macros for FHIR Data
FHIR resources follow consistent patterns (references, codings, extensions) that you will parse repeatedly across models. dbt macros eliminate this repetition:
-- macros/parse_fhir_reference.sql
-- Extract the ID from a FHIR reference string
-- Input: "Patient/abc-123" -> Output: "abc-123"
{%- macro parse_fhir_reference(column_expr) -%}
split_part({{ column_expr }}::string, '/', 2)
{%- endmacro -%}
-- macros/extract_coding.sql
-- Extract code from a FHIR CodeableConcept by system
{%- macro extract_coding(column_expr, system_url) -%}
(
select c.value:code::string
from lateral flatten(
input => {{ column_expr }}:coding
) c
where c.value:system::string = '{{ system_url }}'
limit 1
)
{%- endmacro -%}
-- macros/fhir_extension_value.sql
-- Extract value from a FHIR extension by URL
{%- macro fhir_extension_value(column_expr, ext_url, value_type) -%}
(
select e.value:value{{ value_type }}::string
from lateral flatten(
input => {{ column_expr }}:extension
) e
where e.value:url::string = '{{ ext_url }}'
limit 1
)
{%- endmacro -%}
Putting It All Together: The Complete ELT Pipeline
Here is how the full ELT pipeline works for a healthcare organization:
- Extract: Fivetran, Airbyte, or custom connectors pull raw FHIR data from EHR APIs, raw HL7v2 from integration engines, and raw claims from clearinghouses.
- Load: Raw data lands in the warehouse (Snowflake, BigQuery, or Databricks) as JSON columns. Nothing is transformed. Nothing is lost.
- Validate: Great Expectations checks raw data quality before transformation. Bad batches are quarantined and the team is alerted.
- Transform: dbt models parse raw JSON into staging tables, apply business logic in intermediate tables, and build analytics-ready marts.
- Test: dbt tests run on every execution. Schema tests, relationship tests, and custom clinical tests catch issues automatically.
- Document: dbt docs generate a searchable data catalog with lineage graphs showing how raw data flows to each mart table.
- Serve: Mart tables power Looker dashboards, ML feature stores, and regulatory reports. Clinical researchers query directly.
For the streaming complement to this batch ELT pipeline, see our guide on streaming healthcare data with Kafka and FHIR. For organizations using Delta Lake as the storage layer, see our Delta Lake for healthcare guide.
Frequently Asked Questions
Is ELT more expensive than ETL?
Usually less expensive. ETL requires a separate transformation server (Informatica, Talend) plus the warehouse. ELT uses only the warehouse. Cloud warehouses charge for compute time, and dbt transformations are typically efficient SQL queries that cost pennies per run. The total infrastructure cost is 30-50% lower with ELT for most healthcare data volumes.
How do I handle PHI in an ELT pipeline?
Load raw data into a restricted schema with strict access controls. Apply de-identification in the transformation layer using dbt models that hash, mask, or tokenize PHI fields. The restricted raw schema is accessible only to the service account running dbt. Downstream mart tables contain only de-identified or authorized data. This defense-in-depth approach satisfies HIPAA minimum necessary requirements.
Can dbt handle the volume of healthcare data?
dbt generates SQL that runs on your warehouse engine. If your warehouse can handle the volume (Snowflake, BigQuery, and Databricks all scale to petabytes), dbt can transform it. dbt itself does not process data. It compiles SQL models and executes them against the warehouse. For a 500-bed health system with 50 million observations, a full dbt run typically completes in 15-30 minutes on a medium Snowflake warehouse.
What about real-time transforms?
ELT with dbt handles batch and micro-batch transforms (every 5 minutes to every 6 hours). For true real-time transforms (sub-second), use Kafka Streams or Spark Structured Streaming in combination with ELT. Stream processing handles latency-sensitive events (critical lab alerts), while dbt handles analytical transforms on the same data in the warehouse.
How do I migrate from ETL to ELT?
Run both in parallel during migration. Continue running the existing ETL pipeline while building ELT models in dbt. Compare outputs to verify the dbt models produce identical results. Once validated, decommission the ETL pipeline. This dual-run approach takes 2-4 months for a typical healthcare data warehouse and eliminates the risk of a big-bang migration.
Does dbt support data lineage for compliance?
Yes. dbt automatically generates a DAG (directed acyclic graph) showing how every model depends on upstream models and sources. dbt docs generate creates an interactive documentation site where auditors can trace any metric back to its raw source data. Combined with Git history (every transform change is version-controlled), this provides a complete audit trail of every transformation ever applied to the data.


