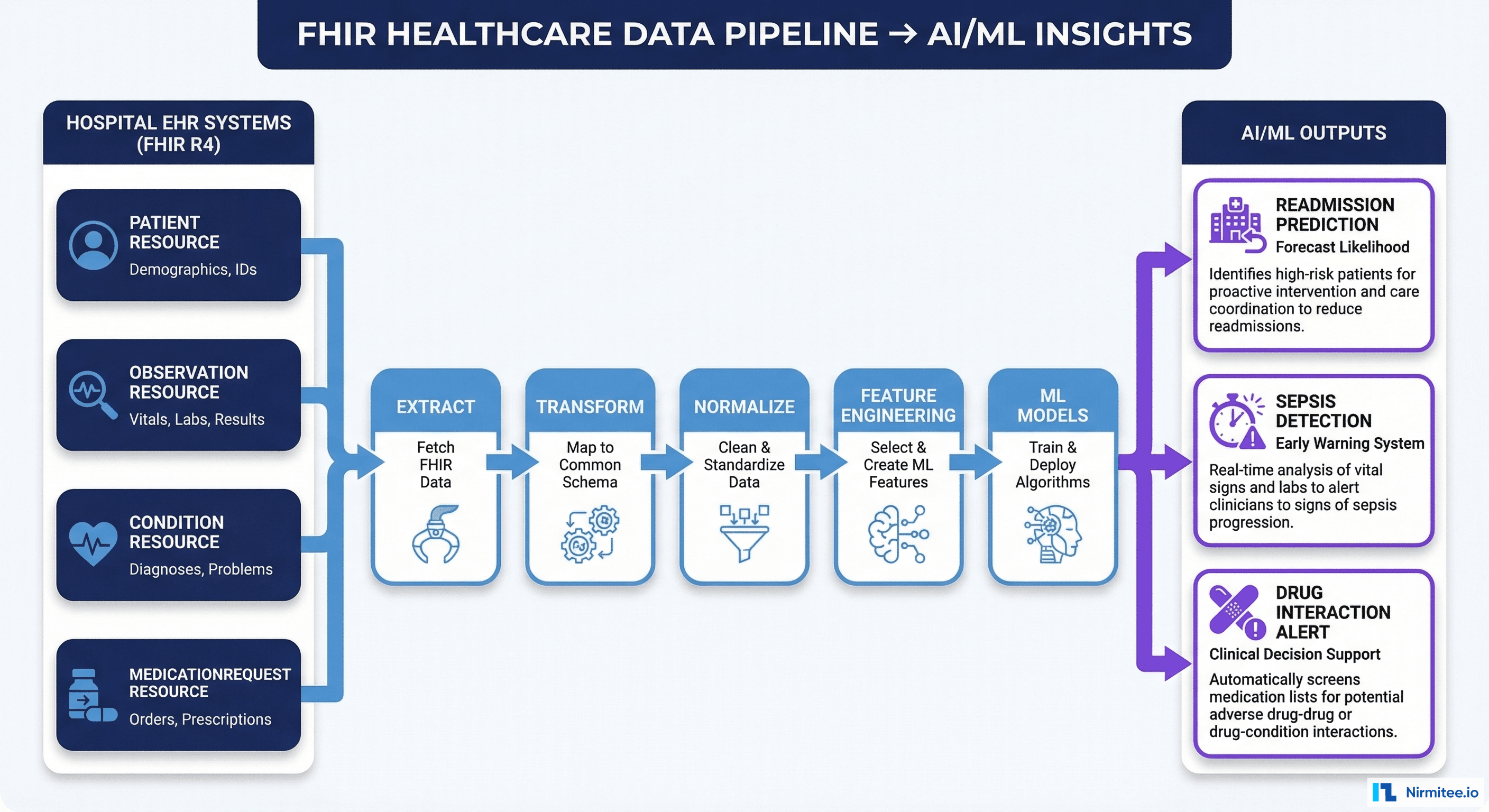
Healthcare organizations are sitting on a goldmine of clinical data — but getting that data into a format that machine learning models can actually use remains one of the hardest engineering challenges in the industry. FHIR (Fast Healthcare Interoperability Resources) has become the universal language for health data exchange, and building robust data pipelines from FHIR to ML is now a critical capability for any health system investing in AI.
This guide walks through the complete architecture for building production-grade FHIR-to-AI/ML data pipelines — from extracting FHIR resources out of EHR systems to training models that generate real clinical value. Whether you're predicting hospital readmissions, detecting sepsis early, or optimizing revenue cycle operations, the pipeline patterns are the same.

Why FHIR Is the Right Foundation for Healthcare AI
Before FHIR, healthcare data pipelines were built on HL7 v2 messages, proprietary CSV exports, and custom database extracts. Each hospital had its own format. Each vendor had its own schema. Building a model that worked across multiple sites meant months of custom mapping work for every new data source.
FHIR changes this fundamentally. Here's why it matters for AI/ML:
- Standardized resource model: Every Patient, Observation, Condition, and MedicationRequest follows the same structure regardless of the source EHR. A lab result from Epic looks structurally identical to one from Cerner or Athena.
- Code system consistency: FHIR resources reference standard terminologies — LOINC for labs, SNOMED CT for diagnoses, RxNorm for medications, ICD-10 for billing codes. This eliminates the "same concept, different code" problem that plagues legacy pipelines.
- RESTful APIs: FHIR Bulk Data Access (the Bulk Data IG) provides a standardized way to export millions of records via asynchronous NDJSON downloads. No more database-level extracts or batch file transfers.
- Temporal richness: FHIR resources carry timestamps, periods, and provenance information that lets you reconstruct the clinical timeline — critical for time-series ML models.
- ONC mandate compliance: Under the 21st Century Cures Act, all certified EHRs must support FHIR R4 APIs. This isn't optional — it's federal law. Your pipeline is building on a guaranteed foundation.
Step 1: Extracting FHIR Data from EHR Systems
The first stage of any FHIR-to-ML pipeline is getting the data out. There are three primary extraction patterns, each suited to different use cases.

Pattern A: FHIR Bulk Data Export (Batch Analytics)
For training ML models, you need historical data in bulk — not one patient at a time. The FHIR Bulk Data Access specification ($export operation) is designed exactly for this. It works as a three-step asynchronous process:
# 1. Kick off the export
curl -X GET 'https://ehr.example.com/fhir/Patient/$export' \
-H 'Accept: application/fhir+json' \
-H 'Prefer: respond-async' \
-H 'Authorization: Bearer {{access_token}}'
# Response: 202 Accepted
# Content-Location: https://ehr.example.com/fhir/bulkstatus/abc123
# 2. Poll for completion
curl -X GET 'https://ehr.example.com/fhir/bulkstatus/abc123' \
-H 'Authorization: Bearer {{access_token}}'
# Response (when ready): 200 OK with NDJSON file URLs
# 3. Download NDJSON files
curl -X GET 'https://ehr.example.com/fhir/bulkfiles/patient-001.ndjson' \
-H 'Authorization: Bearer {{access_token}}' \
-o patients.ndjsonKey considerations for bulk export:
- Scope your export: Use
_typeparameter to request only the resources you need (e.g.,_type=Patient,Observation,Condition). Exporting everything wastes bandwidth and storage. - Incremental exports: Use
_sinceparameter for subsequent runs. Don't re-export your entire dataset every time — that's a common anti-pattern that kills performance. - Backend services auth: Bulk exports use the SMART Backend Services authorization profile with client credentials + JWT assertion. This is system-to-system auth, not patient-facing OAuth.
Pattern B: FHIR Search API (Real-Time Inference)
For real-time ML inference — e.g., a sepsis risk score computed when a nurse opens a patient chart — you need fresh data on demand. The FHIR Search API lets you pull specific resources for a single patient:
# Get last 24 hours of vital signs for a patient
curl -X GET 'https://ehr.example.com/fhir/Observation?patient=12345&category=vital-signs&date=ge2026-03-20' \
-H 'Accept: application/fhir+json' \
-H 'Authorization: Bearer {{access_token}}'
# Get active medications
curl -X GET 'https://ehr.example.com/fhir/MedicationRequest?patient=12345&status=active' \
-H 'Accept: application/fhir+json' \
-H 'Authorization: Bearer {{access_token}}'Pattern C: FHIR Subscriptions (Event-Driven)
FHIR R5 introduces topic-based Subscriptions that let you stream data changes in real time. For ML pipelines that need continuous model updates or real-time feature computation, this is the most efficient pattern:
{
"resourceType": "Subscription",
"status": "requested",
"topic": "http://example.org/SubscriptionTopic/new-lab-result",
"channelType": {
"system": "http://terminology.hl7.org/CodeSystem/subscription-channel-type",
"code": "rest-hook"
},
"endpoint": "https://ml-pipeline.example.com/fhir-webhook"
}Step 2: Data Transformation — From FHIR JSON to ML-Ready Features
Raw FHIR resources are deeply nested JSON documents optimized for interoperability, not for feeding into a pandas DataFrame or a TensorFlow dataset. The transformation layer is where most of the engineering complexity lives.
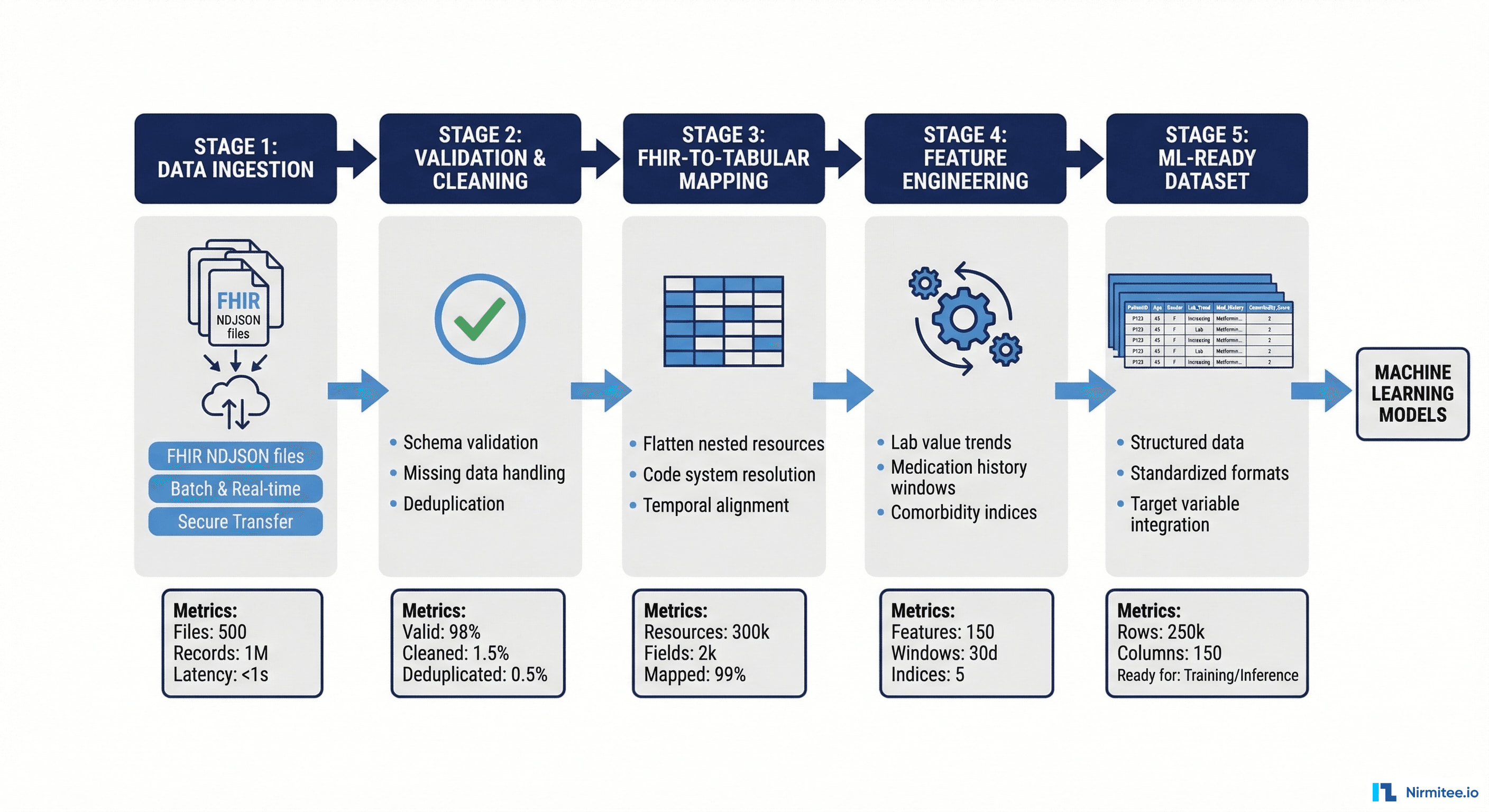
2.1 Flattening Nested FHIR Resources
A single FHIR Observation resource for a blood pressure reading contains nested component arrays, coding arrays within code, and references to other resources. You need to flatten this into tabular columns:
# Raw FHIR Observation (simplified)
{
"resourceType": "Observation",
"code": {
"coding": [
{"system": "http://loinc.org", "code": "85354-9", "display": "Blood pressure panel"}
]
},
"component": [
{
"code": {"coding": [{"system": "http://loinc.org", "code": "8480-6"}]},
"valueQuantity": {"value": 120, "unit": "mmHg"}
},
{
"code": {"coding": [{"system": "http://loinc.org", "code": "8462-4"}]},
"valueQuantity": {"value": 80, "unit": "mmHg"}
}
],
"effectiveDateTime": "2026-03-20T10:30:00Z",
"subject": {"reference": "Patient/12345"}
}
# Flattened output for ML
# patient_id | timestamp | systolic_bp | diastolic_bp
# 12345 | 2026-03-20T10:30:00Z | 120 | 80Use libraries like FHIRPath for consistent extraction across resource types, or build custom extractors with Python's fhirpathpy library. Avoid hand-written JSON traversals — they break when resources contain optional fields or extensions.
2.2 Code System Resolution
FHIR resources reference standardized code systems, but the same clinical concept can be coded differently across institutions. Your pipeline must resolve these to a canonical representation:
import pandas as pd
# Map LOINC codes to ML feature names
LOINC_FEATURE_MAP = {
"8480-6": "systolic_bp",
"8462-4": "diastolic_bp",
"8867-4": "heart_rate",
"8310-5": "body_temperature",
"2708-6": "oxygen_saturation",
"718-7": "hemoglobin",
"4548-4": "hba1c",
"2345-7": "glucose",
"2160-0": "creatinine",
"6299-2": "bun",
}
def resolve_observation(obs: dict) -> dict:
"""Extract feature name and value from a FHIR Observation."""
codings = obs.get("code", {}).get("coding", [])
for coding in codings:
if coding.get("system") == "http://loinc.org":
feature = LOINC_FEATURE_MAP.get(coding["code"])
if feature:
value = obs.get("valueQuantity", {}).get("value")
timestamp = obs.get("effectiveDateTime")
patient_ref = obs.get("subject", {}).get("reference", "").split("/")[-1]
return {
"patient_id": patient_ref,
"feature": feature,
"value": value,
"timestamp": timestamp
}
return None2.3 Temporal Alignment
Clinical data arrives at irregular intervals. A patient might have vitals recorded every 4 hours during an ICU stay, but only once during an outpatient visit. ML models need consistent time windows. Common approaches:
- Snapshot windows: Aggregate values within fixed time buckets (e.g., "last 24 hours", "last 7 days") using mean, min, max, or last-known-value.
- Carry-forward imputation: For slowly-changing values like weight or diagnosis status, carry the last known value forward until a new measurement appears.
- Event-relative alignment: For readmission prediction, align all features to "time relative to discharge" rather than absolute timestamps.
2.4 De-identification (HIPAA Safe Harbor)
Before any data reaches your ML training environment, it must be de-identified per HIPAA Safe Harbor or Expert Determination methods. This means stripping:
- All 18 HIPAA identifiers (names, dates more specific than year, MRNs, SSNs, etc.)
- Free-text clinical notes (these often contain patient names embedded in narrative)
- Geographic data below the state level
- Dates shifted by a consistent random offset per patient (preserving temporal relationships)
# Date shifting example — preserve intervals, remove real dates
import hashlib
from datetime import timedelta
def get_date_shift(patient_id: str, salt: str, max_days: int = 365) -> int:
"""Deterministic date shift per patient using HMAC."""
h = hashlib.sha256(f"{salt}:{patient_id}".encode()).hexdigest()
return int(h[:8], 16) % (max_days * 2) - max_days
def shift_date(date_str: str, patient_id: str, salt: str) -> str:
"""Shift a date deterministically for a given patient."""
from datetime import datetime
dt = datetime.fromisoformat(date_str.replace("Z", "+00:00"))
shift = get_date_shift(patient_id, salt)
shifted = dt + timedelta(days=shift)
return shifted.isoformat()Step 3: Feature Engineering for Clinical ML
With clean, tabular data extracted from FHIR resources, the next stage is engineering features that capture clinically meaningful patterns. Healthcare ML features fall into several categories:
Static Features (from Patient, Condition resources)
- Demographics: Age (calculated from
Patient.birthDate), gender, race/ethnicity (from US Core extensions) - Comorbidity indices: Charlson Comorbidity Index or Elixhauser score computed from active
Conditionresources with ICD-10 codes - Insurance type: From
Coverageresources — Medicare, Medicaid, commercial, self-pay
Temporal Features (from Observation, Encounter resources)
- Vital sign trends: Rate of change in blood pressure, heart rate, or oxygen saturation over sliding windows (6h, 12h, 24h)
- Lab value trajectories: Is creatinine rising or falling? What's the 48-hour delta?
- Visit frequency: Number of ED visits in the past 6 months (from
Encounterresources withclass=emergency)
Medication Features (from MedicationRequest resources)
- Active medication count: Polypharmacy indicator (>5 concurrent medications)
- High-risk medication flags: Anticoagulants, opioids, insulin, immunosuppressants
- Recent medication changes: New starts or dose changes in the past 7 days
def compute_readmission_features(patient_id, discharge_date, fhir_data):
"""Compute feature vector for 30-day readmission prediction."""
features = {}
# Demographics
patient = fhir_data["Patient"]
features["age"] = calculate_age(patient["birthDate"], discharge_date)
features["gender_male"] = 1 if patient.get("gender") == "male" else 0
# Comorbidity burden
conditions = fhir_data["Condition"]
features["charlson_score"] = compute_charlson_index(conditions)
features["active_condition_count"] = len(
[c for c in conditions
if c.get("clinicalStatus", {}).get("coding", [{}])[0].get("code") == "active"]
)
# Recent utilization
encounters = fhir_data["Encounter"]
six_months_ago = shift_date(discharge_date, months=-6)
features["ed_visits_6m"] = len(
[e for e in encounters
if e.get("class", {}).get("code") == "EMER"
and e.get("period", {}).get("start", "") >= six_months_ago]
)
# Medications
meds = fhir_data["MedicationRequest"]
active_meds = [m for m in meds if m.get("status") == "active"]
features["active_med_count"] = len(active_meds)
features["polypharmacy"] = 1 if len(active_meds) > 5 else 0
# Lab trends (last 48 hours)
observations = fhir_data["Observation"]
features["creatinine_last"] = get_latest_value(observations, "2160-0")
features["creatinine_trend_48h"] = compute_trend(observations, "2160-0", hours=48)
features["hemoglobin_last"] = get_latest_value(observations, "718-7")
return featuresStep 4: Healthcare AI Use Cases Powered by FHIR Pipelines
A well-built FHIR data pipeline isn't tied to a single use case — it's a platform that powers multiple AI applications. Here are the highest-impact use cases we see health systems deploying today:

Hospital Readmission Prediction
The most mature use case. Using FHIR resources (Encounter, Condition, Observation, MedicationRequest), gradient boosting models can predict 30-day readmission risk at discharge with AUC scores of 0.72–0.78. Health systems like Mount Sinai have demonstrated 23% readmission reductions by targeting high-risk patients with transitional care interventions.
Sepsis Early Warning
Real-time FHIR Observation data (vitals, labs) feeds LSTM neural networks that detect sepsis 4–6 hours before clinical recognition. Unlike Epic's widely criticized proprietary sepsis model, open FHIR-based models allow external validation and site-specific tuning — addressing the "black box" problem that has undermined trust in healthcare AI.
Clinical Trial Matching
NLP models combined with structured FHIR data (Patient demographics, Condition diagnoses, Procedure history) can automatically screen patients against trial eligibility criteria. Organizations report 3x faster enrollment and significantly reduced screening coordinator workload.
Revenue Cycle Optimization
FHIR's financial resources (Claim, ExplanationOfBenefit) enable ML models that predict claim denials before submission. Random forest classifiers trained on historical denial patterns can flag high-risk claims for pre-submission review, reducing denial rates by 12–15%.
Population Health Risk Stratification
Combining Patient demographics, chronic Conditions, social determinants (from SDOH screening Observations), and utilization patterns enables Cox proportional hazards models that stratify populations into risk tiers — driving proactive care management and value-based contract performance.
Step 5: HIPAA-Compliant ML Infrastructure
Healthcare ML pipelines must operate within strict regulatory boundaries. HIPAA, HITECH, and state privacy laws impose requirements that general-purpose ML platforms (like a vanilla SageMaker or Vertex AI setup) don't satisfy out of the box.
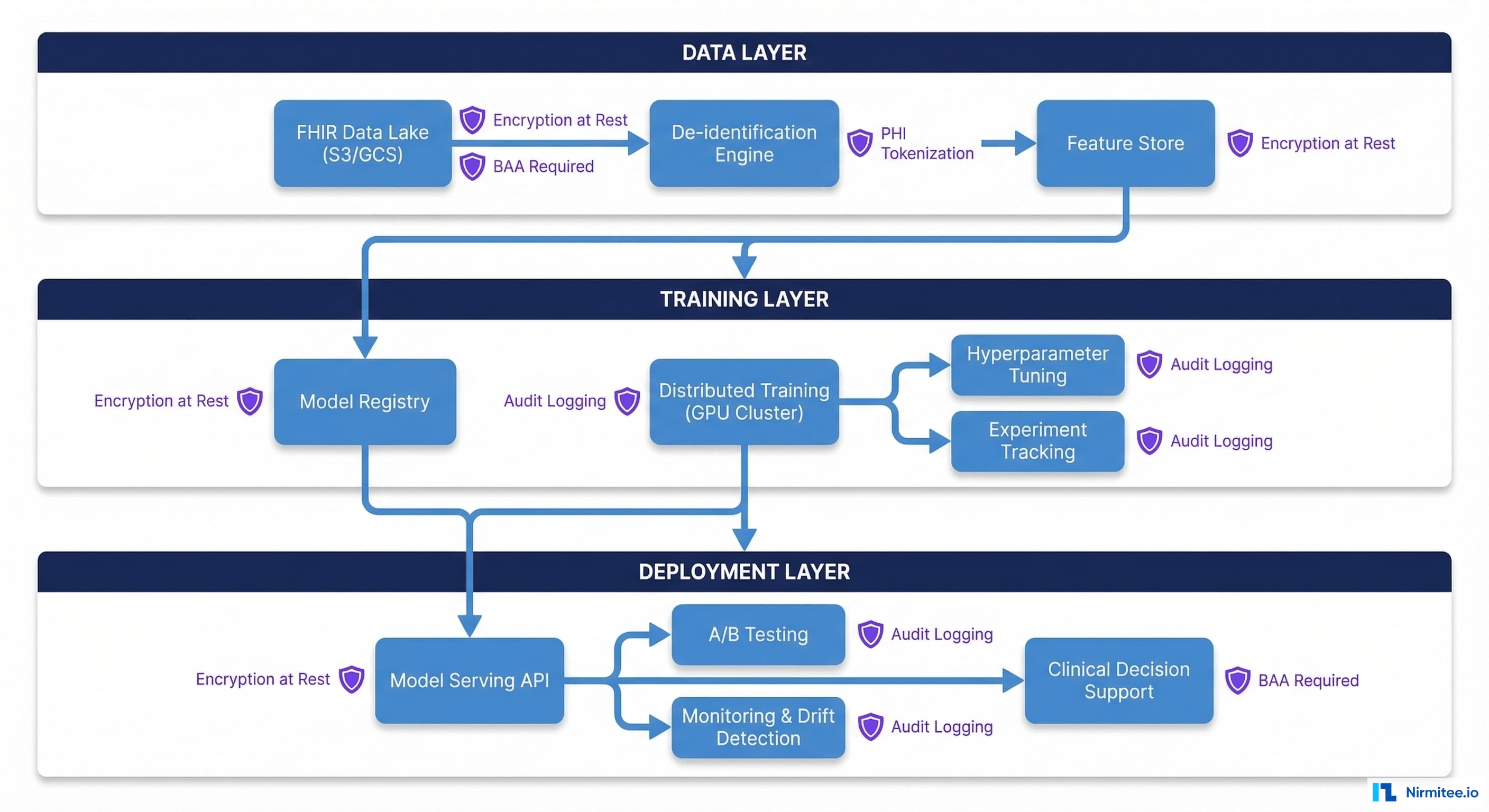
Infrastructure Requirements
- Encryption at rest and in transit: All FHIR data and derived datasets must be encrypted using AES-256 at rest and TLS 1.2+ in transit. This applies to your data lake, feature store, model artifacts, and training logs.
- PHI access controls: Implement role-based access control (RBAC) with the principle of least privilege. Data scientists should access de-identified data only. Only the de-identification pipeline itself should touch PHI.
- Audit logging: Every data access, model training run, and inference request must be logged with who, what, when, and why. Use immutable audit logs (e.g., AWS CloudTrail, GCP Audit Logs) that can't be tampered with.
- Business Associate Agreements: Every cloud service that touches PHI or derived data requires a BAA. AWS, GCP, and Azure all offer HIPAA-eligible services — but you must configure them correctly and sign the BAA.
Recommended Architecture Stack
# Example: HIPAA-compliant pipeline on AWS
# All services configured with BAA + encryption + VPC isolation
Data Lake: S3 (SSE-KMS encryption, VPC endpoint, access logging)
Compute: SageMaker (VPC mode, no internet access, encrypted volumes)
Feature Store: SageMaker Feature Store (encryption, IAM policies)
Model Registry: SageMaker Model Registry (versioned, signed artifacts)
Orchestration: Step Functions (audit trail, error handling)
Monitoring: CloudWatch + custom drift detection Lambda
De-identification: Custom service in ECS (PHI boundary — only component with PHI access)Model Governance
Healthcare AI models require governance beyond what standard MLOps provides:
- Bias testing: Test model performance across demographic subgroups (age, race, gender, insurance type). Disparate impact must be identified and mitigated before deployment. The CMS Health Equity framework provides guidance.
- Clinical validation: Models must be validated by clinical subject matter experts, not just data scientists. A model with 0.85 AUC that triggers alert fatigue is worse than a simpler model with 0.78 AUC that clinicians actually trust.
- Drift monitoring: Healthcare data distributions shift — new EHR versions change coding patterns, seasonal illness patterns change, and patient populations evolve. Monitor feature distributions and model performance weekly.
- FDA considerations: If your model provides a diagnosis or treatment recommendation, it may qualify as a Software as a Medical Device (SaMD) under FDA regulation. Consult regulatory counsel early.
Step 6: Putting It All Together — Reference Architecture
Here's how all the pieces connect in a production FHIR-to-ML pipeline:
┌─────────────────────────────────────────────────────────────────┐
│ FHIR Data Sources │
│ ┌──────────┐ ┌──────────┐ ┌──────────┐ ┌──────────────┐ │
│ │ Epic │ │ Cerner │ │ Athena │ │ Other EHRs │ │
│ └────┬─────┘ └────┬─────┘ └────┬─────┘ └──────┬───────┘ │
│ │ │ │ │ │
│ └──────────────┴──────────────┴───────────────┘ │
│ FHIR R4 APIs │
└──────────────────────────────┬──────────────────────────────────┘
│
┌──────────▼──────────┐
│ Ingestion Layer │
│ • Bulk $export │
│ • Search API │
│ • Subscriptions │
└──────────┬──────────┘
│
┌──────────▼──────────┐
│ De-identification │
│ • Safe Harbor │
│ • Date shifting │
│ • Note scrubbing │
└──────────┬──────────┘
│
┌──────────▼──────────┐
│ Transformation │
│ • FHIR flattening │
│ • Code resolution │
│ • Temporal align │
└──────────┬──────────┘
│
┌──────────▼──────────┐
│ Feature Store │
│ • Static features │
│ • Temporal features │
│ • Online/Offline │
└──────────┬──────────┘
│
┌────────────────┼────────────────┐
│ │ │
┌─────────▼─────────┐ ┌───▼────────┐ ┌────▼──────────┐
│ Batch Training │ │ Real-Time │ │ Monitoring │
│ • Model training │ │ Inference │ │ • Drift det │
│ • Validation │ │ • Scoring │ │ • Bias audit │
│ • Registry │ │ • CDS │ │ • Performance│
└───────────────────┘ └────────────┘ └───────────────┘Common Pitfalls and How to Avoid Them
After building FHIR-to-ML pipelines across multiple health systems, these are the mistakes we see repeatedly:
- Ignoring FHIR profiles: US Core profiles add constraints and extensions that vanilla FHIR R4 doesn't require. If you're working with US health data, validate against US Core profiles, not base FHIR.
- Treating FHIR references as foreign keys: FHIR references can be relative (
Patient/123), absolute (https://ehr.example.com/fhir/Patient/123), or logical (identifier-based). Your resolution logic must handle all three. - Overfitting to single-site data: A model trained on one hospital's FHIR data will embed that hospital's coding patterns, patient population, and clinical workflows. Always validate on held-out data from a different site before claiming generalizability.
- Ignoring data freshness: FHIR Bulk Export runs can take hours for large datasets. If your model needs data fresher than your export cycle, you need a hybrid approach with real-time FHIR Search for recent data, supplementing bulk historical data.
- Skipping clinical stakeholder alignment: The most technically elegant pipeline is useless if clinicians don't trust or use the model outputs. Involve clinical champions from day one — they'll catch data quality issues that data engineers miss.
Getting Started: A Practical Roadmap
If you're building your first FHIR-to-ML pipeline, here's a phased approach:
Phase 1 (Weeks 1–4): Data Access
- Register your application with your EHR's FHIR API
- Implement SMART Backend Services authentication
- Run your first Bulk Data Export with a limited resource set (Patient, Condition, Observation)
- Validate data completeness against known patient counts
Phase 2 (Weeks 5–8): Pipeline Foundation
- Build FHIR-to-tabular transformation for your target resources
- Implement de-identification pipeline (use Safe Harbor for speed)
- Create a feature store with versioned feature definitions
- Set up automated data quality checks
Phase 3 (Weeks 9–12): First Model
- Pick a well-understood use case (readmission prediction is the easiest starting point)
- Train a baseline model (logistic regression first, then gradient boosting)
- Validate with clinical stakeholders — does the model's risk ranking match clinical intuition?
- Set up monitoring for feature drift and model performance
Phase 4 (Ongoing): Production & Scale
- Deploy model behind a HIPAA-compliant inference API
- Integrate with EHR workflows via CDS Hooks or SMART on FHIR apps
- Expand to additional use cases using the same pipeline infrastructure
- Establish a model governance board with clinical and technical representation
Conclusion
Building a FHIR-to-AI/ML data pipeline is a significant engineering investment — but it's an investment that pays compounding returns. Once you have a robust pipeline that extracts FHIR data, transforms it into ML-ready features, and operates within HIPAA boundaries, every subsequent AI use case becomes dramatically easier to deploy.
The healthcare organizations that build this infrastructure now will have a decisive advantage as AI moves from pilot projects to core clinical operations. The data is standardized. The APIs are mandated. The tooling is mature. The only question is whether you'll build the pipeline or let your competitors do it first.
At Nirmitee, we help health systems and health-tech companies build production-grade FHIR integration and AI/ML pipelines. If you're planning a FHIR-to-ML initiative and want to accelerate your timeline, let's talk.
Need expert help with healthcare data integration? Explore our Healthcare Interoperability Solutions to see how we connect systems seamlessly. We also offer specialized Healthcare Software Product Development services. Talk to our team to get started.