"Which LLM should we use for our healthcare product?"
Every clinical AI team asks this question. Few get a good answer. The landscape changes quarterly — new models launch, benchmarks shift, licensing terms evolve, and yesterday's leader becomes today's baseline. In January 2026, Google released MedGemma 1.5 and reset expectations for open-weight medical AI. Three months later, teams are still struggling to choose between it, Meditron, fine-tuned Llama variants, and commercial APIs from OpenAI and Anthropic.
The confusion is understandable. There are now over 50 LLMs with claimed medical capabilities, but fewer than a dozen have been rigorously validated on clinical benchmarks. Marketing claims blur into research results. "Medical-grade AI" appears in press releases for models that have never seen a clinical note.
This guide cuts through the noise. We compare every major healthcare LLM on the metrics that actually matter — benchmark accuracy, licensing, deployment flexibility, HIPAA compliance, fine-tunability, and total cost of ownership. We provide the decision framework, the deployment architectures, and the code to get started. Whether you are building an ambient documentation tool, a clinical decision support system, or a patient-facing triage chatbot, this is the reference you need.
The Healthcare LLM Landscape Map
The healthcare LLM ecosystem divides into four distinct categories, each with different trade-offs for clinical AI teams. Understanding where each model sits determines your architecture, your regulatory posture, and your cost structure.
Category 1: Closed Commercial Models
Models: GPT-4 / GPT-4o (OpenAI), Claude 3.5 Sonnet / Opus (Anthropic), Gemini 1.5 Pro (Google)
These are the most capable general-purpose models available. They achieve strong performance on medical benchmarks without medical-specific training simply because of their scale and training data breadth. GPT-4 scored approximately 86% on MedQA in early evaluations. Claude demonstrates particularly strong clinical reasoning on multi-step diagnostic problems. Gemini 1.5 Pro offers a 1-million-token context window, making it uniquely suited for processing entire patient records.
The trade-off is control. You cannot fine-tune the underlying weights (only prompt-tune or use retrieval augmentation). Your data passes through third-party infrastructure. HIPAA compliance requires Business Associate Agreements (BAAs) — available through Azure OpenAI, AWS Bedrock for Claude, and Google Cloud Vertex AI for Gemini, but not through direct API access.
A notable cautionary signal: in early 2026, FDA's CDER division reportedly switched its internal Elsa AI assistant from Claude to Gemini after encountering hallucination issues in regulatory document analysis. The episode underscores that no commercial model is infallible in clinical contexts, and provider organizations must implement their own real-time AI guardrail systems regardless of which commercial model they choose.
Category 2: Open General-Purpose Models
Models: Llama 3 70B / 405B (Meta), Mixtral 8x22B (Mistral), Gemma 3 (Google), Qwen 2.5 (Alibaba)
Open-weight general models serve as the foundation layer for most custom healthcare AI. Llama 3 has become the de facto base for fine-tuned clinical models — its permissive license, strong reasoning capabilities, and availability in multiple sizes (8B, 70B, 405B) make it the starting point for teams building specialized systems. Gemma 3, Google's open-weight family, provides the base architecture for MedGemma.
These models are not medical models out of the box. Their value is as fine-tuning substrates: you take Llama 3 70B, fine-tune it on your clinical corpus, and get a model that combines Llama's general intelligence with domain-specific knowledge. We cover the fine-tuning process in detail later in this article.
Category 3: Open Medical-Specialized Models
Models: MedGemma 1.5 (Google), Meditron 70B (EPFL/Yale), BioMistral 7B, OpenBioLLM 70B, PMC-LLaMA
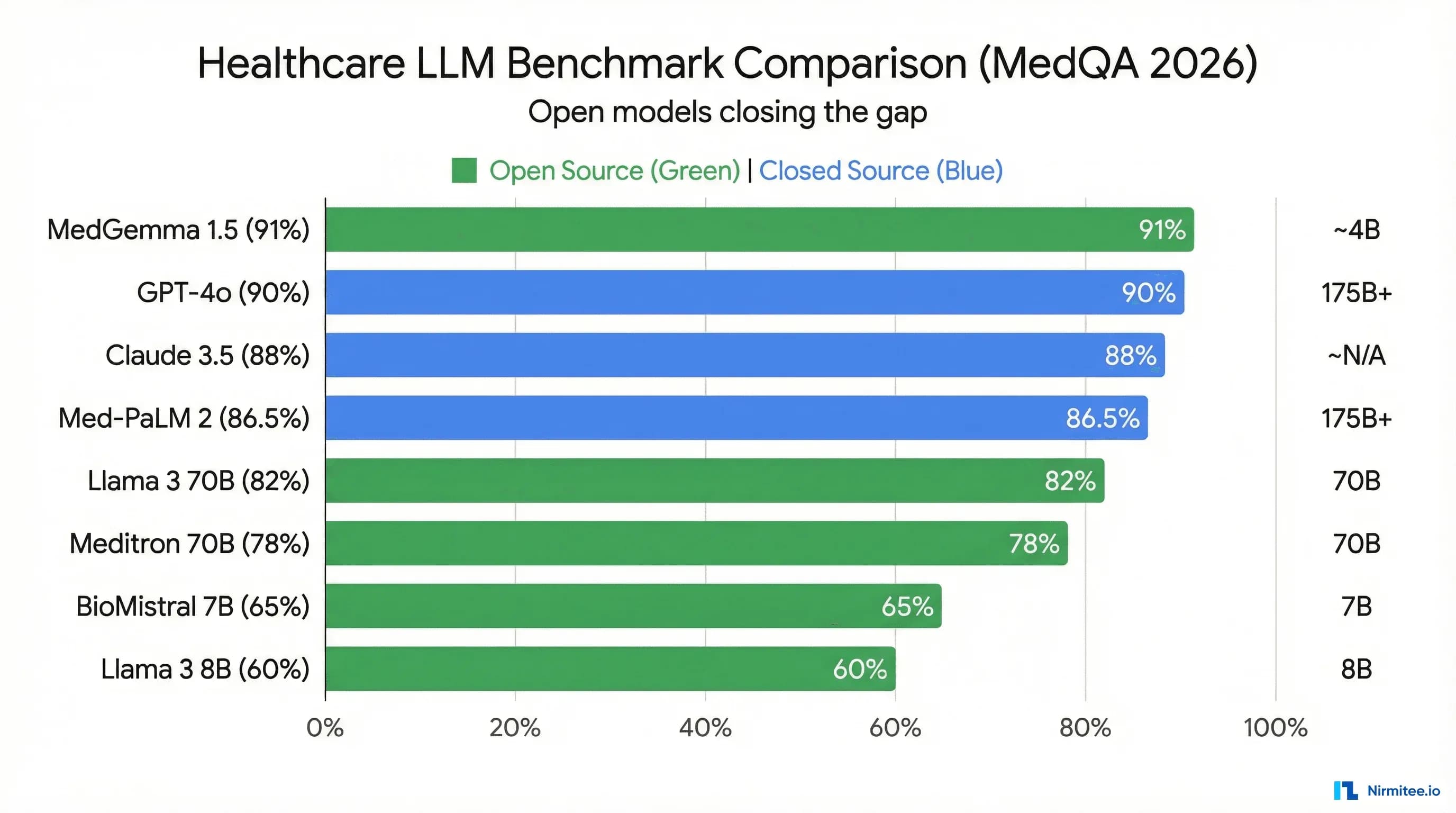
This is where the field is moving fastest. These models are pre-trained or fine-tuned specifically on medical literature, clinical guidelines, and biomedical corpora. MedGemma 1.5, released January 2026, represents the current state of the art among open-weight medical models, achieving approximately 91% on MedQA — surpassing Med-PaLM 2's 86.5% score from 2023.
The diversity here matters. Meditron was trained by academic researchers at EPFL and Yale on curated PubMed and clinical guideline datasets. BioMistral focuses on biomedical text. PMC-LLaMA was trained on PubMed Central articles. Each has different strengths depending on your use case.
Category 4: Specialized Clinical Systems
Examples: Nuance DAX Copilot (ambient documentation), Epic Cognitive Computing (EHR-integrated), Google Health AI (imaging)
These are not standalone LLMs — they are integrated products that use LLMs as components within larger clinical systems. They are mentioned for completeness but are not directly comparable to the models above. If you are building your own clinical AI product, you are choosing from Categories 1-3.
Head-to-Head Benchmark Comparison
Benchmarks are imperfect proxies for clinical utility. MedQA tests multiple-choice medical knowledge, not bedside reasoning. PubMedQA measures biomedical literature comprehension, not clinical note quality. But benchmarks are the only standardized comparison we have, and they correlate meaningfully with downstream task performance when combined with real-world evaluation. For a comprehensive guide on evaluating LLM outputs beyond benchmarks, see our LLM testing at scale guide.
| Model | Parameters | MedQA Score | PubMedQA | License | Multimodal | Fine-Tunable | HIPAA Deployment | Approx. Cost |
|---|---|---|---|---|---|---|---|---|
| MedGemma 1.5 | 4B / 27B | ~91% | ~81% | Gemma License (open-weight) | Yes (text + imaging) | Yes (full weights) | On-prem / private cloud | Free (self-hosted) |
| Med-PaLM 2 | 340B (est.) | 86.5% | 81.8% | Closed (Google) | Yes (Med-PaLM M) | No | Google Cloud (BAA) | Enterprise pricing |
| GPT-4o | Undisclosed | ~86% | ~78% | Closed (OpenAI) | Yes | No (fine-tune API limited) | Azure OpenAI (BAA) | $2.50-$10 / 1M tokens |
| Claude 3.5 Sonnet | Undisclosed | ~84% | ~77% | Closed (Anthropic) | Yes (vision) | No | AWS Bedrock (BAA) | $3-$15 / 1M tokens |
| Meditron 70B | 7B / 70B | ~70% | ~74% | Llama 2 License | No | Yes (full weights) | On-prem / private cloud | Free (self-hosted) |
| Llama 3 70B | 8B / 70B / 405B | ~72% | ~73% | Llama 3 Community License | No (Llama 3.2 adds vision) | Yes (full weights) | On-prem / private cloud | Free (self-hosted) |
| OpenBioLLM 70B | 8B / 70B | ~73% | ~78% | Apache 2.0 | No | Yes (full weights) | On-prem / private cloud | Free (self-hosted) |
| BioMistral 7B | 7B | ~60% | ~71% | Apache 2.0 | No | Yes (full weights) | On-prem / private cloud | Free (self-hosted) |
| PMC-LLaMA | 7B / 13B | ~62% | ~72% | Llama 2 License | No | Yes (full weights) | On-prem / private cloud | Free (self-hosted) |
| Gemini 1.5 Pro | Undisclosed | ~85% | ~79% | Closed (Google) | Yes | No | Vertex AI (BAA) | $1.25-$5 / 1M tokens |
Key takeaways from the table:
- MedGemma 1.5 is the new benchmark leader among open models. At ~91% MedQA, it surpasses both Med-PaLM 2 (86.5%) and GPT-4 (~86%) while being fully open-weight and self-hostable. This is a paradigm shift.
- Open models have closed the gap. Two years ago, open medical models trailed proprietary ones by 20+ percentage points. That gap is now single digits — and MedGemma has eliminated it entirely on MedQA.
- Parameter count is not destiny. MedGemma's 27B variant outperforms models 10x its size because of targeted medical pre-training. Efficient architecture plus domain data beats raw scale.
- Multimodal capability is the next differentiator. MedGemma, GPT-4o, and Gemini support vision. For radiology, pathology, and dermatology use cases, this is a hard requirement.
- HIPAA deployment separates open from closed. Every open model can be deployed on-premises behind your firewall. Commercial models require BAAs with cloud providers, adding cost, complexity, and vendor lock-in.
MedGemma Deep-Dive
MedGemma 1.5 deserves extended analysis because it represents a category shift: the first open-weight model to definitively surpass proprietary medical AI on standard benchmarks. Built on Google's Gemma 3 architecture, it was released in January 2026 and has rapidly become the default starting point for new clinical AI projects.
Architecture and Training
MedGemma is built on Gemma 3, Google's open-weight model family. It comes in two sizes: a 4B parameter variant optimized for edge deployment and resource-constrained environments, and a 27B variant that delivers the headline benchmark results. Both share the same architecture — dense transformer with grouped-query attention, RoPE positional embeddings, and SentencePiece tokenization.
What makes MedGemma different from simply fine-tuning Gemma 3 on medical data is the pre-training stage. Google's Health AI team continued pre-training on a curated corpus of:
- PubMed abstracts and full-text articles — covering biomedical research literature
- Medical textbooks and reference materials — structured medical knowledge
- Clinical practice guidelines — evidence-based treatment protocols from specialty societies
- De-identified clinical notes — real-world clinical language patterns
- Medical imaging datasets — paired with radiologist reports for multimodal understanding
This continued pre-training embeds medical knowledge into the model's weights at a deeper level than instruction fine-tuning alone can achieve. The model learns medical ontologies, drug interactions, diagnostic reasoning patterns, and clinical terminology as core knowledge rather than surface-level pattern matching.
Multimodal Capabilities
MedGemma's multimodal support is its most significant differentiator against text-only medical models like Meditron. The model processes medical images — chest X-rays, CT slices, pathology slides, dermatological photographs — alongside text prompts, enabling use cases that were previously impossible with open-weight models:
- Radiology report generation — Input a chest X-ray, receive a structured findings report
- Pathology slide interpretation — Analyze histopathology images with clinical context
- Dermatology triage — Classify skin lesion images with differential diagnoses
- Clinical document understanding — Process scanned forms, handwritten notes, and lab result images
For teams building imaging-adjacent applications, MedGemma eliminates the need for separate vision and language models. A single model handles both the image understanding and the clinical reasoning. For teams working with medical imaging pipelines, our guide on FHIR-based AI/ML data pipelines covers how to feed structured imaging data into models like MedGemma.
How to Use MedGemma
MedGemma is available through Hugging Face and can be deployed using standard inference frameworks. Here is a minimal setup using the Hugging Face Transformers library:
from transformers import AutoTokenizer, AutoModelForCausalLM
import torch
model_name = "google/medgemma-27b-text-v1.5"
tokenizer = AutoTokenizer.from_pretrained(model_name)
model = AutoModelForCausalLM.from_pretrained(
model_name,
torch_dtype=torch.bfloat16,
device_map="auto"
)
prompt = """You are a clinical assistant. Based on the following patient
presentation, provide a differential diagnosis ranked by likelihood.
Patient: 58-year-old male presenting with acute onset substernal chest
pain radiating to the left arm, diaphoresis, and nausea. History of
hypertension and type 2 diabetes. Current medications: metformin 1000mg
BID, lisinopril 20mg daily.
Differential diagnosis:"""
inputs = tokenizer(prompt, return_tensors="pt").to(model.device)
outputs = model.generate(**inputs, max_new_tokens=512, temperature=0.3)
print(tokenizer.decode(outputs[0], skip_special_tokens=True))For production deployment with higher throughput, use vLLM or Text Generation Inference (TGI) — covered in the deployment section below.
When to Choose MedGemma
MedGemma is the right choice when you need open-weight medical AI with multimodal capabilities and on-premise deployment. Specifically:
- You need to process medical images alongside clinical text
- You require full control over model weights for fine-tuning on your specific clinical domain
- Your deployment must be on-premises or in a private cloud for HIPAA compliance
- You want state-of-the-art medical benchmark performance without commercial API costs
- Your organization's GPU budget can support a 27B parameter model (requires ~60GB VRAM for BF16, ~15GB for 4-bit quantized). For strategies on reducing these resource requirements, see our guide on model compression for healthcare: quantization, distillation, and pruning
MedGemma is not the right choice if you need a model smaller than 4B parameters for edge devices, if you require a model with a proven regulatory track record (it is too new), or if your use case is purely text-based and does not benefit from multimodal capability — in which case, smaller specialized models may be more cost-effective.
Open-Source Medical Models Worth Knowing
MedGemma leads the benchmarks, but it is not the only open medical model worth evaluating. Several alternatives offer unique advantages for specific use cases.
Meditron (EPFL / Yale)
Sizes: 7B and 70B parameters
Base: Llama 2
Training data: 48.1B tokens from PubMed, medical guidelines (WHO, FDA, EMA, NICE), and curated clinical corpora
MedQA: ~70% (70B variant)
Meditron is the most academically rigorous open medical model. Developed by researchers at EPFL (Switzerland) and Yale, it emphasizes evidence-based medical knowledge drawn from authoritative clinical guidelines rather than web-scraped medical content. The training pipeline is fully documented in their published paper, making it the most transparent model in terms of data provenance.
Meditron's strength is in clinical guideline adherence — tasks where you need the model to reason about treatment protocols, contraindications, and evidence levels. Its weakness is that it is built on the older Llama 2 architecture, which limits its context window (4K tokens) and makes it less suitable for processing long clinical documents. There is no multimodal support.
Best for: Teams that prioritize transparency, reproducibility, and guideline-grounded reasoning over raw benchmark scores.
BioMistral 7B
Size: 7B parameters
Base: Mistral 7B
Training data: PubMed Central articles
License: Apache 2.0
MedQA: ~60%
BioMistral is the lightest viable medical model. At 7B parameters, it runs on a single consumer GPU (16GB VRAM at 4-bit quantization), making it accessible for research labs and startups without enterprise GPU infrastructure. Built on Mistral's efficient architecture, it offers faster inference per token than larger alternatives.
The trade-off is clear: benchmark performance lags significantly behind larger models. BioMistral is not suitable as a standalone clinical AI. It works best as a first-pass filter — triaging documents, extracting medical entities, classifying clinical text — where speed and cost matter more than reasoning depth.
Best for: NLP pipeline components, medical entity extraction, document classification, and teams with limited GPU budgets.
OpenBioLLM 70B
Sizes: 8B and 70B parameters
Base: Llama 3
Training data: Curated biomedical and clinical datasets
License: Apache 2.0
MedQA: ~73%
OpenBioLLM is built on the more modern Llama 3 architecture, giving it access to an 8K token context window and improved reasoning capabilities compared to Llama 2-based models like Meditron. Its Apache 2.0 license is the most permissive of any model in this category — no usage restrictions, no commercial limitations, no attribution requirements beyond the license.
Best for: Teams that need a permissively licensed medical model on a modern architecture for commercial products.
PMC-LLaMA
Sizes: 7B and 13B parameters
Base: Llama 2
Training data: 4.8M PubMed Central biomedical papers
License: Llama 2 License
MedQA: ~62%
PMC-LLaMA is notable for its training data focus: exclusively PubMed Central full-text papers. This makes it particularly strong for biomedical research tasks — literature review, study summarization, methodology extraction — rather than clinical practice tasks. If your application involves processing research literature rather than clinical notes, PMC-LLaMA may outperform models trained on broader but shallower medical corpora.
Best for: Biomedical research tools, literature review systems, and evidence synthesis applications.
RAG vs Fine-Tuning: The Decision Tree
Every clinical AI team faces the same architectural fork: do you keep the base model unchanged and augment it with retrieval (RAG), or do you modify the model's weights to embed domain knowledge (fine-tuning)? The answer depends on your data, your use case, and your operational maturity.
Retrieval-Augmented Generation (RAG)
How it works: You keep the base model frozen. At inference time, you retrieve relevant documents from a vector database (clinical guidelines, formularies, patient records) and inject them into the prompt as context. The model reasons over both the prompt and the retrieved documents.
When to use RAG:
- Your knowledge base changes frequently (drug formularies, clinical guidelines, insurance rules)
- You need the model to cite sources and provide verifiable references
- You do not have labeled training data for fine-tuning
- You need to deploy quickly — RAG can be production-ready in weeks, not months
- Your organization lacks ML engineering capacity for training pipelines
RAG limitations: Retrieval quality caps output quality. If the vector search returns irrelevant documents, the model generates incorrect responses with high confidence. Context window limits constrain how much retrieved content the model can process. And RAG cannot change the model's fundamental reasoning patterns — it provides information but does not teach the model to think differently.
Fine-Tuning
How it works: You modify the model's weights by training on your domain-specific dataset. The model internalizes new knowledge and behavioral patterns that persist across all future inferences.
When to use fine-tuning:
- You need the model to adopt a specific clinical reasoning style (radiological report format, SOAP note structure)
- You have a large, high-quality labeled dataset (10K+ examples minimum)
- Your use case involves a narrow, well-defined domain (ophthalmology, dermatology, cardiology)
- You need consistent output formatting without complex prompt engineering
- Latency requirements preclude the retrieval step of RAG
Fine-tuning limitations: Requires ML engineering expertise, GPU infrastructure, and ongoing retraining as your data evolves. Risk of catastrophic forgetting — the model loses general capabilities while gaining domain-specific ones. Quality is entirely dependent on your training data; biased data produces biased models.
The Practical Answer: Start with RAG
For 80% of healthcare AI teams, RAG is the right starting point. It is faster to deploy, cheaper to operate, easier to debug, and simpler to maintain. Your clinical knowledge base is already structured (FHIR resources, clinical guidelines, formulary databases) and changes frequently enough that baking it into model weights creates staleness problems.
Fine-tune only when RAG hits a clear wall: the model cannot produce the output format you need, retrieval quality is insufficient despite optimization, or your domain is specialized enough that general medical knowledge does not transfer. Even then, consider the hybrid approach — fine-tune for behavior and format, use RAG for factual knowledge.
Fine-Tuning a Clinical LLM: Step-by-Step
When RAG is insufficient and fine-tuning is justified, here is the practical pipeline. We use LoRA (Low-Rank Adaptation) because it is the most cost-effective method for clinical teams — you modify less than 1% of the model's parameters while achieving 90-95% of the performance of full fine-tuning.
Step 1: Data Preparation
Clinical fine-tuning data comes from three primary sources:
- MIMIC-IV — The gold standard for de-identified clinical notes. Requires PhysioNet credentialing. Contains discharge summaries, radiology reports, nursing notes, and clinical assessments from Beth Israel Deaconess Medical Center.
- PubMed / PMC — Freely available biomedical literature. Best for embedding research knowledge. Less useful for clinical language patterns.
- Internal clinical notes — Your organization's own data. Requires de-identification per HIPAA Safe Harbor or Expert Determination methods. This is the highest-value training data because it reflects your specific patient population, documentation style, and clinical workflows. Our guide on healthcare data de-identification for AI covers the full pipeline.
Format your training data as instruction-response pairs:
{
"instruction": "Summarize the following discharge note into a concise
clinical summary suitable for a referring physician.",
"input": "DISCHARGE SUMMARY\nPatient: 72F\nAdmission Date: 2026-03-15\n
Discharge Date: 2026-03-19\nPrimary Diagnosis: Community-acquired
pneumonia (CAP), right lower lobe\nHospital Course: Patient presented
with 3-day history of productive cough, fever 101.8F, and dyspnea on
exertion. CXR confirmed RLL consolidation. Started on ceftriaxone
1g IV q24h and azithromycin 500mg IV daily per ATS/IDSA guidelines.
Blood cultures negative x2. Transitioned to oral amoxicillin-clavulanate
875mg BID on hospital day 3. O2 requirement resolved by day 3.
Discharged on 5-day oral course...",
"output": "72-year-old female admitted 3/15-3/19 for right lower lobe
community-acquired pneumonia. Treated per ATS/IDSA protocol with
IV ceftriaxone/azithromycin, transitioned to oral amoxicillin-clavulanate.
Blood cultures negative. O2 requirement resolved day 3. Discharged
stable on 5-day oral antibiotic course. Follow-up CXR recommended
in 6 weeks."
}Step 2: LoRA Fine-Tuning with QLoRA
QLoRA combines LoRA with 4-bit quantization, reducing GPU memory requirements by approximately 75% compared to full fine-tuning. Here is a production-ready training script:
import torch
from datasets import load_dataset
from transformers import (
AutoTokenizer,
AutoModelForCausalLM,
BitsAndBytesConfig,
TrainingArguments,
)
from peft import LoraConfig, get_peft_model, prepare_model_for_kbit_training
from trl import SFTTrainer
# Model and quantization configuration
model_name = "google/medgemma-27b-text-v1.5"
bnb_config = BitsAndBytesConfig(
load_in_4bit=True,
bnb_4bit_quant_type="nf4",
bnb_4bit_compute_dtype=torch.bfloat16,
bnb_4bit_use_double_quant=True,
)
model = AutoModelForCausalLM.from_pretrained(
model_name,
quantization_config=bnb_config,
device_map="auto",
trust_remote_code=True,
)
model = prepare_model_for_kbit_training(model)
tokenizer = AutoTokenizer.from_pretrained(model_name)
tokenizer.pad_token = tokenizer.eos_token
# LoRA configuration — target attention and MLP layers
lora_config = LoraConfig(
r=16, # Rank: 16 is a strong default for medical
lora_alpha=32, # Alpha: 2x rank is standard
target_modules=[
"q_proj", "k_proj", "v_proj", "o_proj", # Attention
"gate_proj", "up_proj", "down_proj", # MLP
],
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
)
model = get_peft_model(model, lora_config)
print(f"Trainable parameters: {model.print_trainable_parameters()}")
# Load your clinical dataset
dataset = load_dataset("json", data_files="clinical_training_data.jsonl")
# Training configuration
training_args = TrainingArguments(
output_dir="./medgemma-clinical-lora",
num_train_epochs=3,
per_device_train_batch_size=4,
gradient_accumulation_steps=4,
learning_rate=2e-4,
weight_decay=0.01,
warmup_ratio=0.03,
lr_scheduler_type="cosine",
logging_steps=10,
save_strategy="epoch",
bf16=True,
gradient_checkpointing=True,
max_grad_norm=0.3,
)
# Format training examples
def format_instruction(example):
return f"""### Instruction:
{example['instruction']}
### Input:
{example['input']}
### Response:
{example['output']}"""
trainer = SFTTrainer(
model=model,
train_dataset=dataset["train"],
tokenizer=tokenizer,
args=training_args,
formatting_func=format_instruction,
max_seq_length=2048,
)
trainer.train()
model.save_pretrained("./medgemma-clinical-lora/final")Hardware requirements for this configuration: A single NVIDIA A100 80GB or two A10G 24GB GPUs. On an A100, training on 10K clinical examples takes approximately 4-6 hours. On cloud infrastructure (AWS p4d.24xlarge), budget approximately $150-200 per training run.
Step 3: Evaluation
Never ship a fine-tuned clinical model based on training loss alone. Your evaluation pipeline must include:
- Benchmark regression testing — Run MedQA and PubMedQA to ensure fine-tuning has not degraded general medical knowledge (catastrophic forgetting)
- Task-specific metrics — ROUGE and BERTScore for summarization tasks, exact match and F1 for extraction tasks, accuracy for classification tasks
- Hallucination rate measurement — Compare model outputs against verified reference data to quantify fabrication frequency
- Clinician evaluation — Board-certified clinicians score a sample of outputs on accuracy, completeness, and safety. No substitute for this step
For a production-grade evaluation framework that scales beyond manual review, see our detailed guide on 7 LLM testing architectures for production healthcare.
Deployment Architecture for Healthcare
Choosing the model is half the decision. The other half is how you deploy it in a way that satisfies HIPAA, scales under production load, and does not bankrupt your GPU budget.
On-Premises Deployment
On-premises deployment gives you complete control over data residency — PHI never leaves your network. For open-weight models like MedGemma and Meditron, this is the default deployment pattern.
Recommended serving frameworks:
- vLLM — The current standard for high-throughput LLM serving. Implements PagedAttention for efficient GPU memory management, continuous batching for maximizing throughput, and supports tensor parallelism across multiple GPUs. Start here for most deployments.
- Text Generation Inference (TGI) — Hugging Face's production serving framework. Strong integration with the Hugging Face ecosystem, built-in quantization support, and production-ready health checks and monitoring. Slightly lower raw throughput than vLLM but easier to operate.
- Ollama — For development and small-scale deployments. Single-binary installation, automatic quantization, and a simple REST API. Not suitable for high-throughput production use but excellent for prototyping and internal tools.
A minimal vLLM deployment of MedGemma:
# Install vLLM
pip install vllm
# Serve MedGemma with 4-bit quantization
python -m vllm.entrypoints.openai.api_server \
--model google/medgemma-27b-text-v1.5 \
--quantization awq \
--max-model-len 8192 \
--tensor-parallel-size 2 \
--port 8000 \
--api-key your-internal-keyThis exposes an OpenAI-compatible API, meaning your application code does not change whether you are hitting vLLM, Azure OpenAI, or any other provider. Future model swaps are a configuration change, not a code rewrite.
Cloud Deployment (HIPAA-Compliant)
For commercial models or when GPU procurement is impractical, cloud deployment with BAAs provides HIPAA compliance:
| Cloud Provider | Models Available | BAA | Key Feature |
|---|---|---|---|
| Azure OpenAI | GPT-4, GPT-4o, GPT-4 Turbo | Yes | Data never used for training; private endpoints |
| Google Vertex AI | Gemini 1.5 Pro, Med-PaLM 2, MedGemma | Yes | Managed MedGemma hosting; integrated with Google Health AI |
| AWS Bedrock | Claude 3.5, Llama 3, Mistral | Yes | Broadest model selection; PrivateLink for VPC isolation |
| AWS SageMaker | Any open model | Yes | Custom model hosting; bring your fine-tuned MedGemma |
Critical HIPAA consideration: A BAA alone is necessary but not sufficient. You must also ensure data is encrypted in transit (TLS 1.2+) and at rest, access is logged and auditable, and your application layer does not cache PHI outside the BAA-covered infrastructure. The direct consumer-facing APIs from OpenAI (api.openai.com) and Anthropic (api.anthropic.com) are not HIPAA-compliant even with an enterprise agreement — you must use the Azure, AWS, or GCP managed offerings.
Hybrid Architecture
The hybrid pattern is increasingly the production architecture of choice. It works like this:
- On-premises: A local MedGemma or Llama 3 instance handles real-time clinical workflows where latency matters and PHI is involved — clinical decision support, note generation, order checking
- Cloud: Commercial APIs (GPT-4 via Azure, Claude via Bedrock) handle batch processing, complex multi-step reasoning, and tasks where the local model's quality is insufficient — prior authorization letter generation, complex coding assistance, research queries
- Router: An intelligent routing layer evaluates each request and directs it to the appropriate model based on latency requirements, PHI sensitivity, task complexity, and cost constraints
This pattern gives you the cost efficiency and data control of local models for high-volume routine tasks, with the quality ceiling of commercial models for complex tasks — all behind a unified API.
Clinical Validation: Beyond Benchmarks
MedQA scores are a starting point, not an endpoint. A model that scores 91% on multiple-choice medical questions can still hallucinate drug dosages, fabricate clinical guidelines, and produce outputs that are plausible but dangerous. Clinical validation requires a multi-layered approach that benchmarks alone cannot provide.
Hallucination Detection
Medical hallucinations are the highest-risk failure mode. Unlike general-purpose applications where a hallucinated fact is embarrassing, a hallucinated drug interaction or fabricated dosing guideline is dangerous. Your validation pipeline must include:
- Fact verification against knowledge bases — Cross-reference model outputs against DailyMed (drug information), UpToDate (clinical guidelines), and ICD/SNOMED ontologies. Automated pipelines can flag outputs that reference non-existent drugs, incorrect dosages, or contraindicated combinations.
- Citation verification — If your model generates references (paper titles, guideline citations), verify they exist. Fabricated references are a telltale sign of hallucination.
- Consistency testing — Present the same clinical scenario multiple times with slight rephrasing. Inconsistent outputs across semantically equivalent prompts indicate unreliable reasoning.
- Adversarial probing — Intentionally present edge cases, ambiguous scenarios, and trick questions to find the model's failure boundaries.
For a comprehensive framework on building real-time guardrail systems that catch hallucinations before they reach clinicians, see our real-time AI guardrails architecture guide.
Bias Testing
Clinical AI models inherit biases from their training data. If the training corpus overrepresents certain demographics, the model's diagnostic suggestions, treatment recommendations, and risk assessments will reflect those biases. Test across:
- Demographic axes — Age, sex, race/ethnicity, socioeconomic indicators. Present identical clinical scenarios with only demographic variables changed and measure output variation.
- Disease presentation patterns — Atypical presentations (heart attack symptoms in women, depression in elderly men) that are systematically underrepresented in medical literature.
- Treatment recommendations — Check for disparities in recommended procedures, medications, and follow-up intensity across demographic groups.
FDA Considerations for SaMD
If your clinical AI product qualifies as Software as a Medical Device (SaMD), the FDA's evolving framework for AI/ML-based devices applies. The key regulatory concept is the Predetermined Change Control Plan (PCCP) — a documented plan for how the model will be updated, retrained, and revalidated after initial clearance. For a deep dive on FDA's current stance on AI-driven medical devices, see our guide on LLM explainability for healthcare: meeting FDA and clinician expectations.
Even if your product does not require FDA clearance (clinical decision support that meets the exemption criteria), building your validation pipeline to FDA standards from day one is a competitive advantage and a risk mitigation strategy.
The Model Selection Decision Framework
After evaluating benchmarks, architectures, and deployment patterns, here is the practical framework for choosing your healthcare LLM. Answer these five questions in order — each answer narrows the option space.
Question 1: What is your GPU/API budget?
| Budget Tier | Recommended Path | Models |
|---|---|---|
| $0-500/month | Self-hosted open model (quantized) | MedGemma 4B, BioMistral 7B, Meditron 7B |
| $500-5,000/month | Self-hosted or cloud API | MedGemma 27B, OpenBioLLM 70B, GPT-4o via Azure |
| $5,000+/month | Production multi-model architecture | MedGemma (local) + GPT-4 (cloud) hybrid |
Question 2: Does your application process PHI?
If yes, your model must either run on infrastructure you control (on-premises, private cloud) or behind a BAA with a HIPAA-compliant cloud provider. This eliminates direct API access to OpenAI, Anthropic, and Google consumer endpoints. The remaining options are: self-hosted open models, Azure OpenAI, AWS Bedrock, Google Vertex AI, or AWS SageMaker with your own models.
Question 3: How narrow is your clinical domain?
Broad domain (general clinical AI, multi-specialty): Use a strong base model with RAG. MedGemma 27B or GPT-4 via Azure, augmented with retrieval from your clinical knowledge base.
Narrow domain (single specialty — dermatology, radiology, ophthalmology): Fine-tune a medical base model on your specialty data. Start with MedGemma or Meditron as the base and fine-tune with LoRA on specialty-specific clinical notes and guidelines.
Question 4: Do you need multimodal capabilities?
If your application processes medical images — radiology, pathology, dermatology, wound care — your options narrow to: MedGemma 1.5 (open-weight, self-hostable), GPT-4o via Azure (commercial, cloud), or Gemini 1.5 Pro via Vertex AI (commercial, cloud). No other models in the comparison table support medical imaging.
Question 5: How frequently does your knowledge base change?
Monthly or more frequently (drug formularies, insurance rules, pricing): RAG is mandatory. Retraining a model monthly is operationally impractical and expensive.
Quarterly or less (clinical guidelines, diagnostic criteria): Fine-tuning is viable. Retrain quarterly with updated guidelines incorporated into the training data.
The synthesis: For most healthcare AI teams in 2026, the optimal starting architecture is MedGemma 27B deployed on-premises via vLLM, augmented with RAG against a clinical knowledge base, with GPT-4 via Azure as a fallback for complex cases. This gives you benchmark-leading medical accuracy, full data control, manageable cost, and a quality ceiling for hard problems. Fine-tune only when you have validated that RAG alone is insufficient for your specific use case.
Frequently Asked Questions
Is MedGemma free for commercial use?
MedGemma is released under the Gemma License, which permits commercial use with some restrictions. It is "open-weight" rather than "open-source" — you can use and fine-tune the model weights commercially, but you must comply with Google's acceptable use policy, which prohibits use in applications that violate applicable laws or generate content to harm individuals. Read the full license before deploying. For most healthcare applications, the license is permissive enough for production use.
Can I use GPT-4 for clinical applications under HIPAA?
Yes, but only through Azure OpenAI Service with a signed BAA. The standard OpenAI API (api.openai.com) does not offer BAAs and is not HIPAA-compliant. Azure OpenAI provides the same models with data residency guarantees, no training on your data, and private endpoint options. The same applies to Claude (use AWS Bedrock, not the Anthropic API directly) and Gemini (use Vertex AI, not the Google AI Studio API).
How much GPU memory do I need to run MedGemma 27B?
In BF16 precision (full quality): approximately 60GB VRAM — a single A100 80GB or two A10G 24GB GPUs. In 4-bit quantization (AWQ or GPTQ): approximately 15-18GB VRAM — a single A10G or RTX 4090. Quantization reduces quality marginally (1-3% on benchmarks) but cuts memory requirements by 75%. For most clinical applications, the quality trade-off is acceptable. For detailed guidance on quantization strategies, see our model compression guide.
Should I fine-tune or use RAG for clinical note summarization?
Start with RAG plus careful prompt engineering. If your summarization output needs to match a specific institutional format (your hospital's discharge summary template, your specialty's referral letter format), then fine-tune on examples of your desired output format. The content accuracy comes from RAG (retrieving relevant patient data); the format consistency comes from fine-tuning. Most teams find the hybrid approach outperforms either method alone.
What is the minimum dataset size for clinically useful fine-tuning?
With LoRA on a medical base model like MedGemma, meaningful fine-tuning results start at approximately 1,000-5,000 high-quality examples for narrow tasks (formatting, classification) and 10,000-50,000 examples for broader clinical reasoning improvements. Quality matters far more than quantity — 5,000 expert-curated, clinician-validated examples outperform 50,000 noisy, inconsistently labeled examples. Always invest in data quality over data volume.
How do I evaluate whether my healthcare LLM is safe for production?
No single metric determines safety. Your validation pipeline must include: (1) benchmark regression testing to ensure general medical knowledge is preserved, (2) automated hallucination detection against verified medical knowledge bases, (3) bias testing across demographic axes, (4) clinician expert review of a statistically significant output sample, and (5) prospective monitoring with automatic escalation triggers after deployment. Build the monitoring infrastructure before you deploy, not after. Our guide on testing LLM outputs at scale provides the complete framework.
Choosing Your Clinical AI Model: What Matters Now
The healthcare LLM landscape in 2026 has reached an inflection point. MedGemma 1.5 has proven that open-weight models can match or exceed proprietary ones on medical benchmarks. The fine-tuning toolchain (LoRA, QLoRA, vLLM) has matured to the point where a two-person ML team can deploy a production clinical LLM. And the RAG pattern has become well-understood enough that teams without ML expertise can build effective clinical AI using off-the-shelf components.
But the technology is only half the challenge. Clinical validation, bias testing, hallucination detection, and regulatory compliance remain hard problems that no benchmark score resolves. The teams that win in healthcare AI are not the ones with the highest MedQA score — they are the ones with the most rigorous validation pipeline, the most thoughtful deployment architecture, and the clearest understanding of where their model's competence ends and clinician judgment begins.
Start with MedGemma and RAG. Validate relentlessly. Deploy incrementally. And never forget that the model serves the clinician, not the other way around.
Nirmitee builds clinical AI systems on FHIR-native architectures with rigorous validation pipelines. If you are evaluating healthcare LLMs for your product or need help deploying MedGemma in a HIPAA-compliant environment, let's talk.