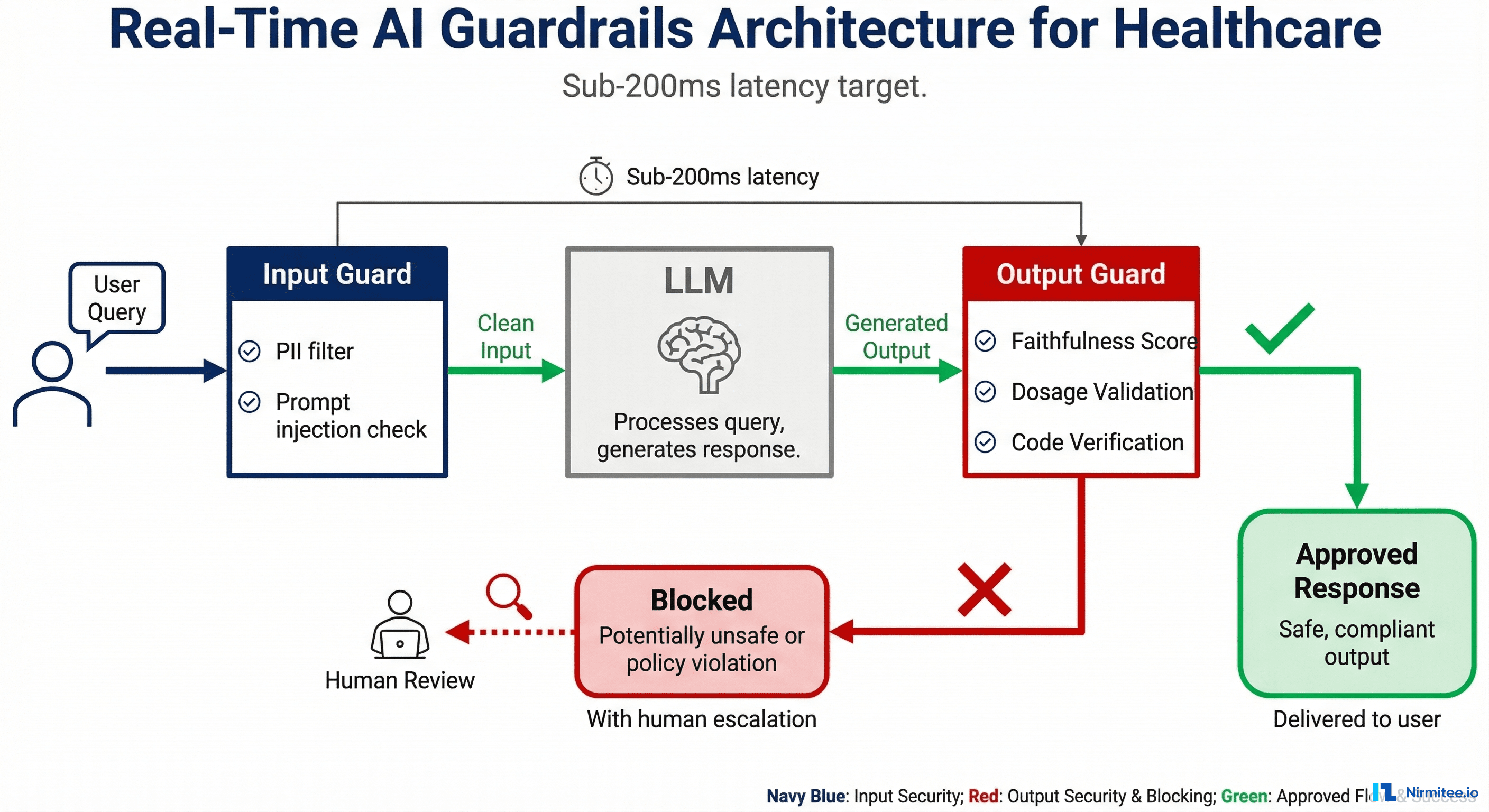
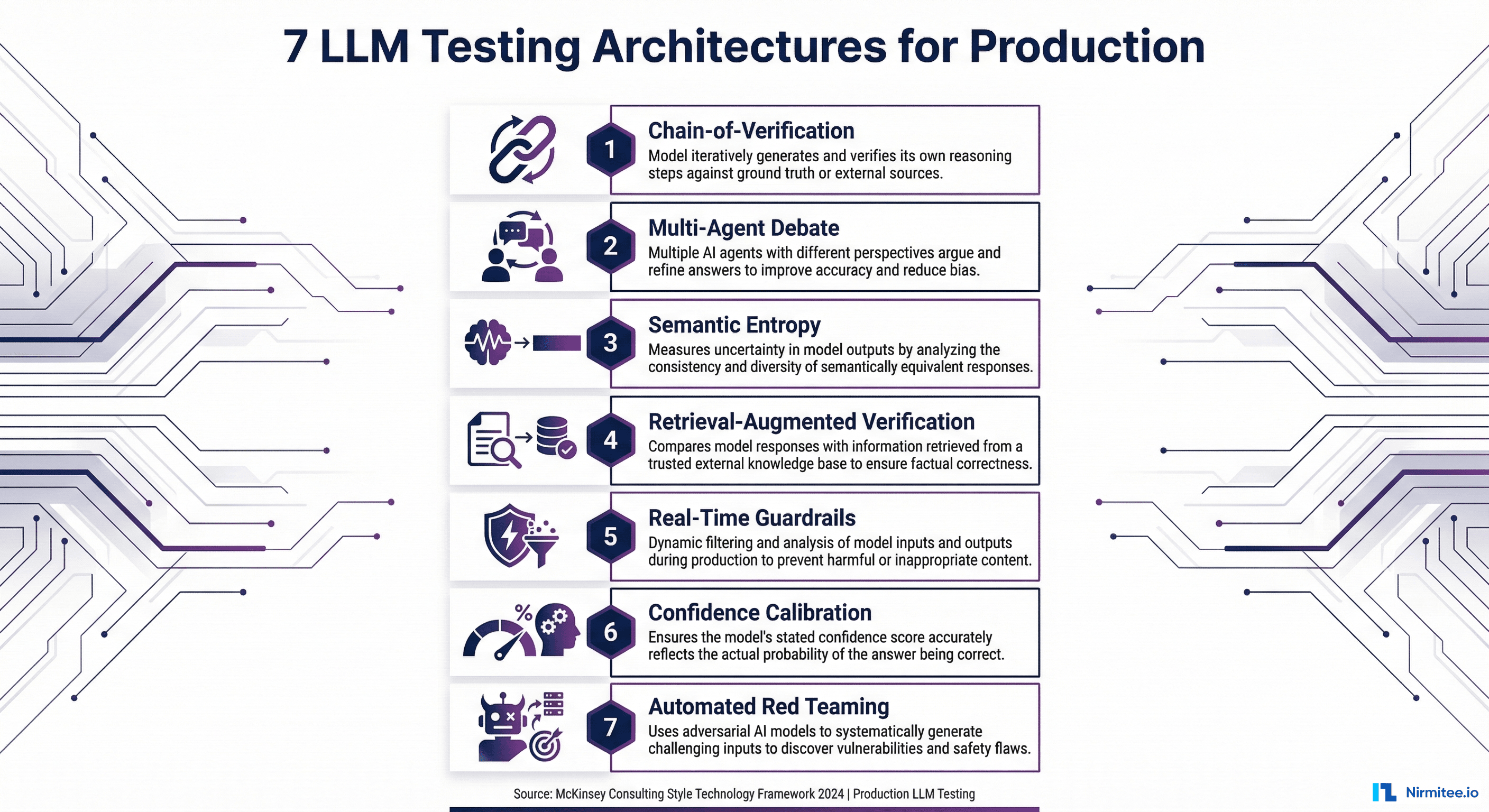
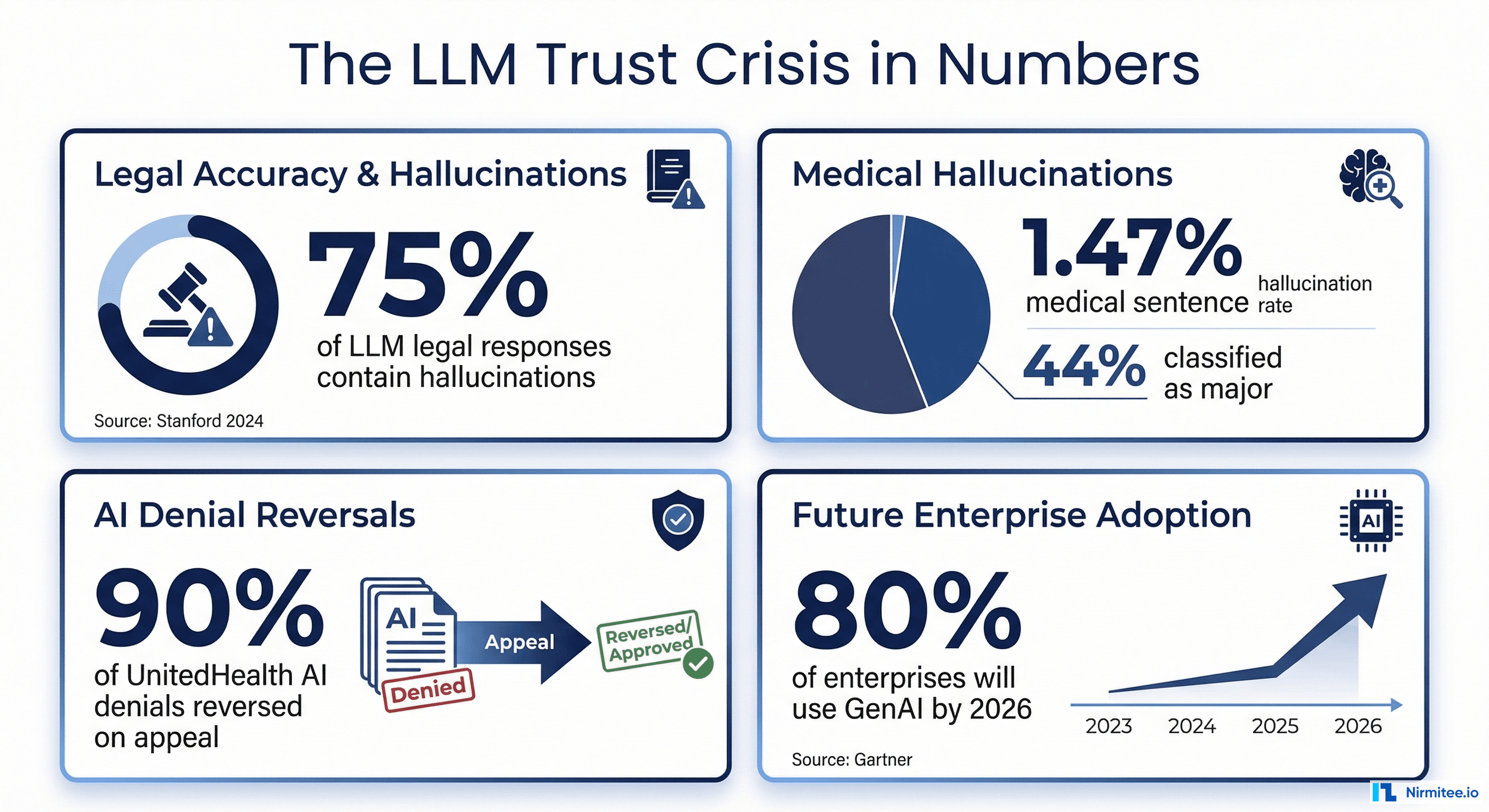
The Explainability Crisis: Healthcare AI That Can't Explain Itself Is Healthcare AI That Can't Be Trusted
A 2025 global survey of clinicians across 15 specialties found that 91.8% had encountered medical hallucinations in AI-generated outputs, and 84.7% believed those hallucinations were capable of causing direct patient harm. The study, published by researchers at MIT Media Lab, surveyed physicians over a 94-day period and included physician-led audits alongside benchmark analysis. Yet despite these alarming numbers, nearly 40% of respondents expressed a high degree of trust in AI outputs.
That gap between trust and verified accuracy is the explainability crisis. And it is not just a technical problem. It is a regulatory, clinical, and liability problem that is about to become significantly more expensive to ignore.
Even when an AI system produces an accurate output, clinicians will not act on it if they cannot see why it reached that conclusion. A confidence score of 0.87 tells a physician nothing. Was the recommendation based on the patient's lab results, a clinical guideline, or a statistical pattern from the training data? Without that context, the number is meaningless. Worse, it creates a false sense of precision.
Regulators have taken notice. The FDA's June 2024 guidance on Transparency for ML-Enabled Medical Devices explicitly requires plain-language descriptions of AI reasoning, including how AI helps achieve a device's intended use, the model inputs and outputs, and known limitations. The December 2024 final guidance on Predetermined Change Control Plans (PCCPs) mandates documentation of anticipated changes and validation protocols. In January 2025, the FDA published draft guidance on AI-Enabled Device Software Functions: Lifecycle Management, extending transparency requirements across the total product lifecycle.
On the European side, the EU AI Act classifies healthcare AI as high-risk and becomes fully applicable in August 2026, requiring technical documentation, conformity assessments, post-market monitoring, and documented human oversight mechanisms. Organizations deploying healthcare AI without explainability infrastructure will face both regulatory penalties and liability exposure.
The gap in most production AI systems is clear: they produce a confidence score but no explanation of what that score means, which evidence supports it, or when it should not be trusted. Closing that gap requires different explainability approaches for different audiences. That is what this guide covers.
Three Audiences, Three Types of Explainability
[IMAGE: https://res.cloudinary.com/dahyfknrf/image/upload/f_auto,q_auto/v1774148430/llm_explainability_three_layers_healthcare_fda_154eb28151.png]
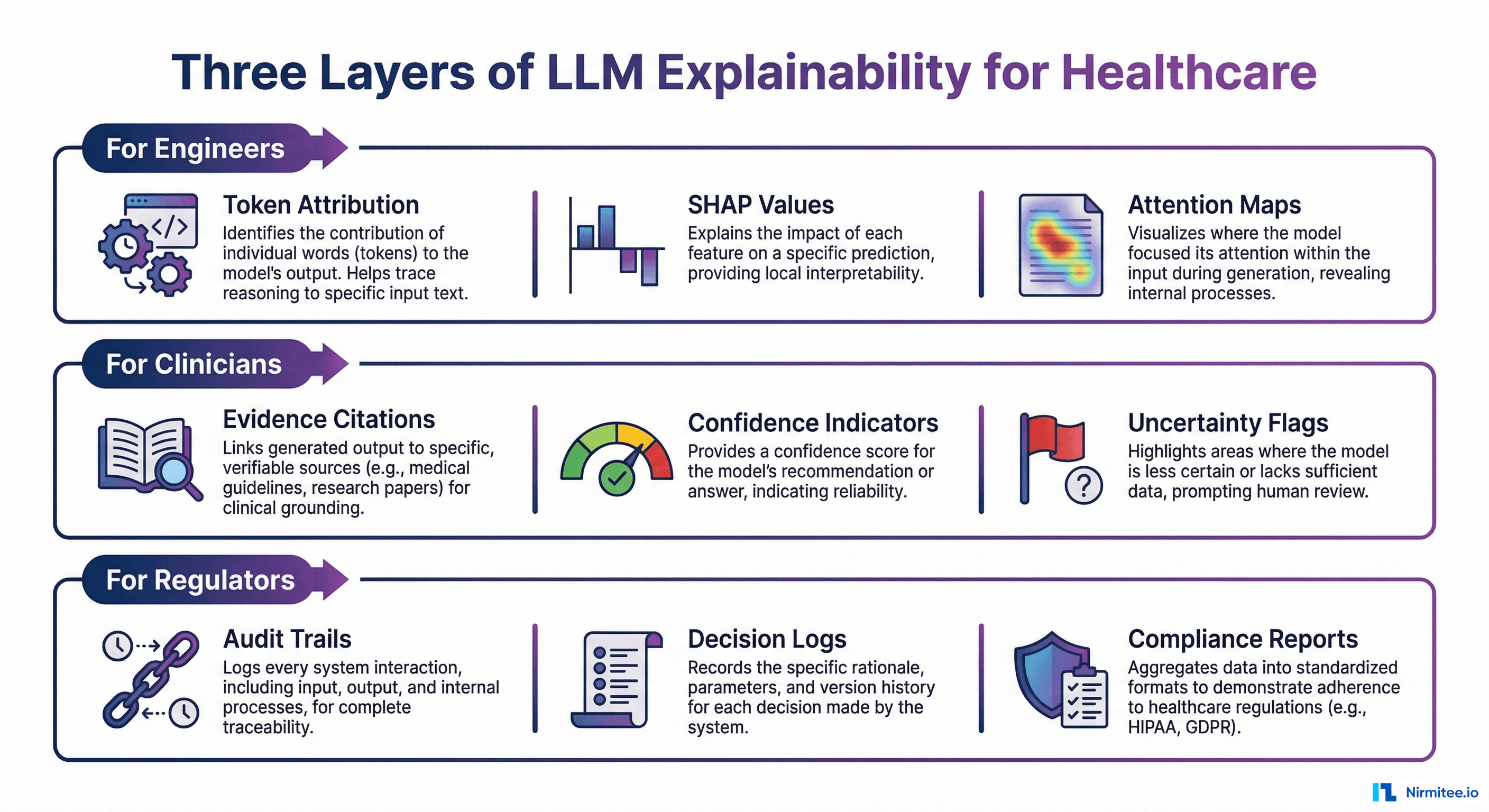
The fundamental mistake most teams make is building a single explainability layer and expecting it to serve every stakeholder. A token attribution map that helps an ML engineer debug a retrieval failure is useless to a cardiologist evaluating a treatment recommendation. An audit trail that satisfies an FDA reviewer provides no clinical decision support. Explainability must be designed for its audience.
For Engineers: Debug-Level Transparency
Engineers need to understand why the model produced a specific output at the mechanistic level. This means token attribution (SHAP values), attention maps, internal representation analysis, and retrieval diagnostics. When an LLM hallucinates a drug interaction that does not exist, engineers need to trace whether the failure was a retrieval problem (the wrong context was fetched), a generation problem (the model ignored correct context), or a training data problem (the model learned an incorrect association).
These tools are essential during development, fine-tuning, failure analysis, and regression testing. They are not meant for production-facing interfaces.
For Clinicians: Evidence-Based Explanations
Clinicians need to see the evidence chain: "This conclusion is based on these specific data points from these specific sources." They do not need attention weights or SHAP values. They need source references, claim-level confidence indicators, uncertainty flags, and clear statements about what the AI does and does not know.
The CLEVER framework (Clinical Large Language Model Evaluation by Expert Review), published in JMIR AI in February 2025, provides a validated methodology for evaluating clinical explainability. CLEVER uses blind, randomized, preference-based evaluation by practicing physicians across factuality, clinical relevance, and conciseness dimensions. Their findings are instructive: medical doctors preferred smaller, healthcare-specialized LLMs over GPT-4o by 45% to 92% on these dimensions, in part because the specialized models produced more grounded, citation-backed outputs rather than fluent but unsupported text.
The implication for explainability design is that clinicians value evidence provenance over confidence scores, and specificity over generality.
For Regulators and Auditors: Decision Audit Trails
Regulators need comprehensive documentation: decision audit trails, evaluation logs, performance monitoring reports, bias assessment records, and compliance documentation. These must be HIPAA-compliant, with 6-year retention per 45 CFR Section 164.316. They must satisfy FDA PCCP requirements, ONC HTI-1 source attribute mandates, and EU AI Act technical documentation requirements.
Every AI-assisted clinical decision must be logged with: the input data, the retrieved context, the model output, the confidence assessment, whether a human reviewed the output, and what action was taken. This is not optional. It is the minimum viable compliance posture for healthcare AI in 2026.
Claim-Level Evidence Chains: The Gold Standard for Healthcare Explainability
The most effective explainability pattern for clinical AI is the claim-level evidence chain. Instead of providing a single confidence score for an entire AI response, the system decomposes the response into individual claims and provides source evidence for each one.
How It Works
The pipeline follows four steps:
- Response generation — The AI system produces its output based on the retrieved context and the clinical query.
- Claim decomposition — The response is broken into individual factual assertions. "The patient should be prescribed metformin 500mg twice daily" becomes three separate claims: (a) metformin is the recommended medication, (b) the dosage should be 500mg, (c) the frequency should be twice daily.
- Evidence matching — Each claim is traced to the specific source data that supports it. Claim (a) maps to a clinical guideline for Type 2 diabetes first-line therapy. Claim (b) maps to the patient's A1C of 8.1% from lab results dated 03/15/2026 and the guideline's dosing recommendations. Claim (c) maps to the same guideline reference.
- Confidence scoring per claim — Each claim receives an individual confidence score based on the strength of the supporting evidence, rather than a single aggregate score for the entire response.
This is the same approach that the RAGAS faithfulness metric uses under the hood. RAGAS decomposes the AI response into individual claims and cross-checks each against the retrieved context, calculating faithfulness as the ratio of supported claims to total claims. But for clinical use, the raw metric is not enough. The output must be formatted so a clinician can inspect it in seconds.
[IMAGE: https://res.cloudinary.com/dahyfknrf/image/upload/f_auto,q_auto/v1774148440/confidence_score_display_clinician_interface_mockup_a38c1c32bf.png]
Structuring Claim-Level Evidence for Clinician Consumption
Here is a practical code structure for a claim-level evidence output that a clinician-facing interface can render:
from dataclasses import dataclass, field
from typing import List, Optional
from enum import Enum
class ConfidenceLevel(Enum):
HIGH = "high" # 0.95+ — strong evidence support
MODERATE = "moderate" # 0.80-0.94 — review recommended
LOW = "low" # below 0.80 — verify independently
@dataclass
class EvidenceSource:
source_type: str # "lab_result", "clinical_guideline", "patient_record"
reference: str # specific document or data point
excerpt: str # the relevant text or value
retrieval_score: float # how well this source matched the claim
@dataclass
class ExplainedClaim:
claim_text: str
confidence: float
confidence_level: ConfidenceLevel
supporting_evidence: List[EvidenceSource]
contradicting_evidence: List[EvidenceSource] = field(default_factory=list)
uncertainty_note: Optional[str] = None
@dataclass
class ExplainableResponse:
original_query: str
response_text: str
claims: List[ExplainedClaim]
overall_confidence: float
timestamp: str
model_version: str
audit_id: str # for compliance trail
def decompose_and_explain(query: str, response: str,
context_docs: list) -> ExplainableResponse:
"""
Core pipeline: decompose response into claims,
match each claim to source evidence,
score confidence per claim.
"""
# Step 1: Claim decomposition via LLM
claims = extract_claims(response)
# Step 2: Evidence matching per claim
explained_claims = []
for claim in claims:
evidence = match_evidence(claim, context_docs)
contradictions = find_contradictions(claim, context_docs)
confidence = calculate_claim_confidence(evidence, contradictions)
explained_claims.append(ExplainedClaim(
claim_text=claim,
confidence=confidence,
confidence_level=classify_confidence(confidence),
supporting_evidence=evidence,
contradicting_evidence=contradictions,
uncertainty_note=generate_uncertainty_note(confidence, evidence)
))
# Step 3: Build auditable response
return ExplainableResponse(
original_query=query,
response_text=response,
claims=explained_claims,
overall_confidence=compute_aggregate_confidence(explained_claims),
timestamp=datetime.utcnow().isoformat(),
model_version=get_model_version(),
audit_id=generate_audit_id()
)
The key design principle: every claim must be traceable to source evidence, and every evidence chain must be stored for audit retrieval. This is not just good engineering. Under ONC's HTI-1 rule, certified health IT that includes predictive decision support interventions must expose up to 31 source attributes, including training data provenance, external validation results, and quantitative performance measures. Claim-level evidence chains provide the infrastructure to meet these requirements.
SHAP and Feature Attribution for LLMs
While claim-level evidence chains serve clinicians, engineers need deeper mechanistic tools to understand and debug model behavior. SHAP (SHapley Additive exPlanations), originally developed for traditional ML models, has been adapted for transformer architectures and LLMs.
The SHAPLLM Framework
The SHAPLLM framework (arXiv 2024) integrates SHAP value computation with LLM-generated natural language narratives through a three-stage pipeline:
- SHAP computation — Calculate feature attribution scores that quantify how much each input token or feature contributed to the output. For a clinical note summarization task, this reveals which parts of the patient record most influenced the generated summary.
- Structured JSON formatting — Transform raw SHAP values into a structured representation that captures each feature's contribution direction (positive or negative), the feature value, and the SHAP magnitude.
- LLM-based narrative generation — Use a language model to translate the structured SHAP data into plain-language explanations. The framework uses instruction tuning to ensure each feature's contribution direction, value, and magnitude are described explicitly.
Token-Level Contribution Insights
The most practical application of SHAP in healthcare LLMs is debugging hallucinations. When SHAP analysis shows that the model attended heavily to irrelevant context passages while ignoring relevant clinical data, that signals a retrieval problem rather than a generation problem. The fix is in the retrieval pipeline (better chunking, improved embeddings, re-ranking), not in the model itself.
Conversely, when SHAP shows the model attended to the correct context but still produced an incorrect output, that indicates a generation-level issue that may require fine-tuning, prompt engineering, or guardrails.
import shap
import numpy as np
def analyze_token_attribution(model, tokenizer, input_text: str,
output_text: str) -> dict:
"""
Compute token-level SHAP values for an LLM response.
Identifies which input tokens most influenced the output.
"""
# Create SHAP explainer for the model
explainer = shap.Explainer(model, tokenizer)
# Compute SHAP values
shap_values = explainer([input_text])
# Extract token-level contributions
tokens = tokenizer.tokenize(input_text)
contributions = []
for i, token in enumerate(tokens):
contributions.append({
"token": token,
"shap_value": float(shap_values.values[0][i]),
"contribution": "positive" if shap_values.values[0][i] > 0
else "negative",
"magnitude": abs(float(shap_values.values[0][i]))
})
# Sort by absolute contribution magnitude
contributions.sort(key=lambda x: x["magnitude"], reverse=True)
# Flag potential retrieval issues
top_contributors = contributions[:10]
context_relevance = assess_context_relevance(top_contributors, output_text)
return {
"token_contributions": contributions,
"top_influencers": top_contributors,
"diagnostic": {
"retrieval_issue": not context_relevance["relevant_context_used"],
"generation_issue": context_relevance["relevant_context_used"]
and not context_relevance["output_faithful"],
"recommendation": context_relevance["fix_recommendation"]
}
}
Practical Limitations
SHAP computation for large language models is computationally expensive. For a 7B parameter model, computing full SHAP values for a single input can take minutes rather than milliseconds. This makes SHAP impractical for real-time production use but invaluable for offline debugging, failure analysis, and model evaluation during development cycles.
Both ISO/IEC 42001 (the international standard for AI management systems) and the NIST AI Risk Management Framework reference SHAP-compatible transparency mechanisms. ISO/IEC 42001 requires organizations to implement processes ensuring AI systems are understandable to stakeholders, with mechanisms explaining how specific recommendations were reached. The NIST AI RMF includes explainability scoring as a core trustworthiness characteristic, helping teams quantify how interpretable model decisions are for different audiences.
Confidence Scoring That Clinicians Actually Understand
The most common failure in healthcare AI interfaces is displaying a raw probability score — 0.87, 0.92, 0.73 — and expecting clinicians to know what to do with it. A probability score without context is not decision support. It is a liability.
The Traffic Light Framework
Translate confidence into actionable clinical indicators that map to clear workflows:
- GREEN (0.95 and above): High confidence. The AI recommendation has strong evidence support from multiple corroborating sources. The clinician can act on the recommendation with standard clinical judgment. Source evidence is displayed but does not require detailed review unless the clinician chooses to inspect it.
- YELLOW (0.80 to 0.94): Moderate confidence. The AI recommendation has partial evidence support or some evidence gaps. Review of the underlying evidence is recommended before acting. The interface highlights which specific claims have weaker evidence so the clinician knows exactly where to focus their review.
- RED (below 0.80): Low confidence. The AI system has identified significant uncertainty, contradictory evidence, or insufficient data to support the recommendation. The clinician must verify independently before acting. In high-stakes scenarios (medication dosing, surgical recommendations), red-zone outputs should be automatically escalated to a second clinician or specialist.
The "Trust or Escalate" Approach
The "Trust or Escalate" framework, published as an oral presentation at ICLR 2025, provides a principled approach to this problem. The method assesses the confidence of AI judge models and selectively decides when to trust their judgment versus when to escalate to human review. Their key innovation is "simulated annotators," a confidence estimation technique that significantly improves model calibration.
In practical terms, the system evaluates whether its own confidence meets a user-specified agreement threshold. If it does, the AI output is presented with full evidence chains. If it does not, the output is automatically routed to human review with a clear explanation of why confidence is low. Their experiments demonstrated that even cost-effective models like Mistral-7B can guarantee over 80% human agreement with nearly 80% test coverage using this cascaded selective evaluation.
Semantic Entropy: Detecting Uncertainty Through Consistency
A complementary approach to confidence estimation is semantic entropy, published in Nature in June 2024 by researchers at the University of Oxford. The method detects hallucinations by measuring uncertainty at the level of meaning rather than at the level of token sequences.
The insight is that when a model is confident about a fact, multiple generations will express the same meaning even if they use different words. When a model is uncertain or confabulating, multiple generations will diverge in meaning. By clustering generations by semantic equivalence and computing entropy over those clusters, the method reliably identifies outputs where the model lacks genuine knowledge.
Semantic entropy works across models (GPT-4, LLaMA 2, Falcon) and generalizes to tasks not seen during development, making it a robust production signal. Subsequent work on Semantic Entropy Probes reduced the computational overhead by approximating semantic entropy from hidden states of a single generation, enabling real-time deployment.
The key design principle for clinician-facing confidence: never show just a number — always show the evidence that produced the number. A confidence score of 0.91 accompanied by "based on 3 corroborating clinical guidelines, patient lab values from 2 visits, and no contradicting evidence" is actionable. The number alone is not.
Meeting FDA and EU AI Act Explainability Requirements
[IMAGE: https://res.cloudinary.com/dahyfknrf/image/upload/f_auto,q_auto/v1774148448/fda_eu_ai_act_explainability_requirements_2026_662e6186f8.png]
Explainability is no longer a nice-to-have. It is a compliance requirement across multiple regulatory frameworks that are converging on healthcare AI simultaneously.
FDA Requirements (2024-2025 Guidance)
Predetermined Change Control Plans (PCCPs). The December 2024 final guidance requires manufacturers to document what changes to AI/ML models are anticipated, the methodology for implementing those changes, and how they will be validated. This means explainability infrastructure must not only explain current model behavior but also document how explanations will change as the model evolves.
Transparency for ML-Enabled Medical Devices. The June 2024 guiding principles, issued jointly by the FDA, Health Canada, and the UK's MHRA, require clear communication of the degree to which information about an ML-enabled device — including its intended use, development, performance, and logic — is made available to relevant audiences. This explicitly includes plain-language descriptions of AI reasoning.
ONC HTI-1 Rule. Effective January 2025, the HTI-1 final rule establishes first-of-its-kind transparency requirements for AI in certified health IT. Predictive Decision Support Interventions (DSIs) must expose up to 31 source attributes covering training data provenance, external validation processes, quantitative performance measures, and fairness assessments. These attributes must be presented in plain language.
Lifecycle Management. The January 2025 draft guidance on AI-Enabled Device Software Functions extends transparency requirements across the total product lifecycle, including post-market performance monitoring and documentation of how the AI system's behavior changes over time.
EU AI Act Requirements (Applicable August 2026 for Healthcare)
The EU AI Act classifies healthcare AI as high-risk and imposes the most comprehensive set of explainability requirements globally:
- Risk assessment documentation — Formal assessment of risks associated with the AI system, including potential harms from unexplainable outputs.
- Technical documentation — Detailed documentation of the AI system's architecture, training methodology, evaluation methodology, and known limitations. This goes beyond what the FDA requires and includes specific documentation of how explainability mechanisms work.
- Post-market monitoring plans — Continuous monitoring of the AI system's performance in production, including monitoring for explainability degradation (when the system becomes less able to justify its outputs over time due to drift).
- Third-party conformity assessments — Healthcare AI systems in MDR risk class IIa or higher must undergo independent conformity assessment, which includes verification of explainability mechanisms.
- Human oversight mechanisms — Documented escalation procedures specifying when and how human reviewers override AI decisions, with audit trails for every override.
Practical Compliance Checklist
For organizations building or deploying HIPAA-compliant AI agents in healthcare, here is the minimum documentation to maintain per AI interaction:
- Input data (de-identified or with access controls per HIPAA)
- Retrieved context documents with source identifiers
- Model version and configuration at time of generation
- Full model output with claim decomposition
- Per-claim confidence scores and evidence chains
- Whether human review occurred and what action was taken
- Timestamp and unique audit identifier
- Retention: minimum 6 years per 45 CFR Section 164.316
Building an Explainability Pipeline: Implementation Guide
Moving from concept to production requires a layered approach. Here is a five-step implementation guide for building explainability into healthcare AI agent architectures.
Step 1: Choose Your Explainability Layers
Map each audience to the explainability methods they need:
EXPLAINABILITY_CONFIG = {
"clinician_layer": {
"methods": ["claim_decomposition", "evidence_matching",
"confidence_traffic_light"],
"output_format": "structured_clinical_display",
"latency_budget_ms": 500, # must be fast for clinical workflow
"always_on": True
},
"engineer_layer": {
"methods": ["shap_attribution", "attention_maps",
"retrieval_diagnostics"],
"output_format": "debug_json",
"latency_budget_ms": 30000, # offline analysis acceptable
"always_on": False, # triggered on failure or scheduled analysis
"trigger_conditions": ["confidence_below_0.80", "contradiction_detected",
"scheduled_evaluation"]
},
"auditor_layer": {
"methods": ["decision_audit_trail", "compliance_report",
"bias_assessment"],
"output_format": "compliance_document",
"storage_retention_years": 6, # HIPAA requirement
"always_on": True # every interaction must be logged
}
}
Step 2: Implement Claim Decomposition and Evidence Matching
The claim decomposition step uses a secondary LLM call (or a fine-tuned smaller model) to break the response into individual assertions, then matches each against the retrieved context:
from typing import List, Tuple
def decompose_claims(response_text: str, llm_client) -> List[str]:
"""Extract individual factual claims from AI response."""
prompt = f"""Decompose the following medical AI response into
individual factual claims. Each claim should be a single,
verifiable assertion.
Response: {response_text}
Return claims as a JSON array of strings."""
result = llm_client.generate(prompt, temperature=0.0)
return parse_json_array(result)
def match_evidence(claim: str, context_docs: List[dict],
embeddings_model) -> List[dict]:
"""Find supporting evidence for a specific claim."""
claim_embedding = embeddings_model.encode(claim)
matches = []
for doc in context_docs:
# Score each context passage against the claim
doc_embedding = embeddings_model.encode(doc["text"])
similarity = cosine_similarity(claim_embedding, doc_embedding)
if similarity > 0.75: # threshold for evidence relevance
matches.append({
"source": doc["metadata"]["source"],
"text": doc["text"],
"similarity": float(similarity),
"source_type": doc["metadata"].get("type", "unknown"),
"date": doc["metadata"].get("date")
})
return sorted(matches, key=lambda x: x["similarity"], reverse=True)
Step 3: Design the Clinician-Facing Confidence Display
The clinician interface must translate the claim-level evidence data into a format that supports rapid clinical decision-making. The display should show the AI recommendation prominently, with expandable evidence chains for each claim:
def render_clinical_display(explainable_response) -> dict:
"""Generate clinician-facing confidence display structure."""
display = {
"recommendation": explainable_response.response_text,
"overall_confidence": {
"score": explainable_response.overall_confidence,
"level": classify_confidence(
explainable_response.overall_confidence
).value,
"action": get_clinical_action(
explainable_response.overall_confidence
)
},
"claims": []
}
for claim in explainable_response.claims:
claim_display = {
"text": claim.claim_text,
"confidence_indicator": claim.confidence_level.value,
"evidence_summary": summarize_evidence(
claim.supporting_evidence
),
"evidence_count": len(claim.supporting_evidence),
"expandable_details": {
"sources": [
format_source(e) for e in claim.supporting_evidence
],
"contradictions": [
format_source(e) for e in claim.contradicting_evidence
] if claim.contradicting_evidence else None,
"uncertainty": claim.uncertainty_note
}
}
display["claims"].append(claim_display)
return display
def get_clinical_action(confidence: float) -> str:
if confidence >= 0.95:
return "Recommendation supported by strong evidence. " \
"Standard clinical judgment applies."
elif confidence >= 0.80:
return "Partial evidence support. Review highlighted " \
"claims before acting."
else:
return "Significant uncertainty detected. Independent " \
"verification required before acting."
Step 4: Build Audit Logging with HIPAA-Compliant Storage
Every AI interaction must be logged with sufficient detail for regulatory audit. The audit log must be immutable (append-only), encrypted at rest, and retained for a minimum of six years:
import hashlib
from datetime import datetime
def create_audit_record(explainable_response,
clinician_action: str) -> dict:
"""Create HIPAA-compliant audit record for AI interaction."""
record = {
"audit_id": explainable_response.audit_id,
"timestamp": datetime.utcnow().isoformat() + "Z",
"model_version": explainable_response.model_version,
"query_hash": hashlib.sha256(
explainable_response.original_query.encode()
).hexdigest(),
"response_hash": hashlib.sha256(
explainable_response.response_text.encode()
).hexdigest(),
"claim_count": len(explainable_response.claims),
"overall_confidence": explainable_response.overall_confidence,
"per_claim_confidence": [
{
"claim_index": i,
"confidence": c.confidence,
"level": c.confidence_level.value,
"evidence_count": len(c.supporting_evidence),
"contradiction_count": len(c.contradicting_evidence)
}
for i, c in enumerate(explainable_response.claims)
],
"clinician_action": clinician_action,
"human_reviewed": clinician_action != "auto_accepted",
"retention_expiry": calculate_retention_expiry(years=6)
}
# Store in append-only, encrypted audit log
store_audit_record(record, encrypt=True, immutable=True)
return record
Step 5: Automated Compliance Report Generation
Build automated report generation that aggregates audit data into formats required by FDA, EU AI Act, and internal governance. This reduces the manual burden of compliance documentation and ensures reports are always current:
def generate_compliance_report(start_date: str, end_date: str,
report_type: str) -> dict:
"""Generate regulatory compliance report from audit logs."""
audit_records = query_audit_logs(start_date, end_date)
report = {
"report_type": report_type,
"period": {"start": start_date, "end": end_date},
"summary": {
"total_interactions": len(audit_records),
"avg_confidence": mean([r["overall_confidence"]
for r in audit_records]),
"low_confidence_rate": sum(
1 for r in audit_records
if r["overall_confidence"] < 0.80
) / len(audit_records),
"human_review_rate": sum(
1 for r in audit_records if r["human_reviewed"]
) / len(audit_records),
},
"confidence_distribution": compute_distribution(
[r["overall_confidence"] for r in audit_records]
),
"escalation_analysis": analyze_escalations(audit_records),
"bias_assessment": run_bias_analysis(audit_records),
"model_drift_indicators": detect_drift(audit_records)
}
if report_type == "fda_pccp":
report["pccp_compliance"] = assess_pccp_requirements(
audit_records
)
elif report_type == "eu_ai_act":
report["conformity_assessment"] = assess_eu_requirements(
audit_records
)
return report
For teams building agentic AI workflows in healthcare, this pipeline integrates directly into the agent's decision loop. Every agent action that affects clinical outcomes should pass through the explainability pipeline before reaching the clinician.
Frequently Asked Questions
Does explainability slow down AI inference in production?
The clinician-facing layer (claim decomposition and evidence matching) adds 200-500ms of latency, which is acceptable for most clinical workflows where the AI is augmenting rather than replacing real-time decisions. SHAP-based engineer-level analysis adds significant overhead and should only run offline for debugging and scheduled evaluations. Audit logging is asynchronous and adds negligible latency to the user-facing response.
How does explainability relate to the LLM evaluation pipeline?
Explainability and LLM evaluation are complementary but distinct. Evaluation measures whether AI outputs are accurate, faithful, and safe across test datasets. Explainability shows why a specific output was produced for a specific input at a specific time. In practice, they share infrastructure: claim decomposition is used by both RAGAS faithfulness evaluation and by the explainability pipeline. Teams should build the shared infrastructure once and use it for both purposes.
What is the minimum viable explainability for FDA compliance?
Based on current FDA guidance, the minimum includes: plain-language descriptions of the AI's intended use and reasoning approach, documentation of model inputs, outputs, and known limitations, a predetermined change control plan if the model will be updated, and performance monitoring infrastructure. For ONC HTI-1 compliance, predictive DSIs must additionally expose up to 31 source attributes covering training data, validation, performance, and fairness. The EU AI Act requires all of the above plus third-party conformity assessment for high-risk healthcare AI systems.
Can smaller open-source models provide better explainability than large proprietary models?
The CLEVER framework research suggests yes, in certain contexts. In their evaluation, healthcare-specialized models with 8B and 70B parameters outperformed GPT-4o on factuality, clinical relevance, and conciseness as judged by practicing physicians. Smaller models fine-tuned on medical data tend to produce more grounded, citation-backed outputs. They are also easier to run SHAP analysis on due to lower computational requirements. Additionally, open-source models can be inspected at every layer, enabling deeper explainability analysis than is possible with API-only access to proprietary models. The trade-off is that they may have weaker general reasoning capabilities on edge cases outside their training distribution.
How should organizations handle AI outputs that fall in the "yellow zone" of moderate confidence?
Yellow-zone outputs represent the highest-value area for explainability investment. These are cases where the AI has partial evidence but not full certainty. The recommended approach is to surface the specific claims that have weaker evidence support, highlight any evidence gaps (for example, a missing recent lab result that would strengthen the recommendation), and provide the clinician with a one-click path to the most relevant source documents. Organizations should track yellow-zone override rates, where clinicians accept or reject moderate-confidence recommendations, and use this data to continuously calibrate confidence thresholds. Over time, these override patterns become training signal for improving the model. For guidance on evaluating the broader vendor and integration landscape, see our CTO framework for evaluating healthcare interoperability vendors.
Making AI Reasoning Visible Is Not Optional — It Is the Next Competitive Advantage
Healthcare organizations that build explainability into their AI infrastructure now will have a significant advantage as regulatory enforcement accelerates through 2026 and 2027. The FDA, EU AI Act, and ONC HTI-1 are converging on a clear expectation: healthcare AI must be able to explain itself to clinicians, auditors, and regulators in terms each audience can act on.
The organizations that treat explainability as an afterthought will face the same reckoning that organizations faced with HIPAA compliance in the early 2000s: retroactive infrastructure buildouts that cost 5-10x more than building it right from the start.
The technology exists. Claim-level evidence chains, semantic entropy, calibrated confidence scoring, and automated audit logging are all production-ready patterns. The question is not whether your healthcare AI needs to explain itself. It is whether you build that capability now, when it is a competitive advantage, or later, when it is a compliance emergency.
Building healthcare AI that needs to explain itself to clinicians, auditors, or the FDA?
Talk to Our Healthcare AI Team