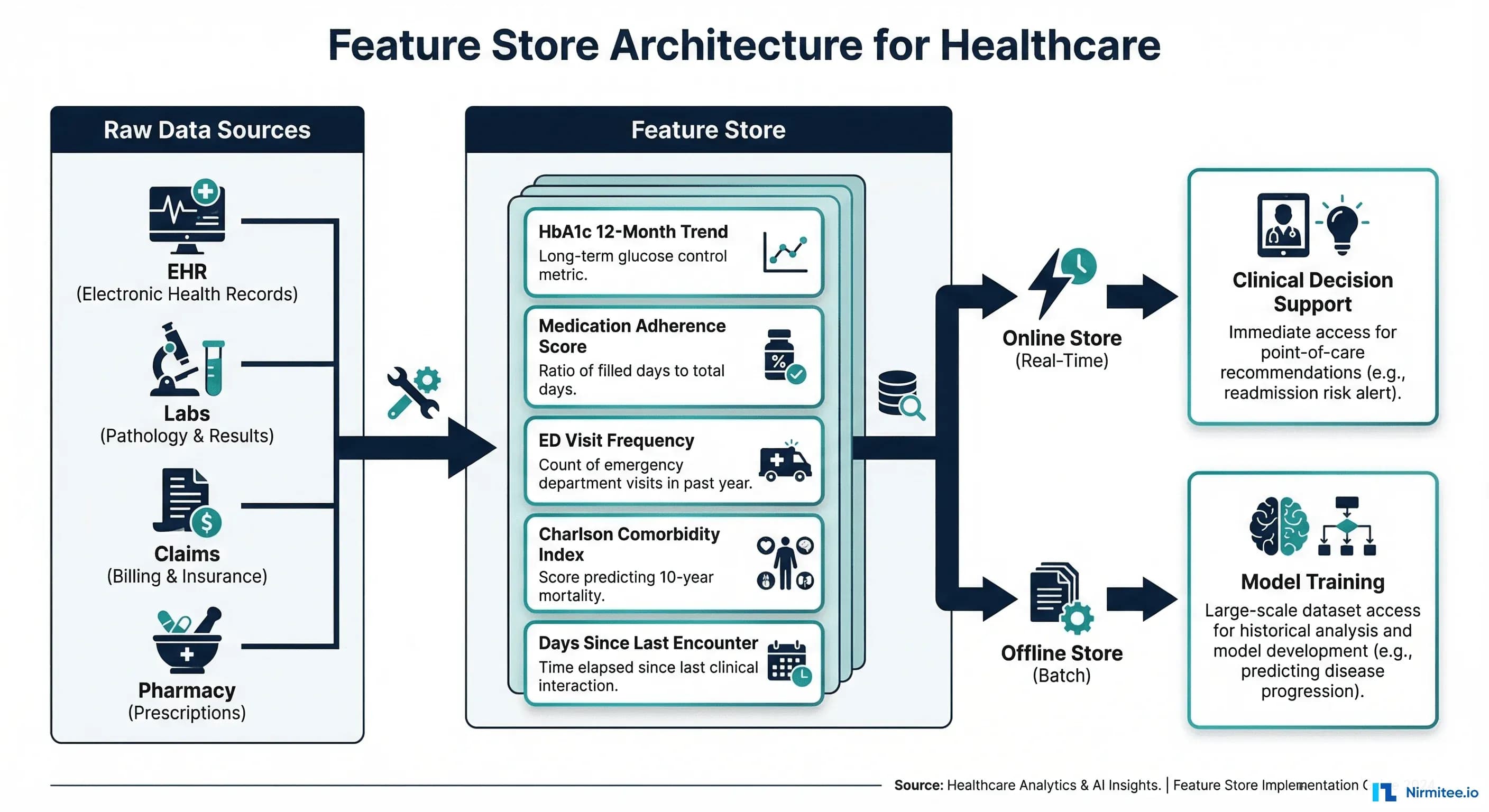
The Feature Engineering Bottleneck in Healthcare ML
Every healthcare ML project hits the same wall. Your data scientist spends three weeks writing SQL queries to extract HbA1c trends from the Observation table, compute medication adherence from MedicationRequest resources, and calculate Charlson comorbidity indices from Condition records. Then a second team building a different model needs the exact same features — and writes them from scratch. A third team deploys to production and discovers their real-time feature values do not match the training data because they computed features differently.
According to Tecton's research, data scientists spend 60-80% of their time on feature engineering rather than model development. In healthcare, this problem is amplified by the complexity of clinical data: FHIR resources are deeply nested, temporal patterns matter enormously (a single lab value means nothing without its trend), and regulatory requirements demand reproducibility across training and serving environments.
A feature store solves this by creating a centralized, versioned, governed repository of pre-computed clinical features that any model can reuse — with guarantees that training-time and serving-time features are computed identically.
This guide covers how to build a healthcare feature store using Feast (open-source), define clinical features from FHIR data, implement both online (real-time) and offline (batch) serving, and establish a feature catalog that accelerates every future ML project in your organization. If you are working with FHIR data pipelines, our guide on healthcare data quality as a prerequisite for AI provides essential context on data preparation.
What Is a Feature Store and Why Healthcare Needs One
A feature store is infrastructure that manages the lifecycle of ML features: computing them from raw data, storing them for both training and real-time inference, versioning them, and serving them with consistent semantics. Think of it as a database purpose-built for machine learning — where "tables" are feature groups and "rows" are entities (patients, encounters, providers).
Healthcare-specific reasons to invest in a feature store:
- Training-serving skew elimination: The number one cause of ML model degradation in production. If you compute "average blood pressure over 30 days" differently during training versus inference, your model predictions will be systematically wrong.
- Regulatory reproducibility: FDA SaMD guidelines and EU AI Act require that you can reproduce exactly how model inputs were derived. A feature store provides this audit trail automatically.
- Feature reuse across models: The Charlson comorbidity index is useful for readmission prediction, mortality risk, surgical outcome models, and resource allocation. Compute it once, reuse everywhere.
- Point-in-time correctness: Healthcare models must avoid data leakage. A feature store ensures that when you train on historical data, you only use features that were available at that point in time — not future lab results that arrived after the prediction window.
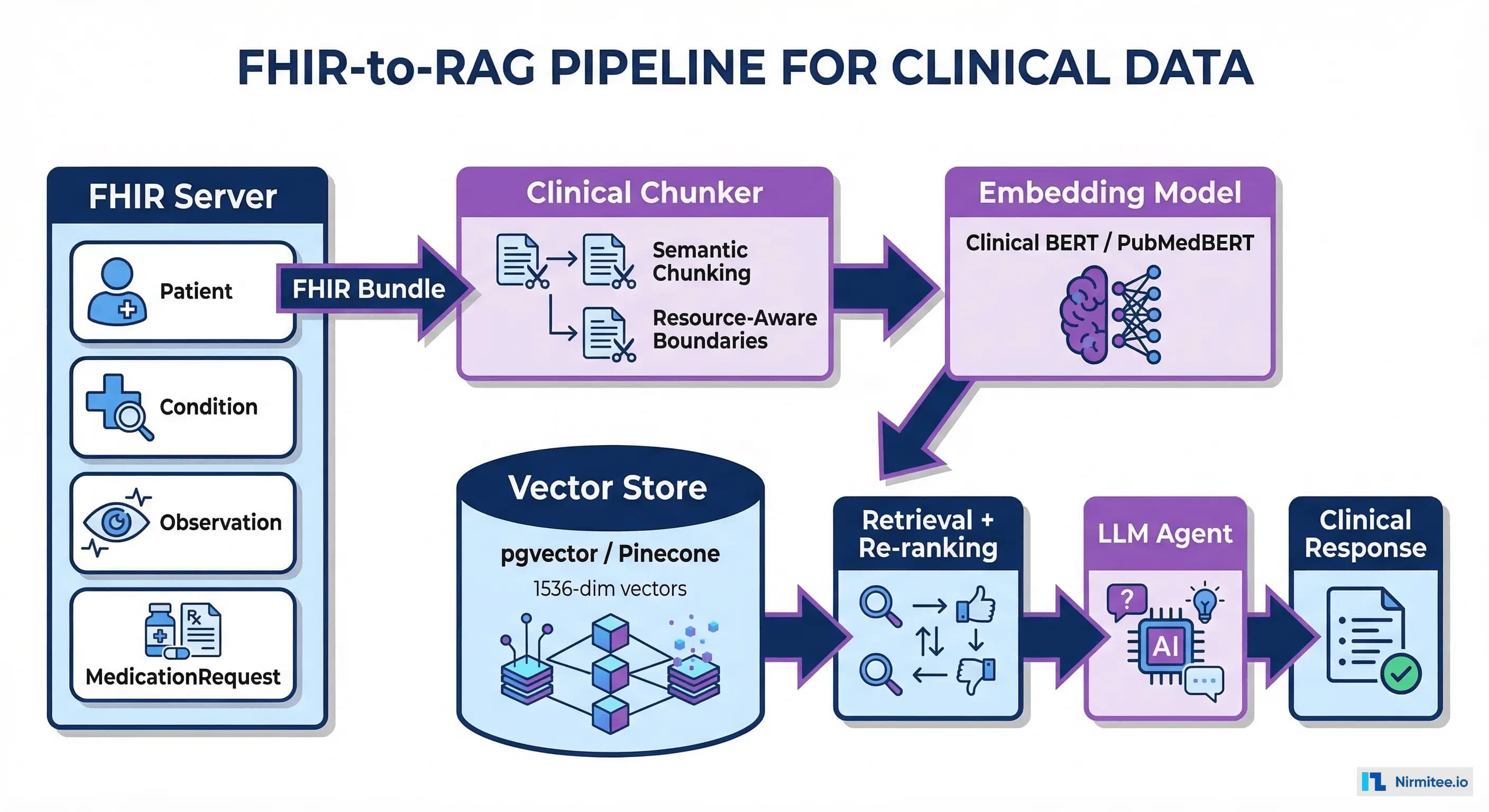
FHIR Resources to Clinical Features: The Mapping
The richness of FHIR makes it an excellent source for clinical features, but the mapping is not straightforward. Each FHIR resource type maps to a different category of features, and the temporal dimension is critical. Our guide on FHIR resource validation covers the data quality checks that must happen before feature extraction.
Observation to Lab Trend Features
Raw lab values are nearly useless for ML — trends are what matter. A single HbA1c of 7.2% tells you very little; three HbA1c values of 6.8, 7.0, 7.2 over 12 months tells you the patient is trending toward uncontrolled diabetes.
import pandas as pd
import numpy as np
from datetime import datetime, timedelta
from typing import Optional
class FHIRLabFeatureExtractor:
"""Extract trend features from FHIR Observation resources."""
def __init__(self, fhir_client):
self.client = fhir_client
def get_lab_observations(self, patient_id: str,
loinc_code: str,
months_back: int = 12) -> pd.DataFrame:
"""Fetch lab observations from FHIR server."""
cutoff = (datetime.now() - timedelta(days=months_back * 30)).isoformat()
bundle = self.client.search(
"Observation",
params={
"patient": patient_id,
"code": f"http://loinc.org|{loinc_code}",
"date": f"ge{cutoff}",
"_sort": "date"
}
)
rows = []
for entry in bundle.get("entry", []):
obs = entry["resource"]
value = obs.get("valueQuantity", {}).get("value")
date = obs.get("effectiveDateTime", "")
if value is not None and date:
rows.append({"date": date, "value": float(value)})
return pd.DataFrame(rows)
def compute_trend_features(self, patient_id: str,
loinc_code: str,
feature_prefix: str) -> dict:
"""Compute trend features from lab observations."""
df = self.get_lab_observations(patient_id, loinc_code)
if df.empty:
return {
f"{feature_prefix}_latest": None,
f"{feature_prefix}_mean_12m": None,
f"{feature_prefix}_slope_12m": None,
f"{feature_prefix}_count_12m": 0,
f"{feature_prefix}_cv": None,
}
df["date"] = pd.to_datetime(df["date"])
df = df.sort_values("date")
# Slope via linear regression (trend direction)
if len(df) >= 2:
days = (df["date"] - df["date"].iloc[0]).dt.days.values
slope = np.polyfit(days, df["value"].values, 1)[0]
slope_per_month = slope * 30
else:
slope_per_month = 0.0
return {
f"{feature_prefix}_latest": df["value"].iloc[-1],
f"{feature_prefix}_mean_12m": df["value"].mean(),
f"{feature_prefix}_slope_12m": slope_per_month,
f"{feature_prefix}_count_12m": len(df),
f"{feature_prefix}_cv": df["value"].std() / df["value"].mean()
if df["value"].mean() > 0 else 0,
}
# Usage: Extract HbA1c features
# extractor = FHIRLabFeatureExtractor(fhir_client)
# features = extractor.compute_trend_features(
# patient_id="patient-123",
# loinc_code="4548-4", # HbA1c LOINC code
# feature_prefix="hba1c"
# )MedicationRequest to Adherence Features
Medication adherence is one of the strongest predictors for hospital readmission and disease progression. The Proportion of Days Covered (PDC) metric is the industry standard, recommended by the Pharmacy Quality Alliance.
class MedicationAdherenceExtractor:
"""Compute adherence features from FHIR MedicationRequest."""
def compute_pdc(self, patient_id: str,
medication_class: str,
period_days: int = 180) -> dict:
"""Proportion of Days Covered (PDC) calculation."""
requests = self.client.search(
"MedicationRequest",
params={
"patient": patient_id,
"status": "active,completed",
"_sort": "-authoredon"
}
)
# Build coverage array
coverage = np.zeros(period_days)
gap_days = []
last_end = None
for entry in requests.get("entry", []):
med = entry["resource"]
start_str = med.get("dispenseRequest", {})\
.get("validityPeriod", {})\
.get("start", "")
supply_days = med.get("dispenseRequest", {})\
.get("expectedSupplyDuration", {})\
.get("value", 30)
if start_str:
start = pd.to_datetime(start_str)
start_idx = max(0, (datetime.now() - start).days)
end_idx = min(period_days, start_idx + int(supply_days))
coverage[start_idx:end_idx] = 1
if last_end is not None:
gap = start_idx - last_end
if gap > 0:
gap_days.append(gap)
last_end = end_idx
pdc = coverage.sum() / period_days
return {
f"med_adherence_pdc_{period_days}d": round(pdc, 4),
f"med_adherence_gap_max": max(gap_days) if gap_days else 0,
f"med_adherence_gap_count": len(gap_days),
f"med_active_count": int(coverage[-1]),
}Encounter to Utilization Features
class EncounterUtilizationExtractor:
"""Compute utilization features from FHIR Encounter resources."""
ENCOUNTER_TYPES = {
"emergency": "EMER",
"inpatient": "IMP",
"outpatient": "AMB",
"observation": "OBSENC"
}
def compute_utilization(self, patient_id: str,
window_days: int = 90) -> dict:
cutoff = (datetime.now() - timedelta(days=window_days)).isoformat()
encounters = self.client.search(
"Encounter",
params={
"patient": patient_id,
"date": f"ge{cutoff}",
"_sort": "-date"
}
)
counts = {t: 0 for t in self.ENCOUNTER_TYPES}
los_days = []
last_encounter_days = None
for entry in encounters.get("entry", []):
enc = entry["resource"]
enc_class = enc.get("class", [{}])[0].get("code", "")
for name, code in self.ENCOUNTER_TYPES.items():
if enc_class == code:
counts[name] += 1
# Length of stay
period = enc.get("period", {})
if period.get("start") and period.get("end"):
start = pd.to_datetime(period["start"])
end = pd.to_datetime(period["end"])
los = (end - start).days
los_days.append(los)
# Days since encounter
if last_encounter_days is None and period.get("start"):
last_encounter_days = (datetime.now() -
pd.to_datetime(period["start"])).days
return {
f"ed_visits_{window_days}d": counts["emergency"],
f"inpatient_admits_{window_days}d": counts["inpatient"],
f"outpatient_visits_{window_days}d": counts["outpatient"],
f"avg_los_days": np.mean(los_days) if los_days else 0,
f"days_since_last_encounter": last_encounter_days or 999,
f"total_encounters_{window_days}d": sum(counts.values()),
}Condition to Comorbidity Features
The Charlson Comorbidity Index (CCI) is one of the most widely used comorbidity measures in clinical research and predictive modeling, validated across hundreds of studies since its introduction in 1987.
class ComorbidityExtractor:
"""Compute Charlson Comorbidity Index from FHIR Condition."""
# Charlson weights by ICD-10 category
CHARLSON_WEIGHTS = {
"myocardial_infarction": (["I21", "I22", "I25.2"], 1),
"congestive_heart_failure": (["I09.9", "I11.0", "I13.0",
"I13.2", "I25.5", "I42",
"I43", "I50"], 1),
"peripheral_vascular": (["I70", "I71", "I73.1", "I73.8",
"I73.9", "I77.1", "I79"], 1),
"cerebrovascular": (["G45", "G46", "I60", "I61", "I62",
"I63", "I64", "I65", "I66", "I67",
"I68", "I69"], 1),
"diabetes_uncomplicated": (["E10.0", "E10.1", "E10.9",
"E11.0", "E11.1", "E11.9",
"E13"], 1),
"diabetes_complicated": (["E10.2", "E10.3", "E10.4",
"E10.5", "E11.2", "E11.3",
"E11.4", "E11.5"], 2),
"renal_disease": (["N01", "N03", "N05", "N07", "N18",
"N19", "N25"], 2),
"cancer": (["C0", "C1", "C2", "C3", "C4", "C5", "C6",
"C70", "C71", "C72", "C73", "C74", "C75",
"C76"], 2),
"metastatic_cancer": (["C77", "C78", "C79", "C80"], 6),
"severe_liver_disease": (["K70.4", "K71.1", "K72.1",
"K72.9", "K76.5", "K76.6",
"K76.7"], 3),
"hiv_aids": (["B20", "B21", "B22", "B24"], 6),
}
def compute_cci(self, patient_id: str) -> dict:
conditions = self.client.search(
"Condition",
params={
"patient": patient_id,
"clinical-status": "active",
}
)
icd_codes = set()
for entry in conditions.get("entry", []):
cond = entry["resource"]
for coding in cond.get("code", {}).get("coding", []):
if "icd" in coding.get("system", "").lower():
icd_codes.add(coding["code"])
# Calculate CCI score
cci_score = 0
matched_categories = []
for category, (prefixes, weight) in self.CHARLSON_WEIGHTS.items():
for code in icd_codes:
if any(code.startswith(p) for p in prefixes):
cci_score += weight
matched_categories.append(category)
break
return {
"charlson_index": cci_score,
"comorbidity_count": len(matched_categories),
"active_condition_count": len(icd_codes),
"has_diabetes": 1 if any("diabetes" in c
for c in matched_categories) else 0,
"has_cancer": 1 if any("cancer" in c
for c in matched_categories) else 0,
}Feature Store Tools Comparison
The feature store ecosystem has matured significantly since 2020. Here is how the major tools compare for healthcare use cases:
| Tool | License | Online Store | Offline Store | HIPAA Compliant | Best For |
|---|---|---|---|---|---|
| Feast | Apache 2.0 | Redis, DynamoDB, Postgres | BigQuery, Redshift, S3 | Self-hosted: Yes | Teams wanting full control and vendor independence |
| Tecton | Commercial | DynamoDB (managed) | Spark + S3 | Yes (BAA available) | Enterprise real-time ML at scale |
| Hopsworks | AGPL / Commercial | RonDB | Hive, Spark | Self-hosted: Yes | End-to-end ML platform with strong Python SDK |
| SageMaker Feature Store | AWS Service | API Gateway + DynamoDB | S3 + Athena | Yes (BAA available) | AWS-native teams already using SageMaker |
| Vertex AI Feature Store | GCP Service | Bigtable | BigQuery | Yes (BAA available) | GCP-native teams with BigQuery data |
For healthcare teams, Feast is typically the best starting point: it is open-source, can be self-hosted for HIPAA compliance, integrates with any storage backend, and does not lock you into a cloud vendor. The rest of this guide uses Feast for implementation examples.
Online vs. Offline Feature Serving
Understanding the distinction between online and offline serving is critical for healthcare, where different use cases have vastly different latency requirements.
| Dimension | Online Store | Offline Store |
|---|---|---|
| Latency | <10ms | Seconds to minutes |
| Data | Latest feature values only | Full historical timeline |
| Use Case | Real-time clinical decision support | Model training, batch scoring, analytics |
| Storage | Redis, DynamoDB, Bigtable | S3, BigQuery, Parquet files |
| Healthcare Example | Sepsis risk score at bedside | Monthly readmission model retraining |
| Update Frequency | Minutes to hours (materialized) | Append-only (event sourced) |
Implementing a Healthcare Feature Store with Feast
Step 1: Define Feature Views
Feast uses Python files to declare features, their data sources, and their serving configuration. Here is a complete feature definition for clinical features derived from FHIR data:
# feature_repo/clinical_features.py
from feast import (
Entity, Feature, FeatureView, FileSource,
ValueType, Field
)
from feast.types import Float64, Int64, String
from datetime import timedelta
# Entity: the patient
patient = Entity(
name="patient_id",
description="FHIR Patient resource ID",
value_type=ValueType.STRING,
)
# Data source: Parquet files generated by FHIR pipeline
lab_features_source = FileSource(
path="s3://healthcare-features/lab_features.parquet",
timestamp_field="event_timestamp",
created_timestamp_column="created_timestamp",
)
# Feature View: Lab trend features
lab_trend_features = FeatureView(
name="lab_trend_features",
entities=[patient],
ttl=timedelta(hours=24), # Features expire after 24h
schema=[
Field(name="hba1c_latest", dtype=Float64),
Field(name="hba1c_mean_12m", dtype=Float64),
Field(name="hba1c_slope_12m", dtype=Float64),
Field(name="hba1c_count_12m", dtype=Int64),
Field(name="egfr_latest", dtype=Float64),
Field(name="egfr_slope_12m", dtype=Float64),
Field(name="creatinine_latest", dtype=Float64),
Field(name="wbc_latest", dtype=Float64),
Field(name="wbc_cv", dtype=Float64),
],
source=lab_features_source,
online=True, # Materialize to online store
tags={"team": "clinical-ml", "domain": "labs"},
)
# Feature View: Medication adherence features
med_adherence_source = FileSource(
path="s3://healthcare-features/med_adherence.parquet",
timestamp_field="event_timestamp",
)
medication_features = FeatureView(
name="medication_adherence_features",
entities=[patient],
ttl=timedelta(hours=12),
schema=[
Field(name="med_adherence_pdc_180d", dtype=Float64),
Field(name="med_adherence_gap_max", dtype=Int64),
Field(name="med_adherence_gap_count", dtype=Int64),
Field(name="med_active_count", dtype=Int64),
],
source=med_adherence_source,
online=True,
tags={"team": "clinical-ml", "domain": "medications"},
)
# Feature View: Encounter utilization features
encounter_source = FileSource(
path="s3://healthcare-features/encounter_util.parquet",
timestamp_field="event_timestamp",
)
utilization_features = FeatureView(
name="encounter_utilization_features",
entities=[patient],
ttl=timedelta(hours=6),
schema=[
Field(name="ed_visits_90d", dtype=Int64),
Field(name="inpatient_admits_90d", dtype=Int64),
Field(name="avg_los_days", dtype=Float64),
Field(name="days_since_last_encounter", dtype=Int64),
Field(name="total_encounters_90d", dtype=Int64),
],
source=encounter_source,
online=True,
tags={"team": "clinical-ml", "domain": "encounters"},
)
# Feature View: Comorbidity features
comorbidity_source = FileSource(
path="s3://healthcare-features/comorbidity.parquet",
timestamp_field="event_timestamp",
)
comorbidity_features = FeatureView(
name="comorbidity_features",
entities=[patient],
ttl=timedelta(days=7), # Comorbidities change slowly
schema=[
Field(name="charlson_index", dtype=Int64),
Field(name="comorbidity_count", dtype=Int64),
Field(name="active_condition_count", dtype=Int64),
Field(name="has_diabetes", dtype=Int64),
Field(name="has_cancer", dtype=Int64),
],
source=comorbidity_source,
online=True,
tags={"team": "clinical-ml", "domain": "conditions"},
)Step 2: Feature Engineering Pipeline
The pipeline that populates the feature store runs on a schedule, extracts data from FHIR, computes features, validates them, and writes Parquet files that Feast ingests. For teams already running event-driven pipelines, this integrates well with the patterns described in our guide on event-driven EHR architecture with FHIR Subscriptions and Kafka.
import pandas as pd
from datetime import datetime
class ClinicalFeaturePipeline:
"""End-to-end pipeline: FHIR -> Features -> Parquet -> Feast."""
def __init__(self, fhir_client, output_path: str):
self.fhir_client = fhir_client
self.output_path = output_path
self.lab_extractor = FHIRLabFeatureExtractor(fhir_client)
self.med_extractor = MedicationAdherenceExtractor()
self.encounter_extractor = EncounterUtilizationExtractor()
self.comorbidity_extractor = ComorbidityExtractor()
def run_for_patient(self, patient_id: str) -> dict:
"""Compute all features for a single patient."""
features = {"patient_id": patient_id,
"event_timestamp": datetime.now()}
# Lab trends
for loinc, prefix in [
("4548-4", "hba1c"), # HbA1c
("33914-3", "egfr"), # eGFR
("2160-0", "creatinine"),
("6690-2", "wbc"), # White blood cell count
]:
features.update(
self.lab_extractor.compute_trend_features(
patient_id, loinc, prefix
)
)
# Medication adherence
features.update(
self.med_extractor.compute_pdc(patient_id, "all")
)
# Encounter utilization
features.update(
self.encounter_extractor.compute_utilization(patient_id)
)
# Comorbidities
features.update(
self.comorbidity_extractor.compute_cci(patient_id)
)
return features
def run_batch(self, patient_ids: list) -> pd.DataFrame:
"""Compute features for all patients and write Parquet."""
all_features = []
for pid in patient_ids:
try:
features = self.run_for_patient(pid)
all_features.append(features)
except Exception as e:
print(f"Error for {pid}: {e}")
df = pd.DataFrame(all_features)
# Validate before writing
self._validate(df)
# Write domain-specific Parquet files
lab_cols = [c for c in df.columns if any(
c.startswith(p) for p in ["hba1c", "egfr", "creatinine", "wbc"]
)]
df[["patient_id", "event_timestamp"] + lab_cols].to_parquet(
f"{self.output_path}/lab_features.parquet"
)
return df
def _validate(self, df: pd.DataFrame):
"""Validate feature quality before ingestion."""
null_pct = df.isnull().mean()
high_null = null_pct[null_pct > 0.30]
if not high_null.empty:
print(f"WARNING: High null rate: {high_null.to_dict()}")
# Range checks for clinical validity
if "hba1c_latest" in df.columns:
invalid = df[
(df["hba1c_latest"] < 3) | (df["hba1c_latest"] > 20)
]
if not invalid.empty:
print(f"WARNING: {len(invalid)} invalid HbA1c values")Step 3: Deploy and Materialize
# Initialize Feast project
feast init healthcare_features
cd healthcare_features
# Copy feature definitions
cp clinical_features.py feature_repo/
# Apply feature definitions to registry
feast apply
# Materialize features to online store
# This copies latest values from offline to online store
feast materialize-incremental $(date -u +"%Y-%m-%dT%H:%M:%S")
# Start the feature server for real-time serving
feast serve --host 0.0.0.0 --port 6566Step 4: Consume Features for Training and Inference
from feast import FeatureStore
import pandas as pd
store = FeatureStore(repo_path="feature_repo/")
# OFFLINE: Get historical features for training
# Point-in-time correct — no data leakage
entity_df = pd.DataFrame({
"patient_id": ["patient-001", "patient-002", "patient-003"],
"event_timestamp": [
pd.Timestamp("2025-06-15"),
pd.Timestamp("2025-06-15"),
pd.Timestamp("2025-06-15"),
]
})
training_df = store.get_historical_features(
entity_df=entity_df,
features=[
"lab_trend_features:hba1c_latest",
"lab_trend_features:hba1c_slope_12m",
"medication_adherence_features:med_adherence_pdc_180d",
"encounter_utilization_features:ed_visits_90d",
"comorbidity_features:charlson_index",
]
).to_df()
print(training_df.head())
# ONLINE: Get latest features for real-time inference
online_features = store.get_online_features(
features=[
"lab_trend_features:hba1c_latest",
"lab_trend_features:hba1c_slope_12m",
"medication_adherence_features:med_adherence_pdc_180d",
"encounter_utilization_features:ed_visits_90d",
"comorbidity_features:charlson_index",
],
entity_rows=[{"patient_id": "patient-001"}]
).to_dict()
print(online_features)Building a Clinical Feature Catalog
A feature catalog is the human-readable index of your feature store — it tells data scientists what features are available, how they are computed, and when they should (and should not) be used. Without a catalog, feature reuse drops and teams duplicate effort. This is similar to how a well-maintained FHIR CapabilityStatement tells developers what an API can do.
| Feature Name | FHIR Source | Computation | Update Frequency | Models Using |
|---|---|---|---|---|
| hba1c_slope_12m | Observation (4548-4) | Linear regression slope of last 12 months | Daily | Diabetes risk, Readmission |
| med_adherence_pdc_180d | MedicationRequest | Proportion of Days Covered, 180-day window | Weekly | Readmission, Chronic disease |
| ed_visits_90d | Encounter (EMER) | Count of ED encounters in 90 days | Daily | Readmission, Risk stratification |
| charlson_index | Condition | Weighted sum of 17 comorbidity categories | Monthly | Mortality, Surgical risk, Cost |
| days_since_last_pcp | Encounter (AMB) | Days since last outpatient primary care | Daily | Care gap detection, Readmission |
| egfr_slope_12m | Observation (33914-3) | eGFR trend over 12 months | Daily | CKD progression, Nephrology referral |
| active_med_count | MedicationRequest | Count of active medications | Real-time | Polypharmacy risk, Drug interaction |
| bmi_trend_6m | Observation (39156-5) | BMI slope over 6 months | Weekly | Metabolic risk, Bariatric screening |
Frequently Asked Questions
Do I need a feature store if I only have one ML model?
Probably not yet — but you should build your feature engineering code as if you will have a feature store. Structure features into logical groups (labs, medications, encounters), use consistent naming conventions, and ensure training and serving code share the same feature computation functions. When you add a second model, migrating to a feature store will be straightforward.
How do feature stores handle HIPAA compliance?
Feature stores themselves are storage and compute infrastructure — HIPAA compliance depends on where they run. Self-hosted Feast on your own infrastructure inherits your existing HIPAA controls. Cloud feature stores (SageMaker, Vertex AI) offer BAAs. Key requirements: encrypt features at rest and in transit, implement access controls per feature group (not all data scientists need access to all features), audit all feature access, and ensure PHI features are properly de-identified for research use cases.
What is training-serving skew and how does a feature store prevent it?
Training-serving skew occurs when features used during model training are computed differently than features used during inference. For example, training code calculates "average blood pressure" using a pandas rolling mean, but the serving code uses a SQL window function with slightly different boundary handling. The results differ by small amounts, but the model was trained on one distribution and sees another. A feature store prevents this by ensuring both training and serving read from the same computed feature values.
How do you handle feature freshness for real-time clinical decisions?
For truly real-time features (e.g., current heart rate for ICU monitoring), feature stores alone are insufficient. Use a streaming feature pipeline (Kafka + Flink/Spark Streaming) that computes features from real-time HL7/FHIR events and writes directly to the online store. For features that tolerate minutes of staleness (e.g., medication adherence, comorbidity index), batch materialization every 15-60 minutes is sufficient.
Can feature stores work with unstructured clinical data?
Feature stores excel at structured, tabular features. For unstructured data (clinical notes, radiology reports), the common pattern is to run NLP/ML embedding models first to extract structured features (sentiment scores, entity counts, embedding vectors), then store those derived features in the feature store. The raw unstructured data stays in its original storage. See our guide on unstructured data pipelines for healthcare AI for the extraction side.
How do you version features when clinical definitions change?
Feature stores support versioning through feature view names and tags. When a feature definition changes (e.g., switching from ICD-9 to ICD-10 for comorbidity scoring), create a new feature view version (comorbidity_features_v2) rather than modifying the existing one. Keep the old version available for models still in production. Deprecate old versions only after all dependent models have been retrained on the new features.