A radiology group spent $180,000 fine-tuning GPT-4 on chest X-ray reports. Six months later, they scrapped the project — not because the model performed poorly, but because a well-crafted RAG pipeline with their existing report templates achieved 94% of the accuracy at 3% of the cost.
Fine-tuning large language models for clinical use cases is one of the most overhyped and simultaneously underutilized capabilities in health technology. Most teams that should fine-tune do not, and most teams that do fine-tune should not have. The difference comes down to understanding when fine-tuning actually provides clinical value that simpler approaches cannot match.
This guide provides a technical framework for making that decision. We cover when fine-tuning is the right approach versus prompt engineering or retrieval-augmented generation, how to prepare clinical data for training, which base models to select, how to evaluate clinical AI with metrics that actually matter, cost analysis at every level, and the FDA regulatory implications of deploying a fine-tuned model in a clinical environment.
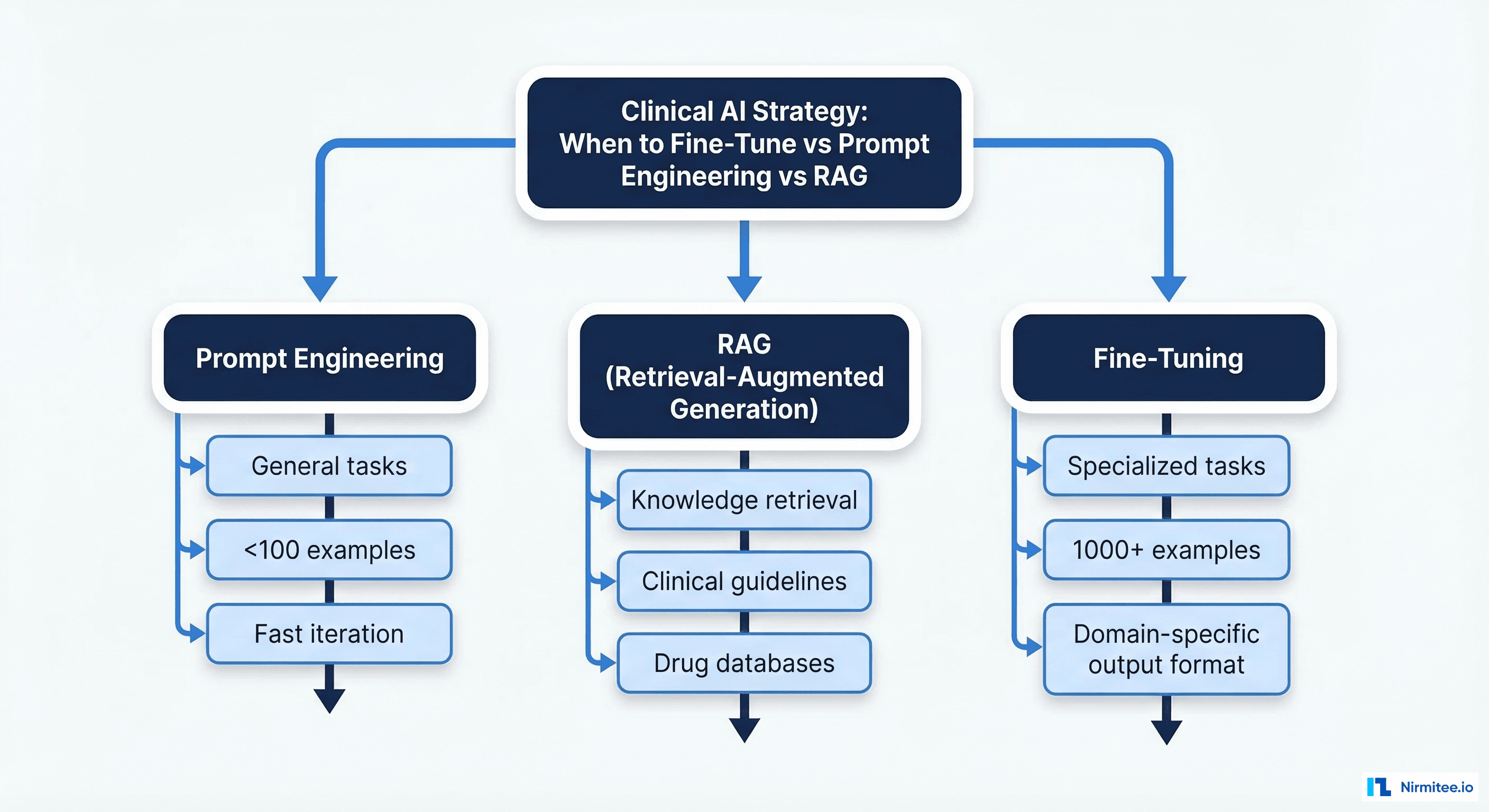
The Three Approaches: Fine-Tuning vs. Prompt Engineering vs. RAG
Before investing in fine-tuning, you need to understand where it sits in the clinical AI toolbox. Each approach solves a different problem, and choosing wrong wastes months and hundreds of thousands of dollars.
Prompt Engineering: The Starting Point
Prompt engineering means crafting input instructions that steer a general-purpose LLM toward clinical outputs without modifying the model itself. This works when:
- The task is general enough that a foundation model already has the knowledge (summarizing clinical notes, answering patient questions about medications)
- You have fewer than 100 example cases
- Output format requirements are flexible
- You need to iterate rapidly (hours, not weeks)
Prompt engineering fails when the model needs to produce outputs in highly specific clinical formats (ICD-10 coding, structured FHIR resources), when it needs knowledge that is not in the training data (proprietary clinical protocols, institution-specific formularies), or when consistency across thousands of outputs is critical.
RAG: Knowledge Retrieval Without Retraining
Retrieval-augmented generation combines a vector database of clinical documents with a foundation model. The model retrieves relevant context at inference time and generates responses grounded in that retrieved knowledge. RAG excels when:
- The knowledge base changes frequently (drug databases, clinical guidelines, insurance formularies)
- You need citations and traceability back to source documents
- The task is primarily knowledge retrieval rather than reasoning
- You want to avoid the cost and complexity of model training
RAG fails when the task requires deep reasoning over clinical patterns (predicting patient trajectories, identifying subtle diagnostic signals), when the output must conform to a rigid schema that the base model cannot reliably produce, or when latency is critical (RAG adds 200-500ms for retrieval).
Fine-Tuning: When the Model Needs New Capabilities
Fine-tuning modifies the model's weights using domain-specific training data, teaching it patterns, formats, and reasoning strategies that the base model does not possess. Fine-tuning is the right choice when:
- You need consistent output in a domain-specific format (structured clinical notes, FHIR-compliant observations, coded diagnoses)
- The task requires clinical reasoning patterns not present in general training data
- You have 1,000+ high-quality annotated examples
- Accuracy requirements exceed what prompt engineering can achieve (typically the 90%+ threshold)
- You are building a product where model performance is a core differentiator
| Approach | Best For | Data Required | Cost | Time to Deploy |
|---|---|---|---|---|
| Prompt Engineering | General clinical tasks | 10-50 examples | $0-$100 | Hours to days |
| RAG | Knowledge retrieval | Document corpus | $200-$2,000/mo | 1-2 weeks |
| LoRA Fine-Tuning | Specialized clinical output | 1,000-5,000 examples | $500-$5,000 | 2-7 days |
| Full Fine-Tuning | Maximum domain adaptation | 10,000+ examples | $5,000-$50,000 | 1-4 weeks |
Clinical Data Preparation: The 80% of the Work
The quality of your fine-tuned model is determined almost entirely by the quality of your training data. In clinical settings, data preparation involves five stages that are each significantly more complex than their non-healthcare equivalents.
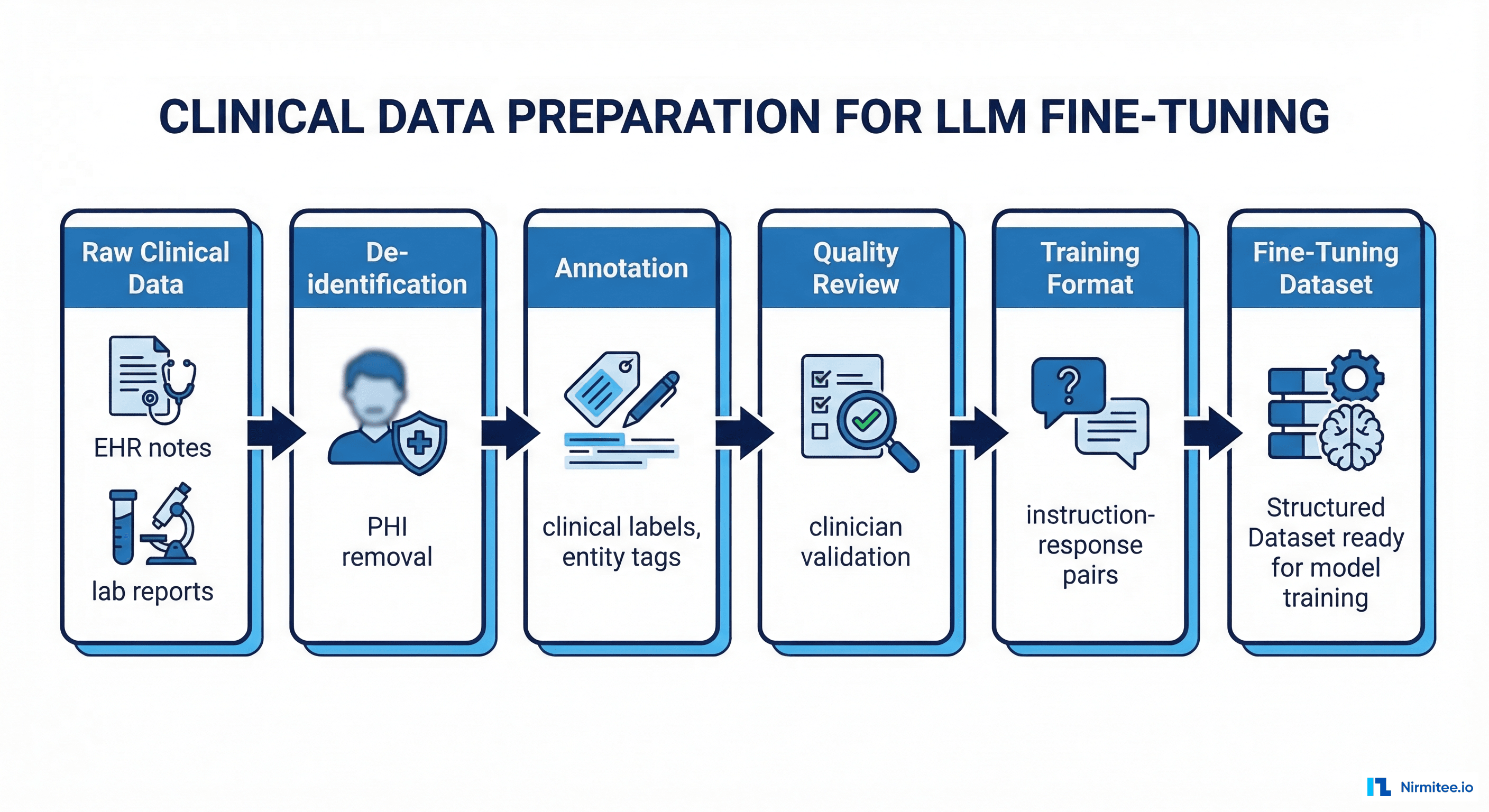
Stage 1: Data Collection from Clinical Sources
Clinical training data comes from three primary sources: EHR systems (clinical notes, discharge summaries, radiology reports), clinical data warehouses (structured diagnoses, medication histories, lab results), and published clinical literature (for supplementary domain knowledge).
The collection process must comply with your institution's IRB (Institutional Review Board) requirements. Most clinical AI projects require either a formal IRB approval or a determination that the project qualifies as quality improvement (exempt from full IRB review). Do not skip this step — training on clinical data without proper authorization creates both legal liability and ethical concerns.
Stage 2: De-Identification
All clinical data must be de-identified before use in model training, per HIPAA Safe Harbor or Expert Determination standards. This means removing 18 categories of identifiers: names, dates, locations, MRNs, phone numbers, email addresses, SSNs, and more.
Automated de-identification tools achieve 95-98% recall, which sounds impressive until you calculate the risk: in a dataset of 50,000 clinical notes containing an average of 12 PHI instances each, 2% leakage means 12,000 PHI instances remaining in your training data. We use a two-pass approach:
# Two-pass de-identification pipeline
import presidio_analyzer
import presidio_anonymizer
from transformers import pipeline
def deidentify_clinical_note(text: str) -> str:
# Pass 1: Rule-based NER (high recall for structured PHI)
analyzer = presidio_analyzer.AnalyzerEngine()
results = analyzer.analyze(
text=text,
entities=["PERSON", "DATE_TIME", "LOCATION",
"PHONE_NUMBER", "EMAIL_ADDRESS", "US_SSN",
"MEDICAL_RECORD", "IP_ADDRESS"],
language="en"
)
anonymizer = presidio_anonymizer.AnonymizerEngine()
pass1 = anonymizer.anonymize(text=text, analyzer_results=results)
# Pass 2: Transformer-based NER (catches contextual PHI)
ner = pipeline("ner", model="obi/deid_roberta_i2b2")
entities = ner(pass1.text)
for ent in reversed(entities): # reverse to preserve positions
if ent["score"] > 0.85:
pass1_text = (
pass1.text[:ent["start"]] +
"[REDACTED]" +
pass1.text[ent["end"]:]
)
return pass1_textStage 3: Clinical Annotation
Fine-tuning requires instruction-response pairs. For clinical tasks, annotation means a domain expert (physician, nurse practitioner, or clinical informaticist) labels each example with the correct output. Annotation strategies vary by task:
- Clinical note summarization: Provide full discharge summary as input, clinician-written one-paragraph summary as output. Requires 2,000-5,000 pairs.
- Diagnosis coding: Clinical note as input, ICD-10 codes with supporting evidence spans as output. Requires 5,000-10,000 pairs due to the large label space (70,000+ ICD-10 codes).
- Clinical question answering: Patient scenario as input, evidence-based clinical recommendation as output. Requires 1,000-3,000 pairs.
Annotation costs range from $15-$50 per example depending on clinical complexity and annotator qualifications. Budget $30,000-$100,000 for a production-quality annotated dataset.
Stage 4: Quality Assurance
Inter-annotator agreement (IAA) testing is non-negotiable for clinical AI. Have at least two clinicians annotate the same 200-300 examples independently, then measure Cohen's kappa. Target kappa > 0.75 before proceeding to training. If agreement is below this threshold, your annotation guidelines need refinement — the model cannot learn patterns that clinicians themselves disagree on.
Stage 5: Training Format Conversion
Convert annotated data into the instruction-following format required by your chosen base model. For most clinical fine-tuning with LoRA, this means Alpaca-style or ChatML formatting:
// Alpaca-style training example for clinical note summarization
{
"instruction": "Summarize the following discharge summary in 2-3 sentences, highlighting the primary diagnosis, key interventions, and follow-up plan.",
"input": "DISCHARGE SUMMARY\nPatient: [REDACTED], 67M\nAdmission Date: [DATE]\nDischarge Date: [DATE]\nPrimary Dx: Acute exacerbation of COPD (ICD-10: J44.1)\nHospital Course: Patient presented to ED with worsening dyspnea and productive cough x 3 days. ABG showed pH 7.31, pCO2 58. Started on BiPAP, IV methylprednisolone 125mg q6h, azithromycin 500mg daily. Transitioned to oral prednisone taper day 3. Weaned off BiPAP day 4. SpO2 94% on 2L NC at discharge.\nDischarge Medications: Prednisone 40mg taper, albuterol MDI, tiotropium, azithromycin x 3 more days.\nFollow-up: PCP in 1 week, pulmonology in 2 weeks, repeat PFTs in 4 weeks.",
"output": "67-year-old male admitted for acute COPD exacerbation with hypercapnic respiratory failure managed with BiPAP, IV steroids, and antibiotics. Clinical improvement allowed transition to oral medications and supplemental oxygen weaning by day 4. Discharged on steroid taper with pulmonology follow-up in 2 weeks and repeat pulmonary function testing at 4 weeks."
}Base Model Selection for Clinical Fine-Tuning
Choosing the right base model determines your ceiling for clinical performance, your infrastructure costs, and your regulatory pathway. Here is what the current landscape looks like for healthcare fine-tuning:
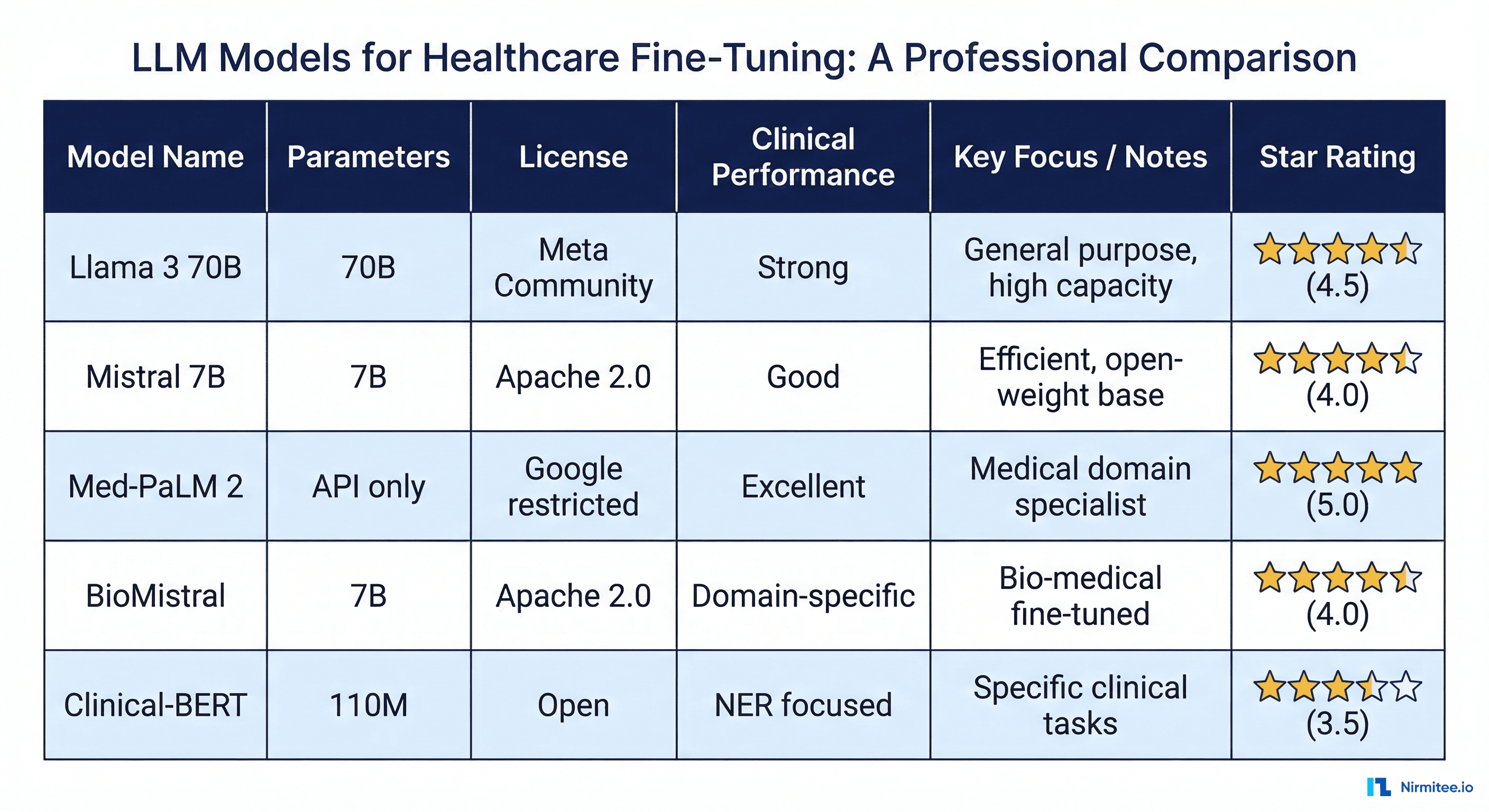
| Model | Parameters | License | Clinical Benchmark (MedQA) | Fine-Tuning Cost (LoRA) |
|---|---|---|---|---|
| Llama 3 70B | 70B | Meta Community | 78.2% | $2,000-$8,000 |
| Llama 3 8B | 8B | Meta Community | 62.5% | $300-$1,200 |
| Mistral 7B | 7B | Apache 2.0 | 58.9% | $200-$800 |
| BioMistral 7B | 7B | Apache 2.0 | 64.1% (pre-clinical) | $200-$800 |
| Gemma 2 27B | 27B | Google Permissive | 71.3% | $800-$3,000 |
Llama 3 70B delivers the strongest baseline performance and is our default recommendation for clinical fine-tuning projects where accuracy is the priority. The 70B parameter count requires multi-GPU setups (minimum 2x A100 80GB for LoRA fine-tuning), but the clinical accuracy gains over smaller models are substantial. We saw a 12-point improvement on our internal clinical benchmark suite compared to Llama 3 8B on a clinical note summarization task.
Mistral 7B and BioMistral 7B offer the best performance-per-dollar for resource-constrained projects. BioMistral comes pre-trained on biomedical literature (PubMed, PMC), giving it a head start on clinical vocabulary and reasoning. For tasks that primarily involve biomedical knowledge (drug interaction checking, clinical guideline retrieval), BioMistral fine-tunes faster and with less data than a general-purpose base model.
Practical recommendation: Start with LoRA fine-tuning on Llama 3 8B using 2,000 examples. Evaluate performance. If it meets your clinical threshold (typically 85%+ accuracy on your specific task), deploy it. If not, move to Llama 3 70B. Full fine-tuning of 70B models is rarely necessary — LoRA achieves 95-98% of full fine-tuning performance at a fraction of the cost.
Clinical Evaluation: Metrics That Actually Matter
Standard NLP evaluation metrics (BLEU, ROUGE, perplexity) are inadequate for clinical AI. A model can achieve high ROUGE scores on clinical note summarization while omitting critical medication allergies or misrepresenting diagnosis severity. Clinical evaluation requires task-specific metrics reviewed by domain experts.
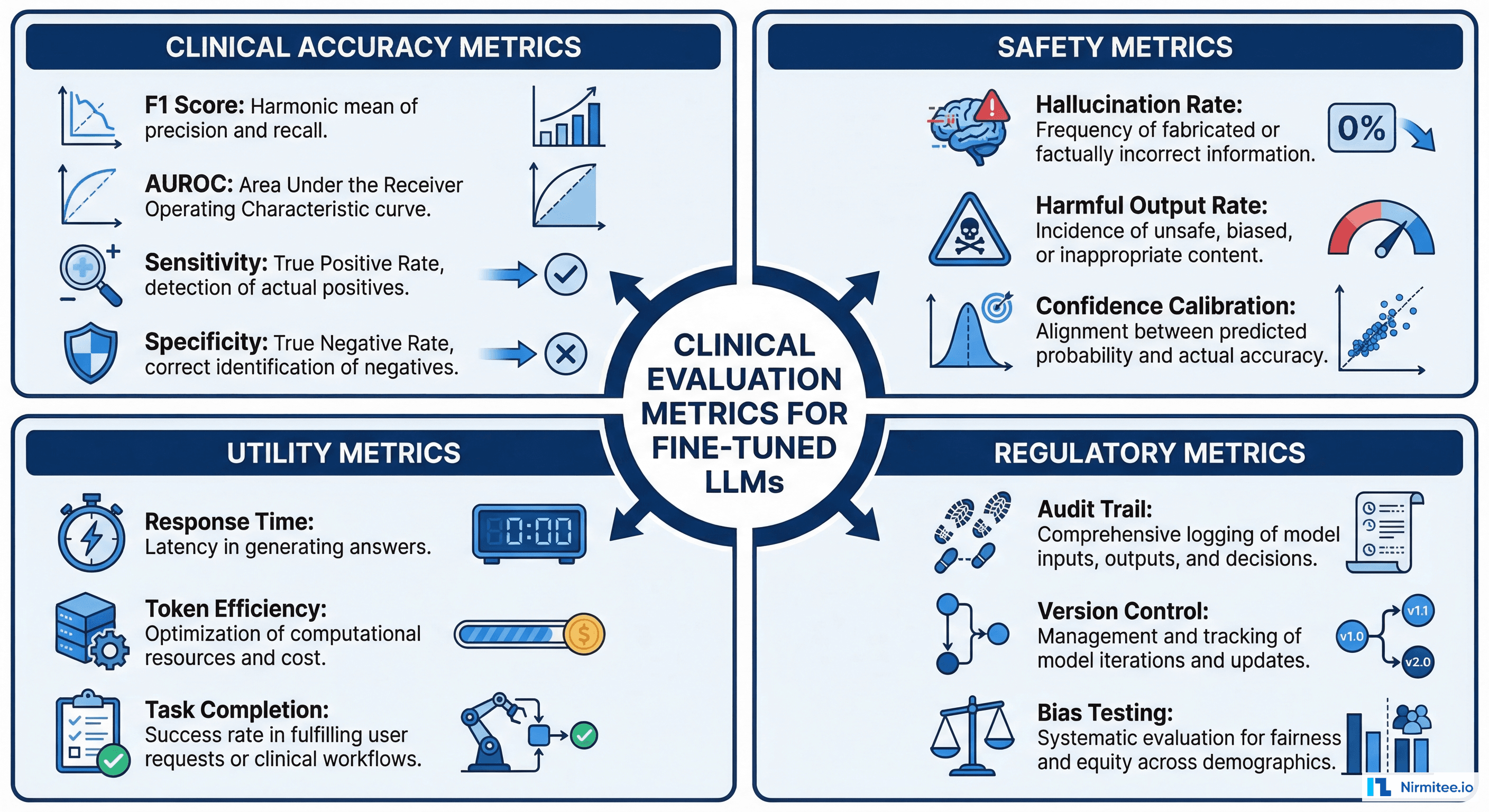
Clinical Accuracy Metrics
- Task-specific F1 score: For classification tasks (diagnosis coding, triage categorization), measure precision and recall on a held-out test set annotated by clinicians. Target F1 > 0.85 for clinical deployment.
- AUROC for risk prediction: For models that output risk scores (readmission prediction, deterioration alerts), AUROC provides a threshold-independent measure of discriminative ability. Target > 0.80, consistent with the benchmark improvements achieved through medallion architecture data pipelines.
- Clinical concordance: Have 3-5 clinicians independently rate model outputs on a 5-point clinical accuracy scale. Calculate Fleiss' kappa across raters. This is the most important metric because it measures whether clinicians trust the output enough to act on it.
Safety Metrics
- Hallucination rate: Percentage of outputs containing clinically fabricated information (medications not in the patient record, diagnoses not supported by available data). Measure by having clinicians flag hallucinations in 500+ model outputs. Target < 2% for clinical deployment.
- Harmful recommendation rate: Percentage of outputs containing recommendations that could cause patient harm if followed. This requires review by a senior clinician. Target: 0%. Any non-zero rate requires root cause analysis and model correction before deployment.
- Confidence calibration: When the model expresses uncertainty ("likely," "possible," "uncertain"), verify that its confidence aligns with actual accuracy. A well-calibrated model that says "likely" should be correct 70-80% of the time on those outputs.
Red Team Testing for Clinical AI
Before deploying any clinical LLM, conduct adversarial testing specifically designed for healthcare scenarios:
- Drug allergy edge cases: present the model with patients who have documented allergies and see if it ever recommends the allergen
- Pediatric dosing: present adult-appropriate medication doses for pediatric patients
- Pregnancy contraindications: present medications contraindicated in pregnancy for patients with pregnancy-related conditions
- Rare disease presentations: test with uncommon conditions that the training data may underrepresent
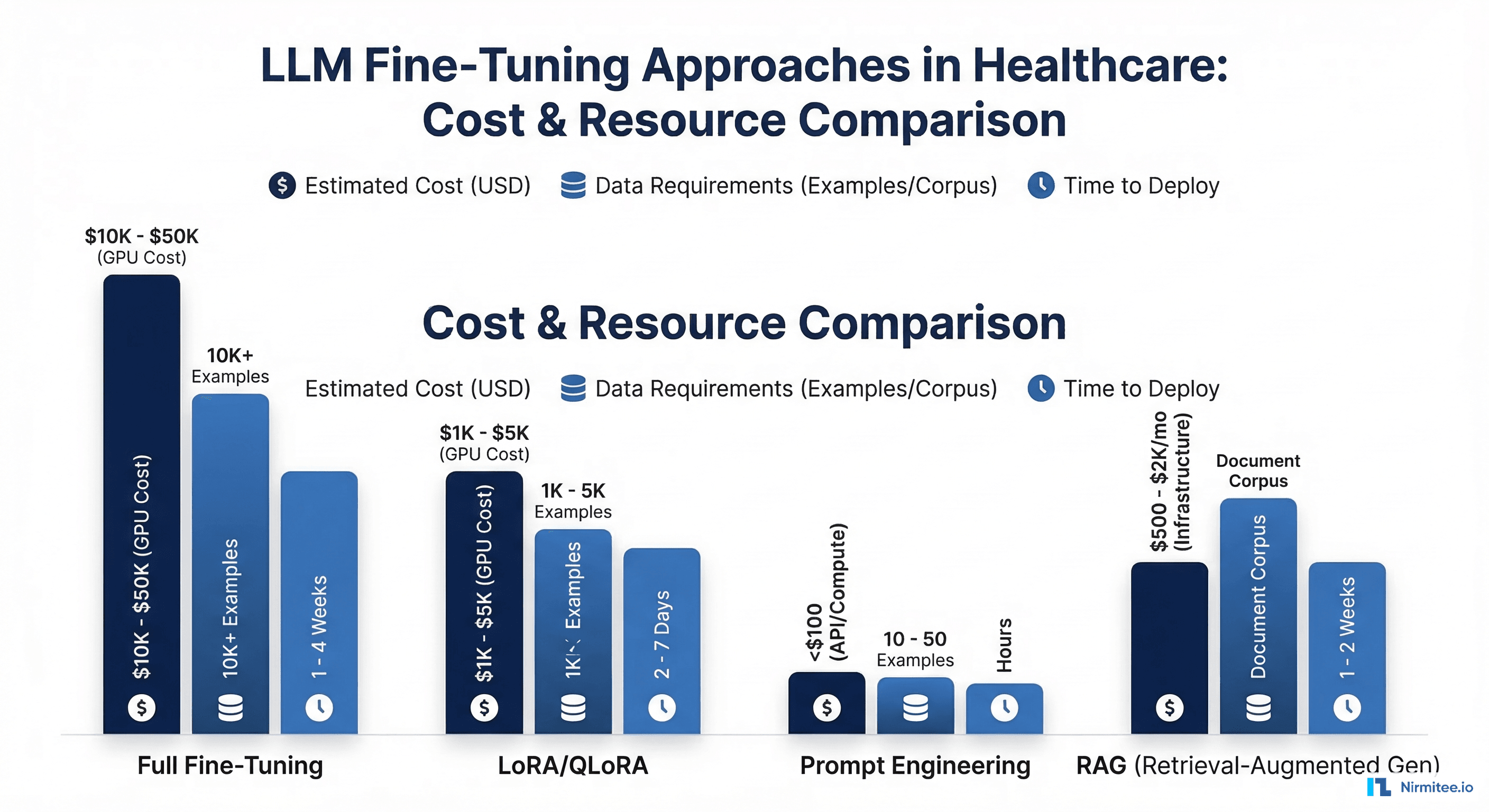
Cost Analysis: What Clinical Fine-Tuning Actually Costs
Here is a realistic budget breakdown for a clinical LLM fine-tuning project, based on our experience delivering these projects for healthtech companies:
Data preparation: $30,000-$100,000. This includes clinician annotation ($15-50 per example x 2,000-5,000 examples), de-identification tooling and validation, quality assurance with inter-annotator agreement testing, and IRB coordination.
Compute for training: $500-$8,000 for LoRA fine-tuning, $5,000-$50,000 for full fine-tuning. Cloud GPU costs on AWS (p4d instances with A100s) or GCP (a2-highgpu) run $25-$35 per hour. A LoRA fine-tune of Llama 3 8B on 3,000 examples takes approximately 4-8 hours. Llama 3 70B takes 24-72 hours.
Evaluation and testing: $10,000-$30,000. Clinical review of model outputs, red team testing, safety evaluation, and documentation for regulatory purposes.
Inference infrastructure: $500-$3,000 per month ongoing. Serving a 7B model requires a single GPU instance ($500-$800/month). Serving a 70B model requires 2-4 GPUs ($2,000-$3,000/month). Quantized deployments (GPTQ, AWQ) can reduce this by 40-60%.
Total project cost: $50,000-$190,000 for a LoRA fine-tuning project with production deployment, or $75,000-$280,000 for full fine-tuning.
FDA Regulatory Implications
The regulatory landscape for fine-tuned clinical LLMs is evolving rapidly. Here is the current framework that determines your compliance obligations:
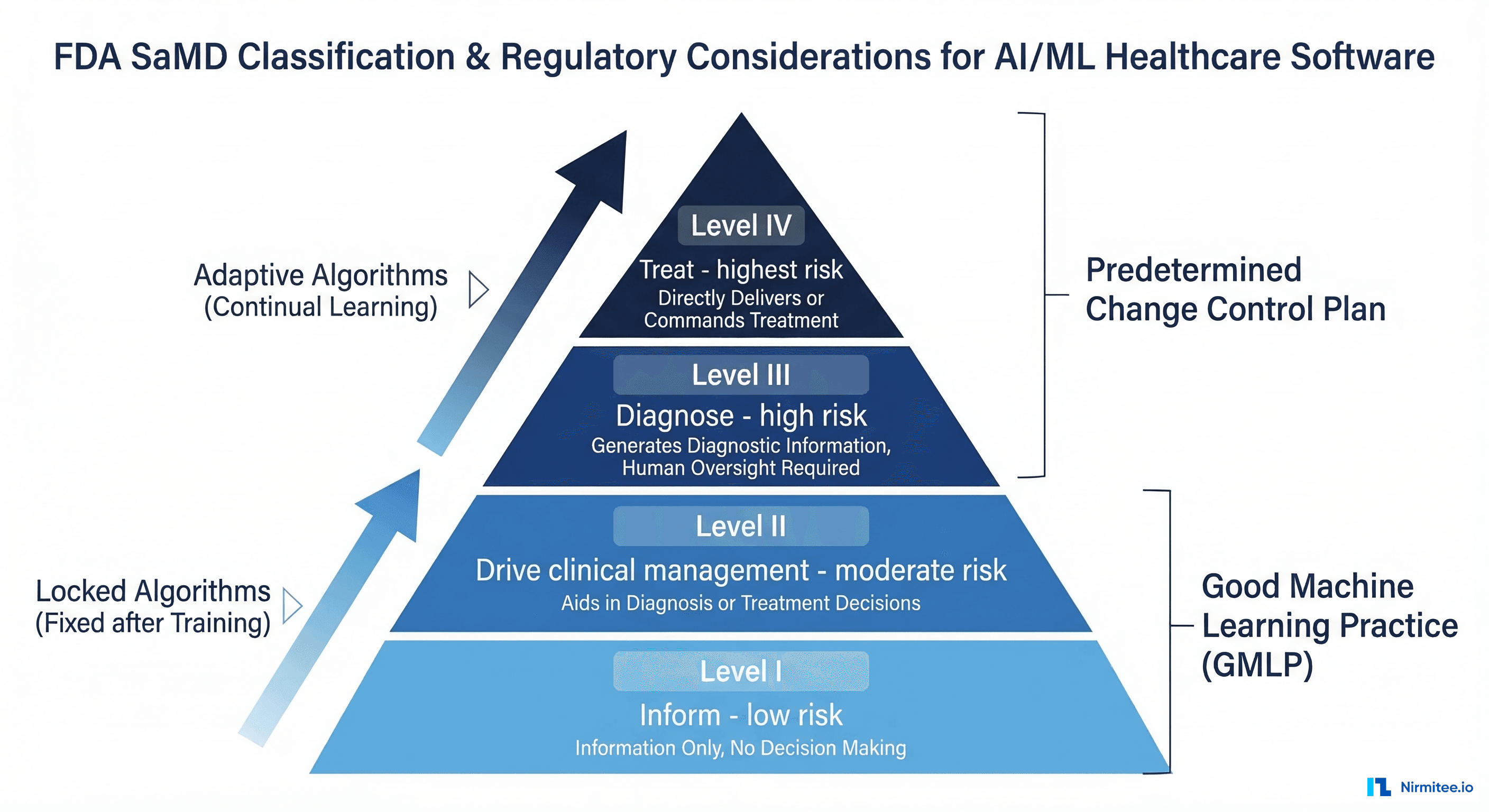
When Fine-Tuned Models Are SaMD
If your fine-tuned model generates outputs that directly influence clinical decisions — diagnosis suggestions, treatment recommendations, risk stratifications that trigger clinical actions — it is almost certainly Software as a Medical Device (SaMD) under FDA's framework.
The FDA classifies SaMD based on the seriousness of the healthcare situation and the significance of the software's contribution to the clinical decision. A fine-tuned model that suggests ICD-10 codes for clinician review (informational, low significance) has a lighter regulatory pathway than one that identifies sepsis risk and triggers automated care escalation (critical situation, high significance).
Predetermined Change Control Plan (PCCP)
The FDA's PCCP framework is specifically designed for AI/ML-based SaMD that improves over time through retraining. Your PCCP specifies:
- The types of modifications you plan to make (retraining on new data, architecture changes, performance improvements)
- Performance boundaries within which the model must operate
- Validation protocols for each modification type
- Real-world performance monitoring strategy
This is particularly relevant for fine-tuned models because you will retrain them as you collect more clinical data. Without a PCCP, each retraining cycle could trigger a new regulatory submission.
Good Machine Learning Practice (GMLP)
The FDA, Health Canada, and UK MHRA jointly published 10 GMLP principles that apply to all clinical AI development. For fine-tuning projects, the most operationally significant principles are:
- Multi-disciplinary expertise: Your team must include clinical domain experts, ML engineers, regulatory affairs, and quality assurance — not just data scientists.
- Representative data: Training data must reflect the intended patient population. A model fine-tuned on data from an academic medical center may perform differently in a community hospital setting.
- Independent test sets: Evaluation data must be strictly separated from training data with no data leakage.
- Ongoing monitoring: Post-deployment performance monitoring must detect model degradation before it impacts patient care.
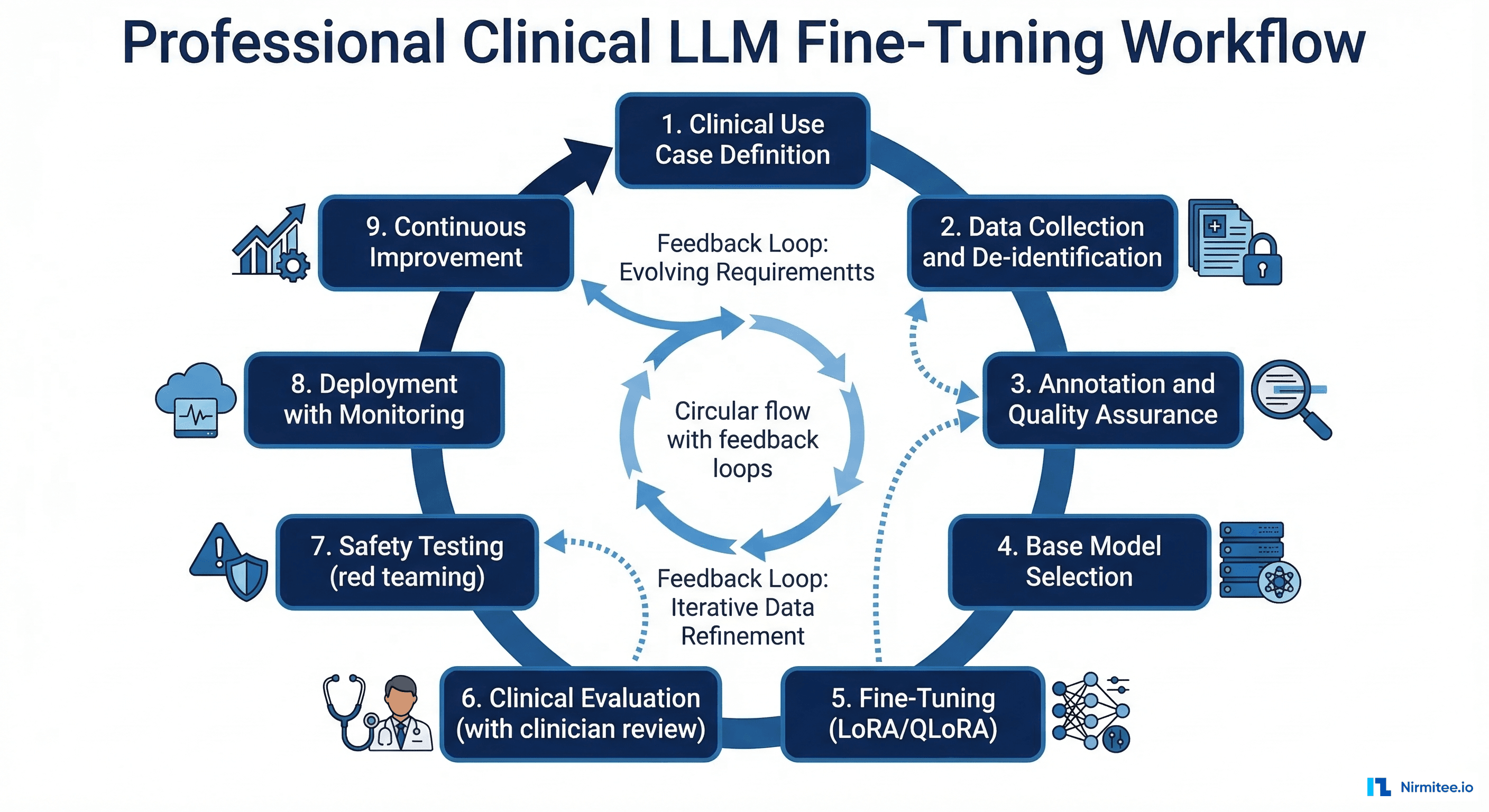
Production Deployment Patterns
Deploying a fine-tuned clinical model requires infrastructure patterns that differ from standard ML serving:
Human-in-the-loop architecture: Every clinical LLM deployment should include a human review step. The model generates suggestions; a clinician accepts, modifies, or rejects them. This is not a compromise — it is both a patient safety requirement and a regulatory expectation.
A/B testing with clinical guardrails: Roll out new model versions to a subset of users while monitoring clinical quality metrics. If the new model's hallucination rate exceeds the established baseline, automatically revert to the previous version.
Audit trail: Every model inference must be logged with the input, output, model version, and (when available) clinician action taken. This audit trail serves both regulatory compliance and continuous improvement — it generates the next round of training data through clinician feedback.
Inference monitoring: Track latency, throughput, error rates, and output distribution shifts in real time. A clinical model that suddenly starts generating unusual output distributions may be experiencing data drift from changes in the input EHR system.
Practical Roadmap: From Concept to Clinical Deployment
If you are planning a clinical LLM fine-tuning project, here is the realistic sequence:
- Weeks 1-2: Define the clinical use case with measurable success criteria. Get buy-in from a clinical champion who will provide domain expertise and annotator access.
- Weeks 3-6: Data collection, IRB coordination, and de-identification pipeline setup.
- Weeks 7-12: Clinical annotation with quality assurance. This is the slowest phase because it depends on clinician availability.
- Weeks 13-14: Base model selection and LoRA fine-tuning. Run experiments with 2-3 model sizes to determine the performance-cost tradeoff.
- Weeks 15-18: Clinical evaluation, safety testing, and red team exercises.
- Weeks 19-22: Deployment infrastructure, monitoring setup, and limited production rollout.
- Ongoing: Performance monitoring, periodic retraining, and regulatory documentation maintenance.
Total timeline: 5-6 months from concept to initial production deployment. This assumes you have access to clinical data and annotators. If IRB approval or data access is uncertain, add 2-4 months for those processes.
Nirmitee has delivered clinical AI projects ranging from multi-agent healthcare systems to fine-tuned models for fraud detection and clinical documentation. Our team combines ML engineering expertise with deep healthcare domain knowledge — the combination that clinical fine-tuning projects require. Talk to our AI team or explore our healthcare AI services to discuss your clinical LLM project.