Why Healthcare AI Needs More Than Git
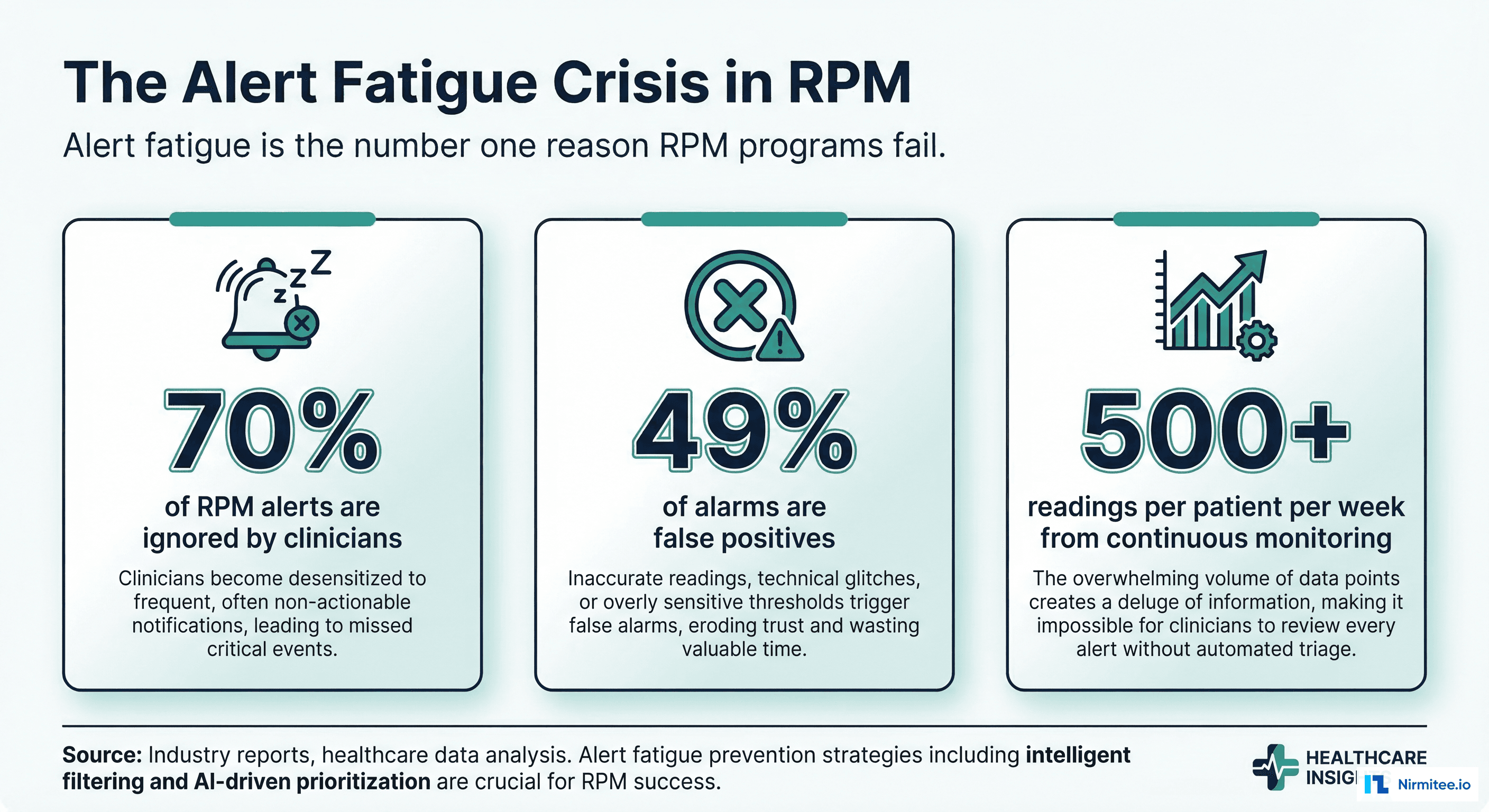
In standard software engineering, version control is a solved problem. Git tracks every line of code, every merge, every rollback. But clinical AI models are not code — they are trained artifacts shaped by data, hyperparameters, compute environments, and random seeds. A model's behavior is determined as much by its training data as by its architecture. And when that model influences clinical decisions — flagging sepsis risk, prioritizing radiology reads, recommending drug dosages — the regulatory bar for tracking changes is orders of magnitude higher than anything a typical engineering team has encountered.
The FDA's 2023 guidance on Predetermined Change Control Plans (PCCPs) for AI-enabled devices makes the requirement explicit: manufacturers must document what changes they plan to make to their models, how those changes will be validated, and what evidence demonstrates that the modified device remains safe and effective. This is not optional guidance — it is the regulatory framework that governs every AI/ML-based Software as a Medical Device (SaMD) on the US market.
A model registry is the system that makes this possible. It is not just a storage layer for model files — it is the single source of truth for what model is running in production, how it got there, who approved it, and what evidence supports that approval.
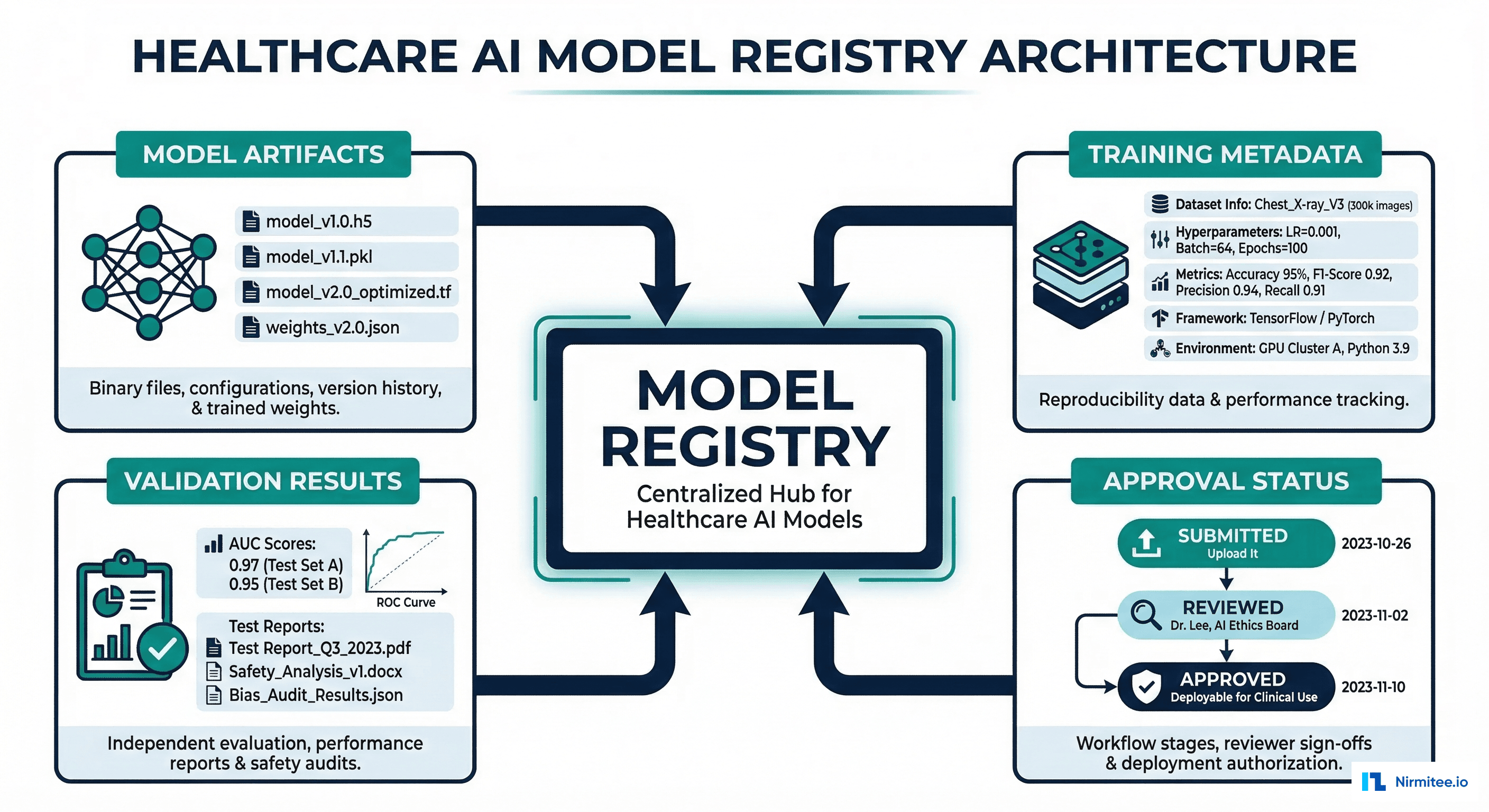
What a Healthcare Model Registry Must Store
Generic ML platforms like MLflow, Weights and Biases, and Neptune.ai handle experiment tracking and model versioning well. But none of them were built for the specific requirements of healthcare AI governance. A healthcare-grade model registry must track five categories of information that generic platforms miss.
1. Model Artifacts with Cryptographic Integrity
Every model version must be stored as an immutable artifact with a SHA-256 hash. The FDA requires that the exact model deployed in production can be reconstructed and re-evaluated at any point during a regulatory review — which may happen years after deployment. This means storing not just the model weights, but the complete inference package: serialized model, preprocessing code, feature engineering pipeline, and inference configuration.
2. Training Metadata and Data Lineage
For every model version, the registry must record: which training dataset was used (by version hash, not just name), the exact hyperparameters, the training infrastructure (GPU type, framework version, random seed), training duration, and convergence metrics. For healthcare, it must also record the data versioning information — the demographics breakdown of the training set, any exclusion criteria applied, and the data quality checks that passed.
3. Validation Results Segmented by Demographics
Aggregate performance metrics (overall AUC, accuracy) are insufficient for clinical models. The registry must store performance broken down by protected classes: age groups, sex, race/ethnicity, and clinical subpopulations. A model with AUC 0.92 overall but AUC 0.71 for Black patients has a bias problem that aggregate metrics hide. The FDA's Action Plan for AI/ML explicitly calls for evaluating algorithmic bias across demographic groups.
4. Approval Chain with Non-Repudiation
Every model version must have a documented approval chain: who submitted it, who reviewed it, who approved it, and when each action occurred. These approvals must be cryptographically signed and immutable — you cannot alter an approval record after the fact. This is the healthcare equivalent of a chain of custody.
5. Deployment and Rollback History
The registry must track every deployment event: when a model was promoted to production, which environment it was deployed to, the canary/shadow deployment results, and any rollback events with their justifications. If the FDA asks "what model was running on March 15th at 2:00 PM when this adverse event occurred?" — the registry must answer definitively.
Why Generic Platforms Fall Short
MLflow, Weights and Biases, and Neptune.ai are excellent tools for ML experiment tracking. But they were designed for tech companies shipping recommendation engines and ad targeting models — not for regulated medical devices.

| Capability | MLflow | W&B | Neptune.ai | Healthcare Requirement |
|---|---|---|---|---|
| Model versioning | Yes | Yes | Yes | Met |
| Experiment tracking | Yes | Yes | Yes | Met |
| Model staging (dev/staging/prod) | Yes | Partial | No | Partially met |
| Multi-step approval workflows | No | No | No | Not met |
| Clinical review committee integration | No | No | No | Not met |
| Demographic-segmented validation | No | No | No | Not met |
| FDA 21 CFR Part 11 audit trail | No | No | No | Not met |
| HIPAA-compliant access controls | Self-hosted only | BAA available | No | Partially met |
| Cryptographic approval signatures | No | No | No | Not met |
The solution is not to replace these platforms — it is to build a healthcare approval layer on top of them. MLflow remains the best open-source option as the model registry backend because of its mature API, model staging support, and self-hosting capability (critical for HIPAA). The approval workflow wraps around MLflow to add the governance layer that healthcare requires.
Building the Healthcare Approval Workflow
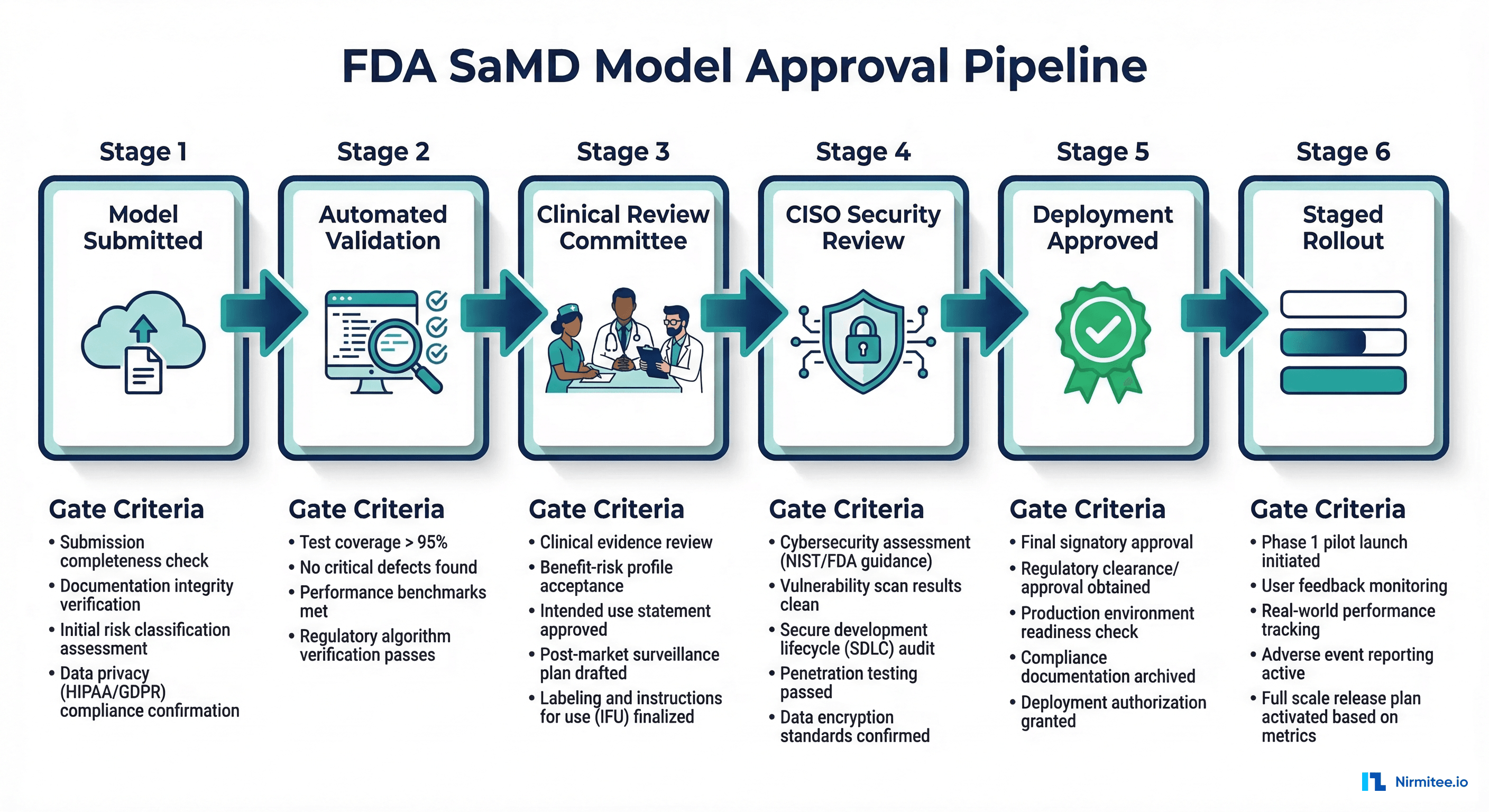
Here is the pattern for a healthcare-grade model approval pipeline built on top of MLflow. The workflow has six stages, each with specific gate criteria that must be satisfied before the model advances.
MLflow Registration with Healthcare Metadata
# healthcare_model_registry.py — Model Registration with Regulatory Metadata
import mlflow
from mlflow.tracking import MlflowClient
import hashlib
import json
from datetime import datetime
from typing import Dict, List, Optional
class HealthcareModelRegistry:
"""MLflow-based model registry with FDA-compliant metadata tracking."""
def __init__(self, tracking_uri: str, registry_uri: str):
mlflow.set_tracking_uri(tracking_uri)
self.client = MlflowClient(tracking_uri=tracking_uri,
registry_uri=registry_uri)
def register_clinical_model(
self,
model_name: str,
model_uri: str,
clinical_use_case: str,
training_data_version: str,
training_data_demographics: Dict,
validation_results: Dict,
model_card: Dict,
submitter_id: str,
) -> str:
"""Register a model with full healthcare metadata.
Returns the model version string.
"""
# Register model in MLflow
mv = mlflow.register_model(model_uri, model_name)
version = mv.version
# Compute artifact hash for integrity verification
artifact_hash = self._compute_model_hash(model_uri)
# Set healthcare-specific tags
tags = {
# Regulatory metadata
"healthcare.clinical_use_case": clinical_use_case,
"healthcare.artifact_sha256": artifact_hash,
"healthcare.submission_timestamp": datetime.utcnow().isoformat(),
"healthcare.submitter_id": submitter_id,
# Training data lineage
"healthcare.training_data_version": training_data_version,
"healthcare.training_data_demographics": json.dumps(
training_data_demographics
),
# Approval status
"healthcare.approval_status": "SUBMITTED",
"healthcare.approval_stage": "AWAITING_AUTOMATED_VALIDATION",
# Validation results (segmented by demographics)
"healthcare.validation_results": json.dumps(validation_results),
# Model card
"healthcare.model_card": json.dumps(model_card),
}
for key, value in tags.items():
self.client.set_model_version_tag(
model_name, version, key, value
)
print(f"Model {model_name} v{version} registered.")
print(f" Artifact SHA-256: {artifact_hash}")
print(f" Status: SUBMITTED -> AWAITING_AUTOMATED_VALIDATION")
return version
def run_automated_validation(
self,
model_name: str,
version: str,
min_auc: float = 0.85,
max_demographic_gap: float = 0.10,
) -> bool:
"""Gate 1: Automated validation checks.
- Overall AUC >= threshold
- No demographic subgroup more than max_gap below overall AUC
- No regression vs current production model
"""
mv = self.client.get_model_version(model_name, version)
results = json.loads(
mv.tags.get("healthcare.validation_results", "{}")
)
overall_auc = results.get("overall_auc", 0.0)
demographic_aucs = results.get("demographic_auc", {})
checks_passed = True
issues = []
# Check 1: Overall AUC threshold
if overall_auc < min_auc:
checks_passed = False
issues.append(
f"Overall AUC {overall_auc:.3f} below threshold {min_auc}"
)
# Check 2: Demographic fairness
for group, auc in demographic_aucs.items():
gap = overall_auc - auc
if gap > max_demographic_gap:
checks_passed = False
issues.append(
f"Demographic gap for {group}: {gap:.3f} "
f"(AUC={auc:.3f}, overall={overall_auc:.3f})"
)
# Check 3: No regression vs production
prod_versions = self.client.get_latest_versions(
model_name, stages=["Production"]
)
if prod_versions:
prod_auc = json.loads(
prod_versions[0].tags.get(
"healthcare.validation_results", "{}"
)
).get("overall_auc", 0.0)
if overall_auc < prod_auc - 0.02: # Allow 2% tolerance
checks_passed = False
issues.append(
f"Regression vs production: {overall_auc:.3f} "
f"vs {prod_auc:.3f}"
)
# Update status
new_status = "PASSED_AUTOMATED" if checks_passed else "FAILED_AUTOMATED"
new_stage = (
"AWAITING_CLINICAL_REVIEW" if checks_passed
else "REJECTED_AUTOMATED"
)
self.client.set_model_version_tag(
model_name, version,
"healthcare.approval_status", new_status
)
self.client.set_model_version_tag(
model_name, version,
"healthcare.approval_stage", new_stage
)
self.client.set_model_version_tag(
model_name, version,
"healthcare.automated_validation_issues",
json.dumps(issues)
)
return checks_passed
def clinical_review_decision(
self,
model_name: str,
version: str,
reviewer_id: str,
decision: str, # "APPROVED" or "REJECTED"
comments: str,
):
"""Gate 2: Clinical review committee decision."""
timestamp = datetime.utcnow().isoformat()
self.client.set_model_version_tag(
model_name, version,
"healthcare.clinical_reviewer_id", reviewer_id
)
self.client.set_model_version_tag(
model_name, version,
"healthcare.clinical_review_decision", decision
)
self.client.set_model_version_tag(
model_name, version,
"healthcare.clinical_review_timestamp", timestamp
)
self.client.set_model_version_tag(
model_name, version,
"healthcare.clinical_review_comments", comments
)
if decision == "APPROVED":
self.client.set_model_version_tag(
model_name, version,
"healthcare.approval_stage", "AWAITING_SECURITY_REVIEW"
)
else:
self.client.set_model_version_tag(
model_name, version,
"healthcare.approval_stage", "REJECTED_CLINICAL"
)
def promote_to_production(
self,
model_name: str,
version: str,
deployer_id: str,
deployment_strategy: str = "canary",
):
"""Final gate: promote model to production after all approvals."""
mv = self.client.get_model_version(model_name, version)
stage = mv.tags.get("healthcare.approval_stage", "")
if stage != "APPROVED_FOR_DEPLOYMENT":
raise ValueError(
f"Model not approved. Current stage: {stage}"
)
# Archive current production model
prod_versions = self.client.get_latest_versions(
model_name, stages=["Production"]
)
for pv in prod_versions:
self.client.transition_model_version_stage(
model_name, pv.version, "Archived"
)
# Promote new version
self.client.transition_model_version_stage(
model_name, version, "Production"
)
self.client.set_model_version_tag(
model_name, version,
"healthcare.deployment_timestamp",
datetime.utcnow().isoformat()
)
self.client.set_model_version_tag(
model_name, version,
"healthcare.deployer_id", deployer_id
)
self.client.set_model_version_tag(
model_name, version,
"healthcare.deployment_strategy", deployment_strategy
)
@staticmethod
def _compute_model_hash(model_uri: str) -> str:
"""Compute SHA-256 hash of model artifacts."""
import os
sha256 = hashlib.sha256()
model_path = mlflow.artifacts.download_artifacts(model_uri)
for root, dirs, files in os.walk(model_path):
for fname in sorted(files):
fpath = os.path.join(root, fname)
with open(fpath, "rb") as f:
for chunk in iter(lambda: f.read(8192), b""):
sha256.update(chunk)
return sha256.hexdigest()
Healthcare Model Cards
Google introduced Model Cards in 2019 as a standardized way to document ML model characteristics. For healthcare, the model card needs significant extensions to satisfy regulatory requirements and clinical stakeholder needs.
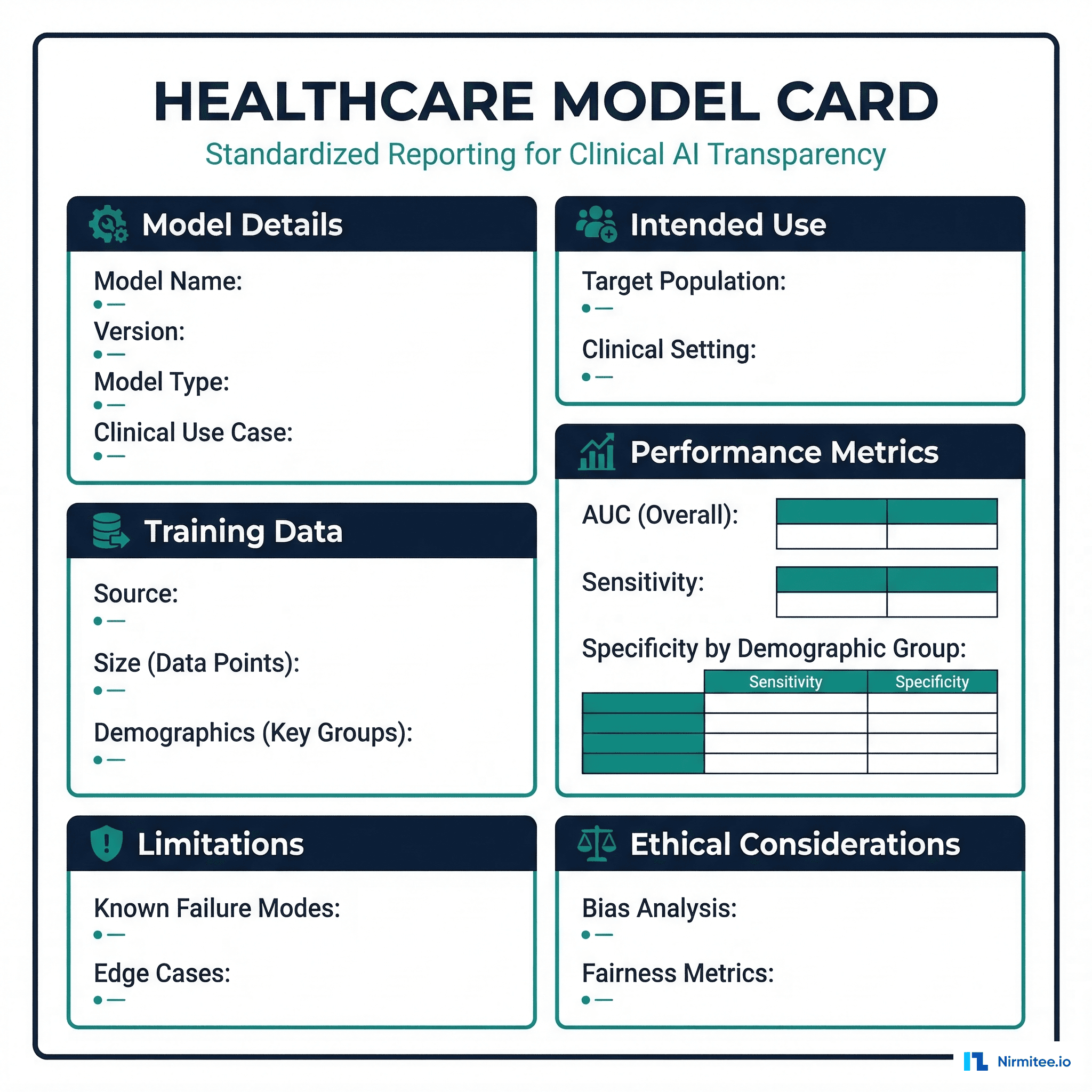
Here is a model card template adapted for healthcare AI:
# healthcare_model_card.py — Model Card Template for Clinical AI
def generate_model_card(
model_name: str,
version: str,
model_type: str,
clinical_use_case: str,
training_data: dict,
performance: dict,
limitations: list,
) -> dict:
"""Generate a healthcare-adapted model card."""
return {
"model_details": {
"name": model_name,
"version": version,
"type": model_type,
"framework": "PyTorch 2.1",
"clinical_use_case": clinical_use_case,
"fda_classification": "Class II SaMD",
"predicate_device": "K231234 (if applicable)",
},
"intended_use": {
"primary_use": "Clinical decision support for 30-day "
"readmission risk stratification",
"target_population": "Adult patients (18+) discharged "
"from acute care hospitals",
"clinical_setting": "Inpatient discharge planning",
"intended_users": ["Hospitalists", "Discharge planners",
"Care coordinators"],
"out_of_scope": ["Pediatric patients", "Psychiatric "
"admissions", "Planned readmissions"],
},
"training_data": {
"source": training_data.get("source", "Multi-site EHR"),
"size": training_data.get("size", "150,000 encounters"),
"date_range": training_data.get("range", "2020-2024"),
"demographics": {
"age_mean": 62.3,
"age_std": 15.7,
"female_pct": 52.1,
"race_distribution": {
"White": 58.2,
"Black": 22.1,
"Hispanic": 12.4,
"Asian": 4.8,
"Other": 2.5,
},
},
"exclusions": ["Patients < 18 years",
"In-hospital mortality",
"Transfers to other acute care"],
},
"performance_metrics": {
"overall": {
"auc_roc": performance.get("overall_auc", 0.89),
"sensitivity_at_90_specificity": 0.72,
"ppv_at_10_threshold": 0.34,
"calibration_slope": 1.02,
},
"by_demographic": performance.get("demographic_auc", {}),
"by_clinical_subgroup": {
"heart_failure": 0.91,
"copd": 0.88,
"diabetes": 0.87,
"surgical": 0.85,
},
},
"limitations": limitations or [
"Performance degrades for patients with < 2 prior "
"encounters in the system",
"Not validated for psychiatric readmissions",
"Training data predominantly from urban academic "
"centers; rural generalizability unconfirmed",
],
"ethical_considerations": {
"bias_analysis": "Demographic parity gap < 0.05 across "
"all racial groups for FPR",
"fairness_metric": "Equalized odds within 5% across "
"age and race groups",
"human_oversight": "Model output is advisory only; "
"all clinical decisions require "
"physician review",
},
}
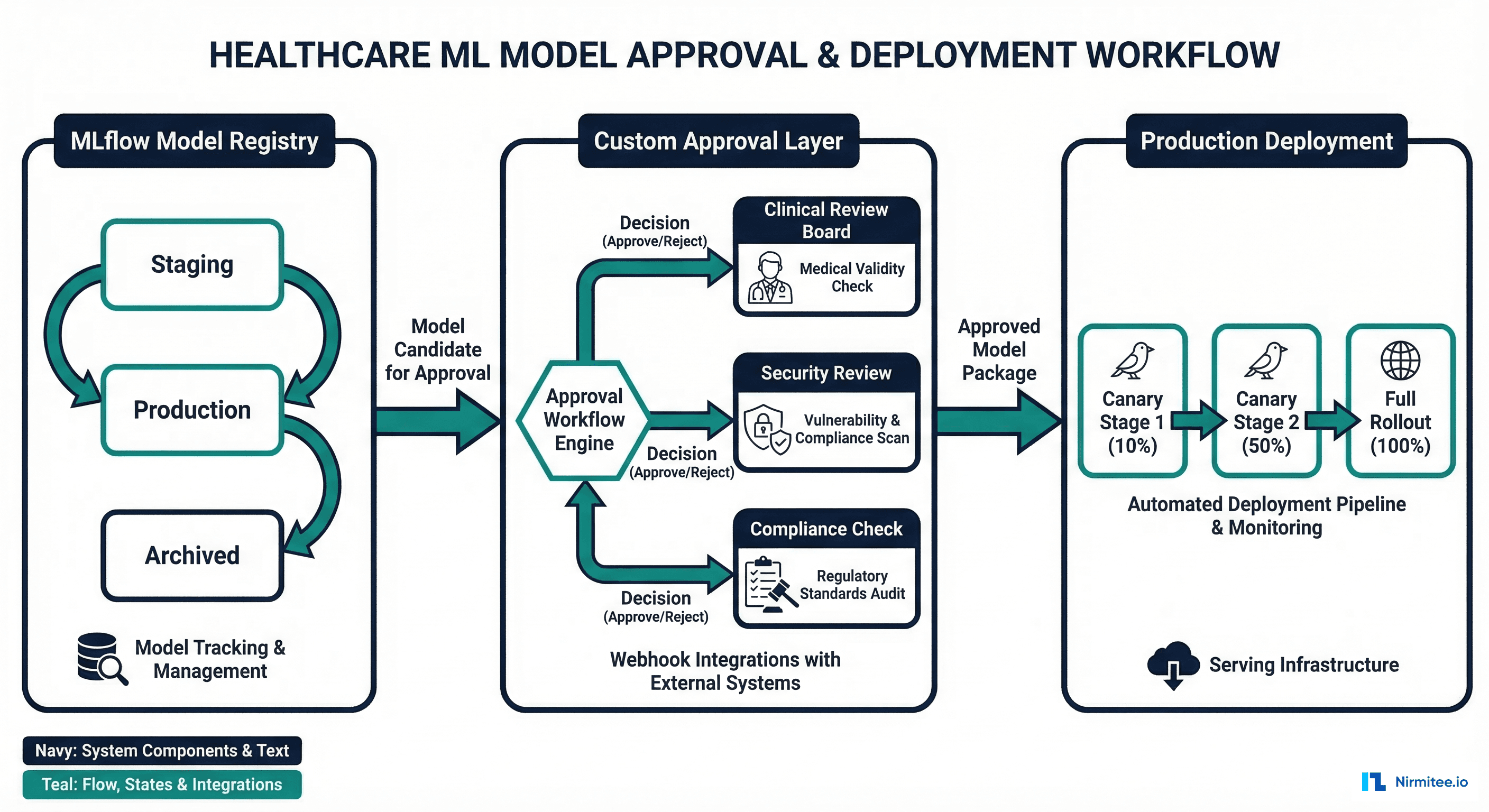
The Six-Stage Approval Pipeline
A healthcare model registry must enforce a multi-stage approval pipeline. No model reaches production without passing every gate. This is not bureaucracy — it is the process that protects patients and satisfies regulators.
Stage 1: Model Submission
The data science team submits a trained model to the registry with all required metadata. The system validates completeness — if the model card is missing fields, if validation results are not segmented by demographics, or if the training data version is not recorded, the submission is rejected automatically.
Stage 2: Automated Validation
A test suite runs automatically against the submitted model. The suite includes performance benchmarks (AUC thresholds), fairness checks (demographic parity), regression tests (comparison to current production model), and adversarial robustness tests. For a CI/CD pipeline perspective on automating these checks, see our guide on CI/CD for healthcare ML.
Stage 3: Clinical Review Committee
A committee of clinicians (typically the CMIO, a domain-expert physician, and a clinical informaticist) reviews the model card, validation results, and clinical relevance. They assess whether the model's performance translates to clinical value — a statistically significant AUC improvement may not matter if it does not change clinical workflows.
Stage 4: CISO Security Review
The security team reviews the model for adversarial vulnerabilities, data leakage risks (can the model memorize and regurgitate training data?), and infrastructure security of the deployment environment.
Stage 5: Deployment Approved
With all reviews complete, the model receives deployment approval. The approval record includes all reviewer decisions, timestamps, and the deployment strategy (canary, shadow, or full rollout).
Stage 6: Staged Rollout
The model is deployed using a canary strategy: 5% of traffic for 48 hours, then 25%, then 100%. At each stage, monitoring dashboards track prediction distribution, latency, error rates, and clinical outcome proxies. Any anomaly triggers an automatic rollback to the previous production model.
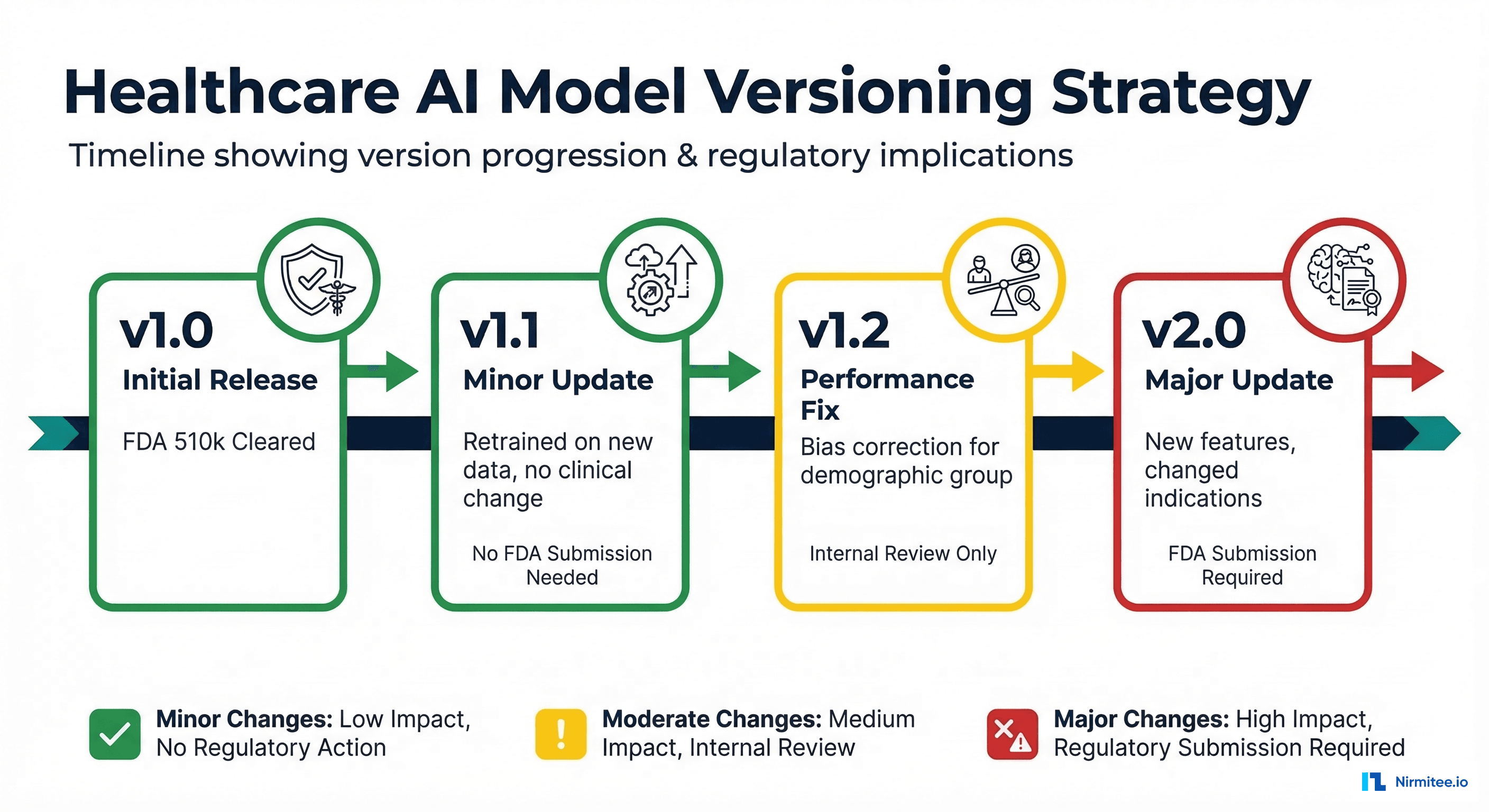
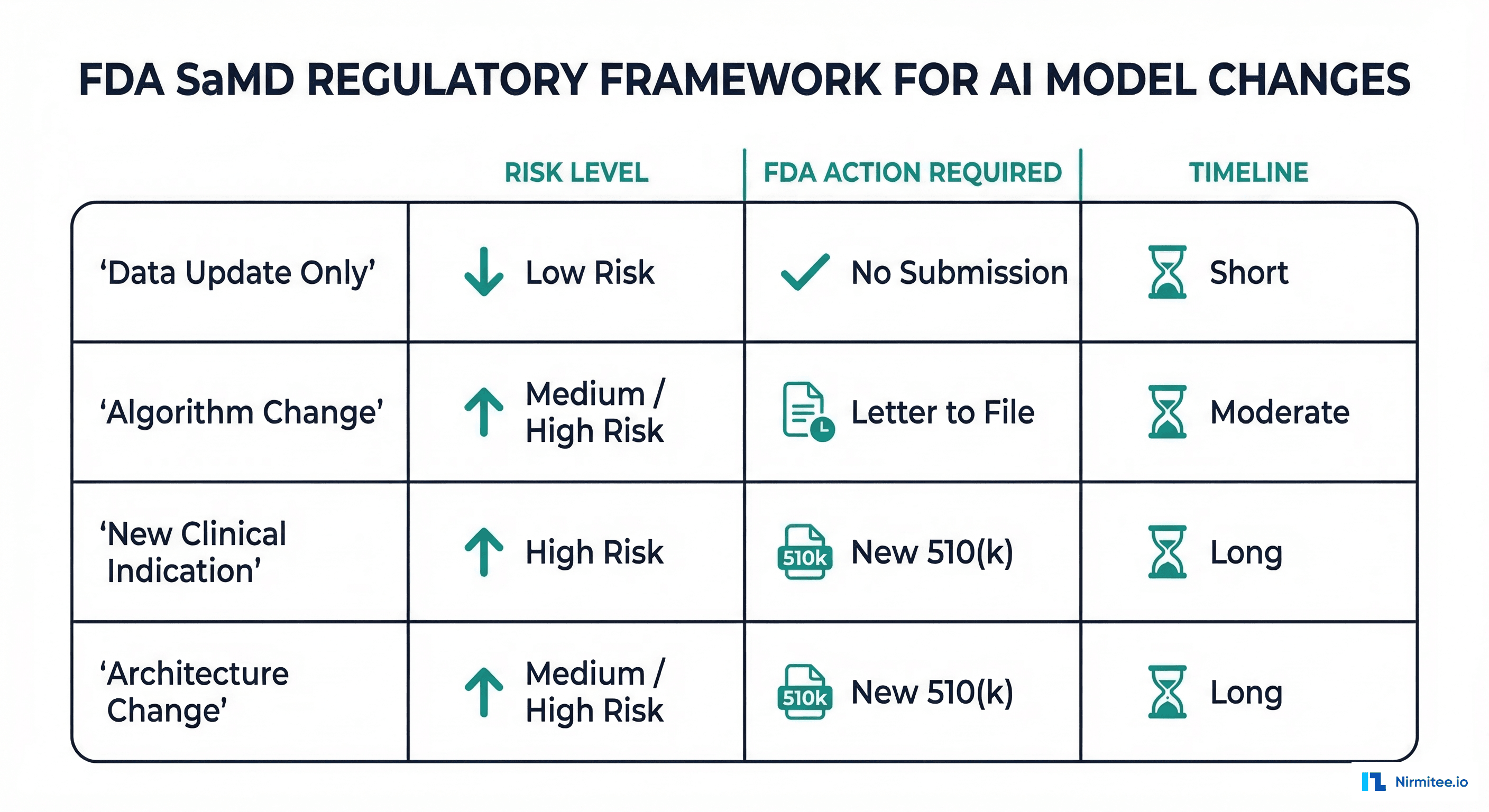
Version Classification for Regulatory Compliance
Not every model update requires the same level of scrutiny. The FDA's PCCP framework distinguishes between changes that need a new regulatory submission and changes that can be managed through internal processes.
| Change Type | Example | Risk Level | FDA Action | Internal Process |
|---|---|---|---|---|
| Data update only | Retrained on 6 months of new data, same architecture | Low | None (if within PCCP) | Full automated validation + clinical review |
| Hyperparameter tuning | Changed learning rate, batch size, regularization | Low | None (if within PCCP) | Automated validation + clinical sign-off |
| Feature engineering | Added new lab values or clinical features | Medium | Letter to file | Full pipeline + CISO review |
| Architecture change | Switched from logistic regression to neural network | High | New 510(k) likely | Full pipeline + external validation |
| New clinical indication | Expanded from readmission to mortality prediction | High | New 510(k) required | Full pipeline + new clinical studies |
Audit Trail Requirements
FDA 21 CFR Part 11 governs electronic records and signatures for regulated industries. While originally designed for pharmaceutical manufacturing, its principles apply to healthcare AI model management. The key requirements are:
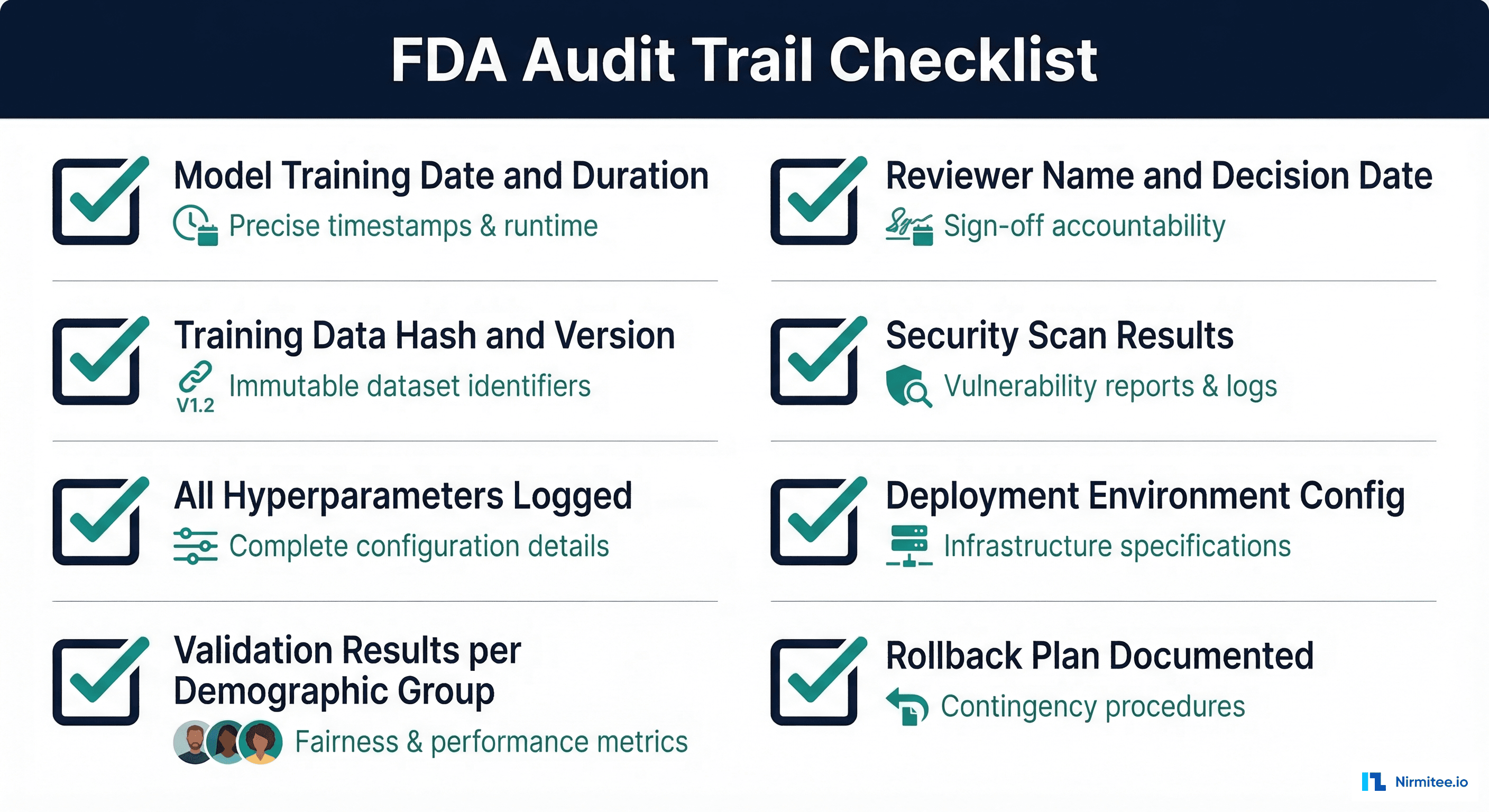

- Immutability: Records cannot be modified or deleted after creation. Use append-only storage (event sourcing pattern).
- Attribution: Every action is attributed to a specific user with authenticated identity.
- Timestamping: All events have accurate, tamper-proof timestamps (use a trusted time source, not local system clock).
- Reason for change: Every status transition must include a documented rationale.
- Access controls: Role-based access ensures only authorized personnel can approve models or modify registry records.
Implementation Recommendations
Based on the patterns above, here is the recommended technology stack for a healthcare model registry:
- Model storage: MLflow Model Registry (self-hosted on your HIPAA-compliant infrastructure, never SaaS)
- Experiment tracking: MLflow Tracking Server with PostgreSQL backend for durability
- Approval workflow: Custom service (Python/Go) that wraps MLflow with approval gates, built on an event-sourcing architecture for immutable audit trails
- Notifications: Integration with Slack/Teams/email for review requests and approval notifications
- Deployment: Integration with your Kubernetes-based ML serving infrastructure for staged rollouts
- Monitoring: Post-deployment drift detection feeds back into the registry, triggering retraining workflows when performance degrades
Frequently Asked Questions
Do we really need a custom approval layer, or can MLflow's built-in staging suffice?
MLflow's staging (Staging/Production/Archived) is a model lifecycle management feature, not an approval workflow. It has no concept of multi-step reviews, role-based approvals, or audit trails. For non-regulated ML applications, MLflow staging is sufficient. For healthcare SaMD, you need the custom layer.
How does the PCCP framework affect model registry design?
The FDA's Predetermined Change Control Plan requires that you define in advance what types of changes your model will undergo, how you will validate them, and what evidence you will generate. Your model registry must classify every change according to the PCCP categories and automatically route it through the appropriate approval path.
Should model cards be static documents or dynamic registry entries?
Dynamic. A model card should be auto-generated from the registry metadata whenever requested, not manually maintained as a separate document. This ensures the model card always reflects the current state of the model and eliminates the risk of documentation drift.
How do you handle rollbacks when the previous model version had known issues?
Maintain a "rollback target" tag in the registry that identifies the last known-good version for each model. This may not always be the immediately preceding version — if v3 had bias issues and v4 introduced a regression, the rollback target might be v2. The rollback target is updated as part of the deployment approval process.
What happens when a model is retrained on federated data from multiple institutions?
The registry must track the data lineage for each participating institution separately: dataset version, sample size, and demographic breakdown per site. For more on this topic, see our guide on federated learning for healthcare. The model card should disclose which institutions contributed to training and how their data was weighted.
Conclusion
A model registry for healthcare AI is not a nice-to-have — it is a regulatory requirement for any organization deploying clinical AI models. The gap between what generic ML platforms offer and what healthcare demands is significant: multi-stage approval workflows, demographic-segmented validation, cryptographic audit trails, and FDA-compliant version control.
The pattern outlined here — MLflow as the storage and versioning backend, wrapped with a custom approval layer that enforces healthcare governance — provides a practical path forward. It leverages the best open-source tooling while adding the regulatory compliance layer that the healthcare industry demands. Organizations that build this infrastructure now will be positioned to scale their clinical AI programs with confidence, knowing that every model in production has been validated, reviewed, and approved through a process that satisfies both clinical stakeholders and regulatory bodies.